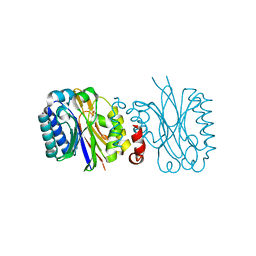
5JQN
 
 | |
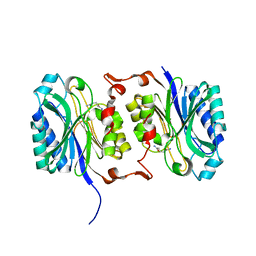
6YPA
 
 | |
4LF0
 
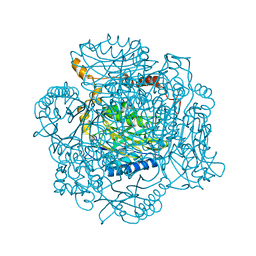 | | The E142D mutant of the amidase from Geobacillus pallidus | | Descriptor: | Aliphatic amidase | | Authors: | Sewell, B.T, Weber, B.W, Kimani, S.W, Cowan, D.A, Hunter, R, Venter, G.A, Gumbart, J.C, Thuku, R.N, Varsani, A. | | Deposit date: | 2013-06-26 | | Release date: | 2013-08-21 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | The mechanism of the amidases: mutating the glutamate adjacent to the catalytic triad inactivates the enzyme due to substrate mispositioning.
J.Biol.Chem., 288, 2013
|
|
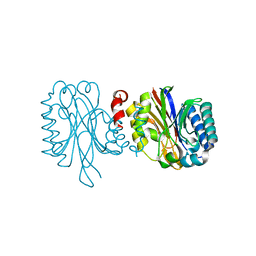
3HKX
 
 | |
8P4I
 
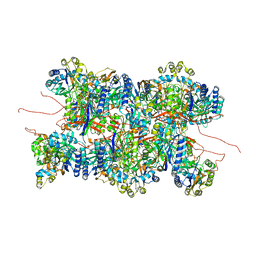 | | Cyanide dihydratase from Bacillus pumilus C1 | | Descriptor: | Cyanide dihydratase | | Authors: | Mulelu, A.E, Reitz, J, van Rooyen, J.M, Scheffer, M, Frangakis, A.S, Dlamini, L.S, Woodward, J.D, Benedik, M.J, Sewell, B.T. | | Deposit date: | 2023-05-22 | | Release date: | 2023-08-16 | | Method: | ELECTRON MICROSCOPY (3.83 Å) | | Cite: | The Role of Histidine Residues in the Oligomerization of Cyanide Dihydratase from Bacillus pumilus C1
To Be Published
|
|
7Z0V
 
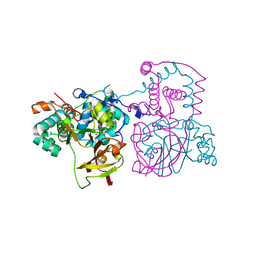 | | A mutant of the nitrile hydratase from Geobacillus pallidus having enhanced thermostability | | Descriptor: | CHLORIDE ION, COBALT (II) ION, Nitrile hydratase, ... | | Authors: | Van Wyk, J.C, Cowan, D.A, Danson, M.J, Tsekoa, T.L, Sayed, M.F, Sewell, B.T. | | Deposit date: | 2022-02-23 | | Release date: | 2023-03-08 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Engineering enhanced thermostability into the Geobacillus pallidus nitrile hydratase.
Curr Res Struct Biol, 4, 2022
|
|
9ER3
 
 | |
3O6X
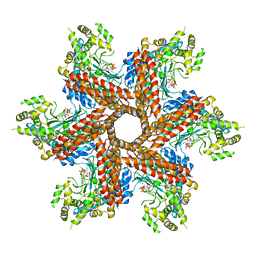 
 | | Crystal Structure of the type III Glutamine Synthetase from Bacteroides fragilis | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CHLORIDE ION, Glutamine synthetase, ... | | Authors: | van Rooyen, J.M, Belrhali, H, Abratt, V.R, Sewell, B.T. | | Deposit date: | 2010-07-29 | | Release date: | 2011-03-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Crystal Structure of Type III Glutamine Synthetase: Surprising Reversal of the Inter-Ring Interface.
Structure, 19, 2011
|
|
8C5I
 
