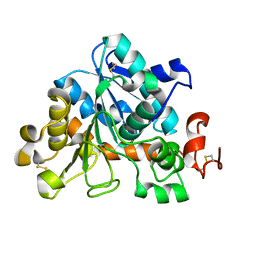
4ZV7
 
 | |
8ECE
 | | E. coli L-asparaginase II mutant (V27T) in complex with L-Glu | | Descriptor: | 1,2-ETHANEDIOL, GLUTAMIC ACID, L-asparaginase 2 | | Authors: | Strzelczyk, P, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2022-09-01 | | Release date: | 2022-11-16 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | The E. coli L-asparaginase V27T mutant: structural and functional characterization and comparison with theoretical predictions.
Febs Lett., 596, 2022
|
|
8ECD
 
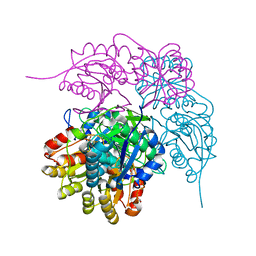 | | E. coli L-asparaginase II mutant (V27T) in complex with L-Asp | | Descriptor: | 1,2-ETHANEDIOL, ASPARTIC ACID, CITRIC ACID, ... | | Authors: | Strzelczyk, P, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2022-09-01 | | Release date: | 2022-11-16 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | The E. coli L-asparaginase V27T mutant: structural and functional characterization and comparison with theoretical predictions.
Febs Lett., 596, 2022
|
|
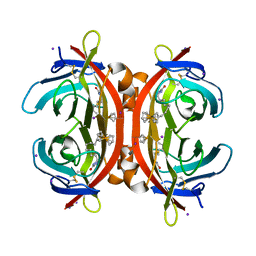
5MYQ
 
 | |
5CHK
 
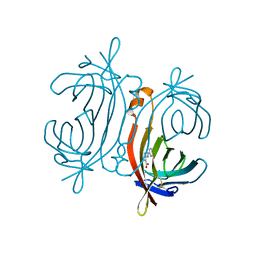 | | Crystal structure of avidin - HABA complex (hexagonal crystal form) | | Descriptor: | 2-[2-(4-oxocyclohexa-2,5-dien-1-ylidene)hydrazinyl]benzoic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, Avidin | | Authors: | Strzelczyk, P, Bujacz, A, Bujacz, G. | | Deposit date: | 2015-07-10 | | Release date: | 2016-03-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure and ligand affinity of avidin in the complex with 4-hydroxyazobenzene-2-carboxylic acid
J.Mol.Struct., 1109, 2016
|
|
5IRW
 
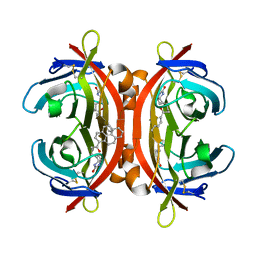 | |
5IRU
 
 | | Crystal structure of avidin in complex with 1-biotinylpyrene | | Descriptor: | 1-biotinylpyrene, 2-acetamido-2-deoxy-beta-D-glucopyranose, Avidin | | Authors: | Strzelczyk, P, Bujacz, G. | | Deposit date: | 2016-03-14 | | Release date: | 2016-10-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Characterization of the Avidin Interactions with Fluorescent Pyrene-Conjugates: 1-Biotinylpyrene and 1-Desthiobiotinylpyrene.
Molecules, 21, 2016
|
|
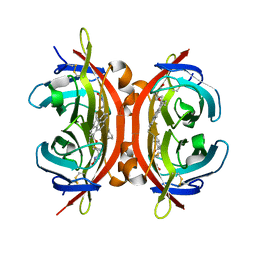
4I60
 
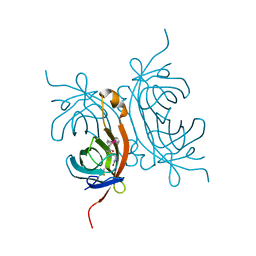 | | Crystal structure of avidin - biotinylruthenocene complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Avidin, [(1,2,3,4,5-eta)-cyclopentadienyl][(1,2,3,4,5-eta)-{5-[(3aS,4S,6aR)-2-oxohexahydro-1H-thieno[3,4-d]imidazol-4-yl]pentanoyl}cyclopentadienyl]ruthenium | | Authors: | Strzelczyk, P, Bujacz, A, Bujacz, G. | | Deposit date: | 2012-11-29 | | Release date: | 2013-08-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural investigation of the interactions of biotinylruthenocene with avidin.
Chem.Biol.Interact, 204, 2013
|
|
7R6A
 
