4AV9
 
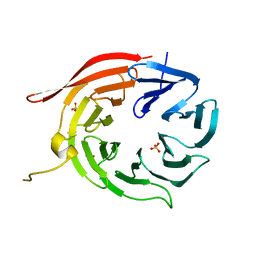 | | Kluyveromyces lactis Hsv2 | | Descriptor: | SULFATE ION, SVP1-LIKE PROTEIN 2 | | Authors: | Krick, R, Busse, R.A, Scacioc, A, Stephan, M, Janshoff, A, Thumm, M, Kuhnel, K. | | Deposit date: | 2012-05-24 | | Release date: | 2012-06-13 | | Last modified: | 2012-08-15 | | Method: | X-RAY DIFFRACTION (3.001 Å) | | Cite: | Structural and Functional Characterization of the Two Phosphoinositide Binding Sites of Proppins, a Beta-Propeller Protein Family.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4AV8
 
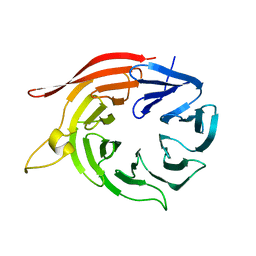 | | Kluyveromyces lactis Hsv2 complete loop 6CD | | Descriptor: | SVP1-LIKE PROTEIN 2 | | Authors: | Krick, R, Busse, R.A, Scacioc, A, Stephan, M, Janshoff, A, Thumm, M, Kuhnel, K. | | Deposit date: | 2012-05-24 | | Release date: | 2012-06-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.35 Å) | | Cite: | Structural and Functional Characterization of the Two Phosphoinositide Binding Sites of Proppins, a Beta-Propeller Protein Family.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4RZ2
 
 | |
4RZ3
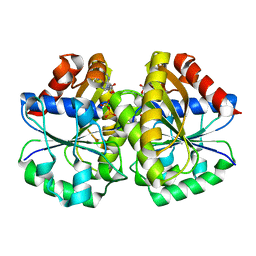 
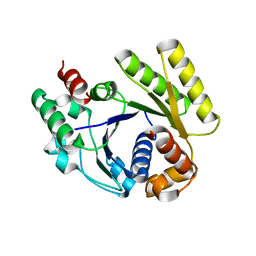
 | | Crystal structure of the MinD-like ATPase FlhG | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Site-determining protein | | Authors: | Schuhmacher, J.S, Bange, G. | | Deposit date: | 2014-12-18 | | Release date: | 2015-03-04 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | MinD-like ATPase FlhG effects location and number of bacterial flagella during C-ring assembly.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