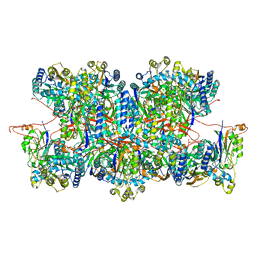 | | Cyanide dihydratase from Bacillus pumilus C1 variant - Q86R,H305K,H308K,H323K | | Descriptor: | Cyanide dihydratase | | Authors: | Mulelu, A.E, Reitz, J, van Rooyen, J, Scheffer, M, Frangakis, A.S, Dlamini, L.S, Woodward, J.D, Benedik, M.J, Sewell, B.T. | | Deposit date: | 2023-01-09 | | Release date: | 2023-01-18 | | Method: | ELECTRON MICROSCOPY (3.15 Å) | | Cite: | The Role of Histidine Residues in the Oligomerization of Cyanide Dihydratase from Bacillus pumilus C1
To Be Published
|
|
4APE
 
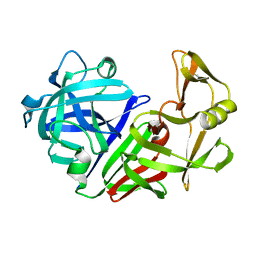 | | THE ACTIVE SITE OF ASPARTIC PROTEINASES | | Descriptor: | ENDOTHIAPEPSIN | | Authors: | Pearl, L.H, Sewell, B.T, Jenkins, J.A, Cooper, J.B, Blundell, T.L. | | Deposit date: | 1986-06-09 | | Release date: | 1986-07-14 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Active Site of Aspartic Proteinases
FEBS Lett., 174, 1984
|
|
2PLQ
 
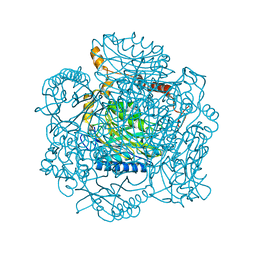 | | Crystal structure of the amidase from geobacillus pallidus RAPc8 | | Descriptor: | Aliphatic amidase | | Authors: | Kimani, S.W, Sewell, B.T, Agarkar, V.B, Sayed, M.F, Cowan, D.A. | | Deposit date: | 2007-04-20 | | Release date: | 2007-05-01 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The quaternary structure of the amidase from Geobacillus pallidus RAPc8 is
revealed by its crystal packing.
Acta Crystallogr.,Sect.F, 62, 2006
|
|
2IUL
 
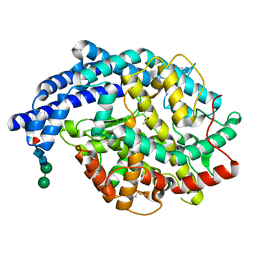 | | Human tACE g13 mutant | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, ANGIOTENSIN-CONVERTING ENZYME, ... | | Authors: | Watermeyer, J.M, Sewell, B.T, Natesh, R, Corradi, H.R, Acharya, K.R, Sturrock, E.D. | | Deposit date: | 2006-06-06 | | Release date: | 2006-10-25 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Structure of Testis Ace Glycosylation Mutants and Evidence for Conserved Domain Movement.
Biochemistry, 45, 2006
|
|
6ZBY
 
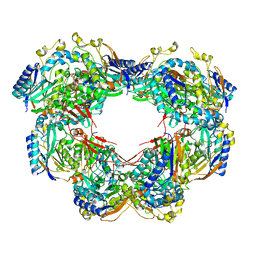 | |
4EWL
 
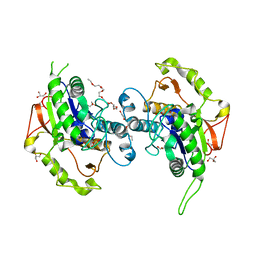 | | Crystal Structure of MshB with glycerol and Acetate bound in the active site | | Descriptor: | 1D-myo-inositol 2-acetamido-2-deoxy-alpha-D-glucopyranoside deacetylase, 2-{2-[2-(2-{2-[2-(2-ETHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, ACETATE ION, ... | | Authors: | Broadley, S.G, Sewell, B.T, Weber, B.W, Marakalala, M.J, Steenkamp, D.J. | | Deposit date: | 2012-04-27 | | Release date: | 2012-09-26 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | A new crystal form of MshB from Mycobacterium tuberculosis with glycerol and acetate in the active site suggests the catalytic mechanism.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3BKL
 
