2V9U
 
 | | Rim domain of main porin from Mycobacteria smegmatis | | Descriptor: | MSPA | | Authors: | Grueninger, D, Ziegler, M.O.P, Koetter, J.W.A, Treiber, N, Schulze, M.-S, Schulz, G.E. | | Deposit date: | 2007-08-27 | | Release date: | 2008-01-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Designed Protein-Protein Association.
Science, 319, 2008
|
|
8B7H
 
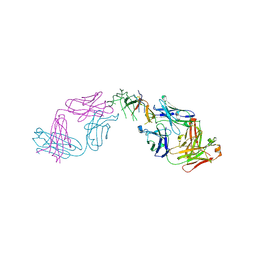 | |
8CDG
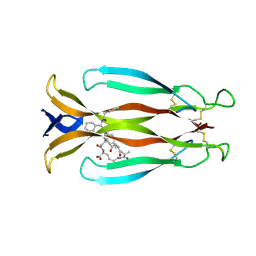 
 | | Crystal structure of human IL-17A cytokine in complex with macrocycle | | Descriptor: | (9~{S},12~{R},19~{S})-9-[[4-[[(2~{S})-2-cyclohexyl-2-(2-phenylethanoylamino)ethanoyl]amino]phenyl]methyl]-12-methyl-7,10,13,21-tetrakis(oxidanylidene)-8,11,14,20-tetrazaspiro[4.17]docosane-19-carboxylic acid, Interleukin-17A | | Authors: | Schulze, M.-S. | | Deposit date: | 2023-01-30 | | Release date: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Modulation of IL-17 backbone dynamics reduces receptor affinity and reveals a new inhibitory mechanism.
Chem Sci, 14, 2023
|
|
6GY1
 
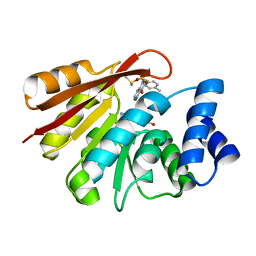 | | rat COMT in complex with inhibitor | | Descriptor: | 7-fluoranyl-5-(4-methylphenyl)sulfonyl-quinolin-8-ol, Catechol O-methyltransferase, DIMETHYL SULFOXIDE, ... | | Authors: | Schulze, M.-S. | | Deposit date: | 2018-06-27 | | Release date: | 2018-10-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Optimization of 8-Hydroxyquinolines as Inhibitors of Catechol O-Methyltransferase.
J. Med. Chem., 61, 2018
|
|
8P41
 
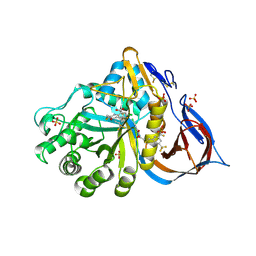 | | Crystal structure of glucocerebrosidase in complex with allosteric activator | | Descriptor: | 2-[2-[[3,5-bis(trifluoromethyl)phenyl]methylsulfanyl]ethanoylamino]-5-chloranyl-benzoic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Schulze, M.-S. | | Deposit date: | 2023-05-18 | | Release date: | 2024-03-06 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Identification of ss-Glucocerebrosidase Activators for Glucosylceramide hydrolysis.
Chemmedchem, 19, 2024
|
|
8P3E
 
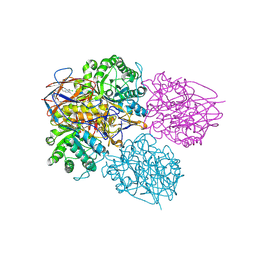 | | Crystal structure of glucocerebrosidase in complex with allosteric activator | | Descriptor: | 2-[[3-[(4-chlorophenyl)carbamoyl]phenyl]sulfonylamino]benzoic acid, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Schulze, M.-S. | | Deposit date: | 2023-05-17 | | Release date: | 2024-03-06 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Identification of ss-Glucocerebrosidase Activators for Glucosylceramide hydrolysis.
Chemmedchem, 19, 2024
|
|
6NDR
 
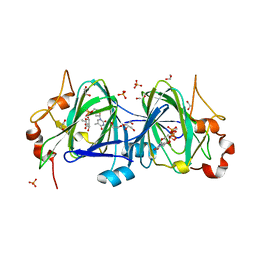 | |
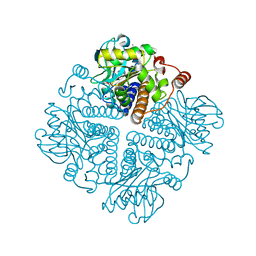
2V9F
 
 | |
2UYV
 
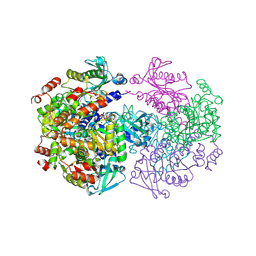 | |
2V9G
 
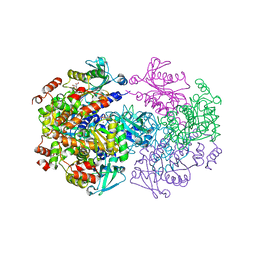 | |
2UYU
 
 | |
2V9I
 
 | |
2V9O
 
 | |
2V9E
 
 | |
2V9M
 
 | |
2V9L
 
 | |
2V7G
 
 | |
2V2B
 
 | |