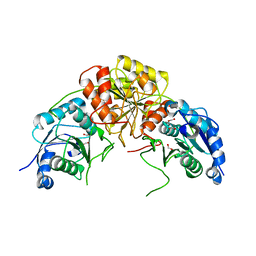 | |
7R6B
 
 | | Crystal structure of mutant R43D/L124D/R125A/C273S of L-Asparaginase I from Yersinia pestis | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, L-asparaginase I | | Authors: | Strzelczyk, P, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2021-06-22 | | Release date: | 2022-07-06 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | The dimeric form of bacterial l-asparaginase YpAI is fully active.
Febs J., 290, 2023
|
|
7R69
 
 | |
6WZ6
 | | Complex of mutant (K173M) of Pseudomonas 7A Glutaminase-Asparaginase with L-Glu at pH 5. Covalent acyl-enzyme intermediate | | Descriptor: | 1,2-ETHANEDIOL, GLUTAMIC ACID, Glutaminase-asparaginase | | Authors: | Strzelczyk, P, Zhang, D, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2020-05-13 | | Release date: | 2020-10-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Generalized enzymatic mechanism of catalysis by tetrameric L-asparaginases from mesophilic bacteria.
Sci Rep, 10, 2020
|
|
6WYW
 
 | | Crystal structure of Pseudomonas 7A Glutaminase-Asparaginase in complex with L-Asp at pH 4.5 | | Descriptor: | ASPARTIC ACID, Glutaminase-asparaginase | | Authors: | Strzelczyk, P, Zhang, D, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2020-05-13 | | Release date: | 2020-10-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.13 Å) | | Cite: | Generalized enzymatic mechanism of catalysis by tetrameric L-asparaginases from mesophilic bacteria.
Sci Rep, 10, 2020
|
|
6WYX
 
 | | Crystal structure of Pseudomonas 7A Glutaminase-Asparaginase in complex with L-Asp at pH 5.0 | | Descriptor: | ASPARTIC ACID, GLYCEROL, Glutaminase-asparaginase | | Authors: | Strzelczyk, P, Zhang, D, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2020-05-13 | | Release date: | 2020-10-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Generalized enzymatic mechanism of catalysis by tetrameric L-asparaginases from mesophilic bacteria.
Sci Rep, 10, 2020
|
|
6WZ8
 
 | | Complex of Pseudomonas 7A Glutaminase-Asparaginase with Citrate Anion at pH 5.5. Covalent Acyl-Enzyme Intermediate | | Descriptor: | CITRIC ACID, Glutaminase-asparaginase | | Authors: | Strzelczyk, P, Zhang, D, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2020-05-13 | | Release date: | 2020-10-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Generalized enzymatic mechanism of catalysis by tetrameric L-asparaginases from mesophilic bacteria.
Sci Rep, 10, 2020
|
|
6WYY
 
 | | Crystal structure of Pseudomonas 7A Glutaminase-Asparaginase in complex with L-Glu at pH 6.5 | | Descriptor: | 1,2-ETHANEDIOL, GLUTAMIC ACID, GLYCEROL, ... | | Authors: | Strzelczyk, P, Zhang, D, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2020-05-13 | | Release date: | 2020-10-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Generalized enzymatic mechanism of catalysis by tetrameric L-asparaginases from mesophilic bacteria.
Sci Rep, 10, 2020
|
|
6WZ4
 
 | | Complex of mutant (K173M) of Pseudomonas 7A Glutaminase-Asparaginase with L-Asp at pH 6. Covalent acyl-enzyme intermediate | | Descriptor: | ASPARTIC ACID, Glutaminase-asparaginase | | Authors: | Strzelczyk, P, Zhang, D, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2020-05-13 | | Release date: | 2020-10-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Generalized enzymatic mechanism of catalysis by tetrameric L-asparaginases from mesophilic bacteria.
Sci Rep, 10, 2020
|
|
6WYZ
 
 | | Crystal structure of Pseudomonas 7A Glutaminase-Asparaginase (mutant K173M) in complex with D-Glu at pH 5.5 | | Descriptor: | D-GLUTAMIC ACID, Glutaminase-asparaginase | | Authors: | Strzelczyk, P, Zhang, D, Wlodawer, A, Lubkowski, J. | | Deposit date: | 2020-05-13 | | Release date: | 2020-10-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Generalized enzymatic mechanism of catalysis by tetrameric L-asparaginases from mesophilic bacteria.
Sci Rep, 10, 2020
|
|
5HLM
 
 | | Crystal structure of avidin complex with a ferrocene biotin derivative | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Avidin, [(1,2,3,4,5-eta)-cyclopentadienyl]{(1,2,3,4,5-eta)-1-[1-hydroxy-5-(2-oxohexahydro-1H-thieno[3,4-d]imidazol-4-yl)pentyl]cyclopentadienyl}iron | | Authors: | Strzelczyk, P, Bujacz, G. | | Deposit date: | 2016-01-15 | | Release date: | 2017-01-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Ferrocene-Biotin Conjugates: Synthesis, Structure, Cytotoxic Activity and Interaction with Avidin
Chempluschem, 81, 2016
|
|
4JHQ
 
 | | Crystal structure of avidin - 6-(6-biotinamidohexanamido)hexanoylferrocene complex | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Avidin, [(1,2,3,4,5-eta)-cyclopentadienyl][(1,2,3,4,5-eta)-{6-[(6-{[5-(2-oxohexahydro-1H-thieno[3,4-d]imidazol-4-yl)pentanoyl]amino}hexanoyl)amino]hexanoyl}cyclopentadienyl]iron | | Authors: | Strzelczyk, P, Bujacz, A, Bujacz, G. | | Deposit date: | 2013-03-05 | | Release date: | 2013-11-20 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Ferrocene-Biotin Conjugates Targeting Cancer Cells: Synthesis, Interaction with Avidin, Cytotoxic Properties and the Crystal Structure of the Complex of Avidin with a Biotin-Linker-Ferrocene Conjugate
Organometallics, 32, 2013
|
|