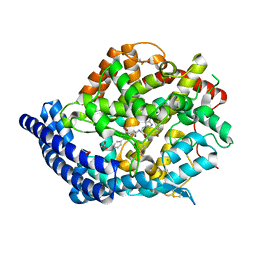 | | Testis ACE co-crystal structure with ketone ACE inhibitor kAW | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, ... | | Authors: | Watermeyer, J.M, Kroger, W.L, O'Neill, H.G, Sewell, B.T, Sturrock, E.D. | | Deposit date: | 2007-12-07 | | Release date: | 2008-06-10 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Probing the basis of domain-dependent inhibition using novel ketone inhibitors of Angiotensin-converting enzyme
Biochemistry, 47, 2008
|
|
3BKK
 
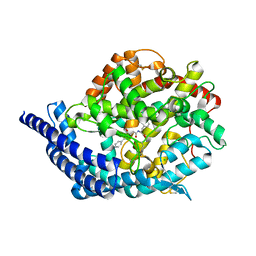 | | Tesis ACE co-crystal structure with ketone ACE inhibitor kAF | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme, ... | | Authors: | Watermeyer, J.M, Kroger, W.L, O'Neill, H.G, Sewell, B.T, Sturrock, E.D. | | Deposit date: | 2007-12-07 | | Release date: | 2008-06-10 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Probing the basis of domain-dependent inhibition using novel ketone inhibitors of Angiotensin-converting enzyme
Biochemistry, 47, 2008
|
|
2DPP
 
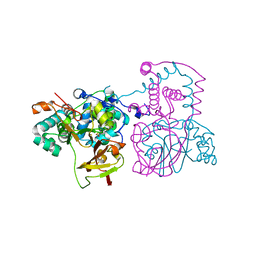 | | Crystal structure of thermostable Bacillus sp. RAPc8 nitrile hydratase | | Descriptor: | COBALT (II) ION, Nitrile hydratase alpha subunit, Nitrile hydratase beta subunit | | Authors: | Tsekoa, T.L, Tastan-Bishop, A.O, Cameron, R.A, Sewell, B.T, Sayed, M.F, Cowan, D.A. | | Deposit date: | 2006-05-12 | | Release date: | 2007-05-22 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.52 Å) | | Cite: | Crystal structure of thermostable Bacillus sp. RAPc8 nitrile hydratse
To be Published
|
|
3HGJ
 
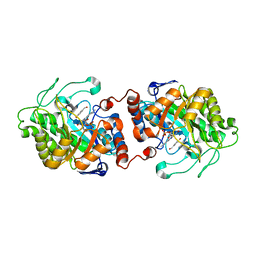 | | Old Yellow Enzyme from Thermus scotoductus SA-01 complexed with p-hydroxy-benzaldehyde | | Descriptor: | Chromate reductase, FLAVIN MONONUCLEOTIDE, P-HYDROXYBENZALDEHYDE | | Authors: | Opperman, D.J, Sewell, B.T, Litthauer, D, Isupov, M.N, Littlechild, J.A, van Heerden, E. | | Deposit date: | 2009-05-14 | | Release date: | 2010-02-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of a thermostable old yellow enzyme from Thermus scotoductus SA-01
Biochem.Biophys.Res.Commun., 393, 2010
|
|
3HHT
 
