6QK8
 
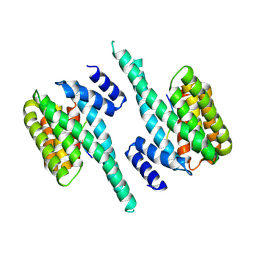 | |
5M4A
 
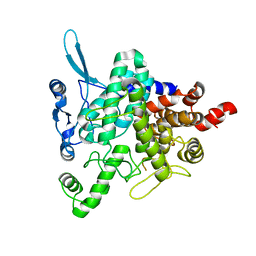 | | Neutral trehalase Nth1 from Saccharomyces cerevisiae in complex with trehalose | | Descriptor: | Neutral trehalase, alpha-D-glucopyranose-(1-1)-alpha-D-glucopyranose | | Authors: | Smidova, A, Alblova, M, Obsilova, V, Obsil, T. | | Deposit date: | 2016-10-18 | | Release date: | 2017-11-01 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Molecular basis of the 14-3-3 protein-dependent activation of yeast neutral trehalase Nth1.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
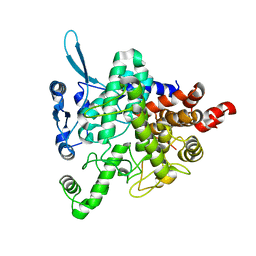
5JTA
 
 | |
5N6N
 
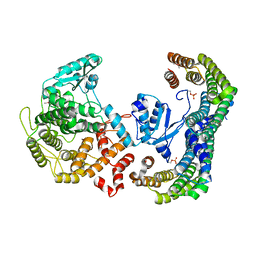 | | CRYSTAL STRUCTURE OF THE 14-3-3:NEUTRAL TREHALASE NTH1 COMPLEX | | Descriptor: | CALCIUM ION, Neutral trehalase, Protein BMH1, ... | | Authors: | Alblova, M, Smidova, A, Obsilova, V, Obsil, T. | | Deposit date: | 2017-02-15 | | Release date: | 2017-11-01 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Molecular basis of the 14-3-3 protein-dependent activation of yeast neutral trehalase Nth1.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5NIS
 
 | |
6GKF
 
 | |
6GKG
 
 | |