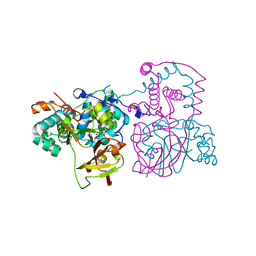 | | A mutant of the nitrile hydratase from Geobacillus pallidus having enhanced thermostability | | Descriptor: | CHLORIDE ION, COBALT (III) ION, Nitrile hydratase alpha subunit, ... | | Authors: | Van Wyk, J.C, Sewell, B.T, Cowan, D.A, Sayed, M.F, Tsekoa, T.L, Tastan Bishop, A.O. | | Deposit date: | 2009-05-17 | | Release date: | 2009-07-14 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.16 Å) | | Cite: | The high-resolution structure of a Cobalt containing nitrile hydratase
To be published
|
|
3HF3
 
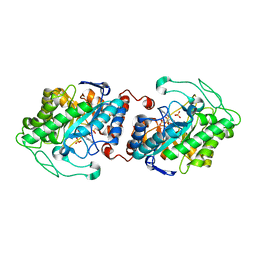 | | Old Yellow Enzyme from Thermus scotoductus SA-01 | | Descriptor: | Chromate reductase, FLAVIN MONONUCLEOTIDE, SULFATE ION | | Authors: | Opperman, D.J, Sewell, B.T, Litthauer, D, Isupov, M.N, Littlechild, J.A, van Heerden, E. | | Deposit date: | 2009-05-11 | | Release date: | 2010-02-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of a thermostable old yellow enzyme from Thermus scotoductus SA-01
Biochem.Biophys.Res.Commun., 393, 2010
|
|
3L3N
 
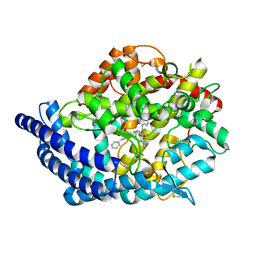 | | Testis ACE co-crystal structure with novel inhibitor lisW | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, Angiotensin-converting enzyme, CHLORIDE ION, ... | | Authors: | Watermeyer, J.M, Kroger, W.L, O'Neil, H.G, Sewell, B.T, Sturrock, E.D. | | Deposit date: | 2009-12-17 | | Release date: | 2010-04-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Characterization of domain-selective inhibitor binding in angiotensin-converting enzyme using a novel derivative of lisinopril.
Biochem.J., 428, 2010
|
|
6T1T
 
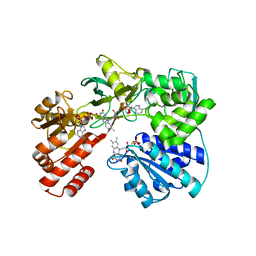 | |
6T1U
 
 | | Cytochrome P450 reductase from Candida tropicalis | | Descriptor: | FLAVIN MONONUCLEOTIDE, FLAVIN-ADENINE DINUCLEOTIDE, NADPH--cytochrome P450 reductase | | Authors: | Opperman, D.J, Sewell, B.T. | | Deposit date: | 2019-10-06 | | Release date: | 2020-01-08 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Biochemical and structural insights into the cytochrome P450 reductase from Candida tropicalis.
Sci Rep, 9, 2019
|
|
7QOP
 
 | | A mutant of the nitrile hydratase from Geobacillus pallidus having enhanced thermostability | | Descriptor: | CHLORIDE ION, COBALT (II) ION, MAGNESIUM ION, ... | | Authors: | Van Wyk, J.C, Cowan, D.A, Danson, M.J, Tsekoa, T.L, Sayed, M.F, Sewell, B.T. | | Deposit date: | 2021-12-24 | | Release date: | 2023-01-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Engineering enhanced thermostability into the Geobacillus pallidus nitrile hydratase.
Curr Res Struct Biol, 4, 2022
|
|
7QOV
 
 | | The wild type nitrile hydratase from Geobacillus pallidus | | Descriptor: | CHLORIDE ION, COBALT (III) ION, Nitrile hydratase, ... | | Authors: | Van Wyk, J.C, Cowan, D.A, Danson, M.J, Tsekoa, T.L, Sayed, M.F, Sewell, B.T. | | Deposit date: | 2021-12-29 | | Release date: | 2023-01-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Engineering enhanced thermostability into the Geobacillus pallidus nitrile hydratase.
Curr Res Struct Biol, 4, 2022
|
|
