4UE3
 
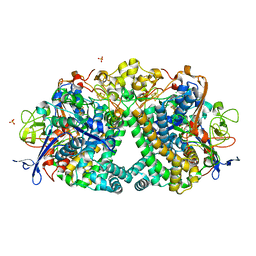 | | The Mechanism of Hydrogen Activation by NiFe-hydrogenases and the Importance of the active site Arginine | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, CHLORIDE ION, FE3-S4 CLUSTER, ... | | Authors: | Evans, R.M, Wehlin, S.A.M, Nomerotskaia, E, Sargent, F, Carr, S.B, Phillips, S.E.V, Armstrong, F.A. | | Deposit date: | 2014-12-15 | | Release date: | 2014-12-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Mechanism of Hydrogen Activation by [Nife] Hydrogenases.
Nat.Chem.Biol., 12, 2016
|
|
4C3O
 
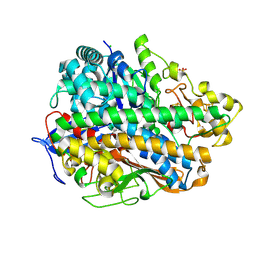 | | Structure and function of an oxygen tolerant NiFe hydrogenase from Salmonella | | Descriptor: | CHLORIDE ION, FE3-S4 CLUSTER, FE4-S3 CLUSTER, ... | | Authors: | Bowman, L, Flanagan, L, Fyfe, P.K, Parkin, A, Hunter, W.N, Sargent, F. | | Deposit date: | 2013-08-26 | | Release date: | 2014-01-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | How the Structure of the Large Subunit Controls Function in an Oxygen-Tolerant [Nife]-Hydrogenase.
Biochem.J., 458, 2014
|
|
1E5K
 
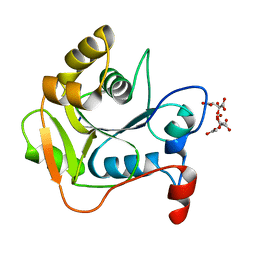 | | CRYSTAL STRUCTURE OF THE MOLYBDENUM COFACTOR BIOSYNTHESIS PROTEIN MOBA (PROTEIN FA) FROM ESCHERICHIA COLI AT NEAR ATOMIC RESOLUTION | | Descriptor: | CITRIC ACID, LITHIUM ION, MOLYBDOPTERIN-GUANINE DINUCLEOTIDE BIOSYNTHESIS PROTEIN A | | Authors: | Stevenson, C.E.M, Sargent, F, Buchanan, G, Palmer, T, Lawson, D.M. | | Deposit date: | 2000-07-27 | | Release date: | 2000-11-07 | | Last modified: | 2019-05-08 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Crystal Structure of the Molybdenum Cofactor Biosynthesis Protein Moba from Escherichia Coli at Near Atomic Resolution
Structure, 8, 2000
|
|
2JSX
 
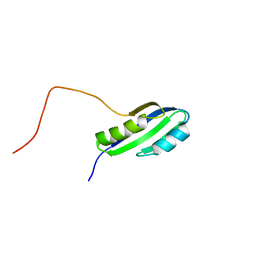 | |
2UXT
 
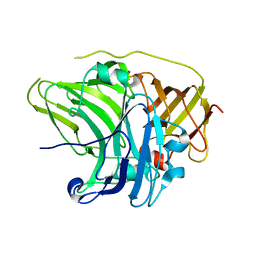 | | SufI Protein from Escherichia Coli | | Descriptor: | PROTEIN SUFI | | Authors: | Tarry, M.J, Roversi, P, Sargent, F, Berks, B.C, Lea, S.M. | | Deposit date: | 2007-03-29 | | Release date: | 2008-05-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Escherichia Coli Cell Division Protein and Model Tat Substrate Sufi (Ftsp) Localizes to the Septal Ring and Has a Multicopper Oxidase-Like Structure.
J.Mol.Biol., 386, 2009
|
|
2UXV
 
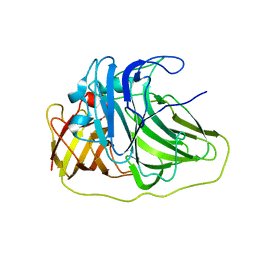 | | SufI Protein from Escherichia Coli | | Descriptor: | PROTEIN SUFI | | Authors: | Tarry, M.J, Roversi, P, Sargent, F, Berks, B.C, Lea, S.M. | | Deposit date: | 2007-03-30 | | Release date: | 2008-05-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | The Escherichia Coli Cell Division Protein and Model Tat Substrate Sufi (Ftsp) Localizes to the Septal Ring and Has a Multicopper Oxidase-Like Structure.
J.Mol.Biol., 386, 2009
|
|
5A4I
 | | The mechanism of Hydrogen activation by NiFE-hydrogenases | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, CHLORIDE ION, DODECYL-BETA-D-MALTOSIDE, ... | | Authors: | Evans, R.M, Brooke, E.J, Wehlin, S.A.M, Nomerotskaia, E, Sargent, F, Carr, S.C, Phillips, S.E.V, Armstrong, F.A. | | Deposit date: | 2015-06-10 | | Release date: | 2015-11-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.23 Å) | | Cite: | Mechanism of hydrogen activation by [NiFe] hydrogenases.
Nat. Chem. Biol., 12, 2016
|
|
5A4F
 
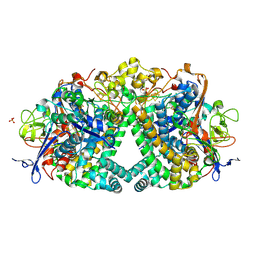 | | The mechanism of Hydrogen Activation by NiFe-hydrogenases. | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, CHLORIDE ION, FE3-S4 CLUSTER, ... | | Authors: | Evans, R.M, Brooke, E.J, Wehlin, S.A.M, Nomerotskaia, E, Sargent, F, Carr, S.B, Phillips, S.E.V, Armstrong, F.A. | | Deposit date: | 2015-06-09 | | Release date: | 2015-11-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Mechanism of hydrogen activation by [NiFe] hydrogenases.
Nat. Chem. Biol., 12, 2016
|
|
5ADU
 | | The Mechanism of Hydrogen Activation by NiFe-hydrogenases | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, CHLORIDE ION, DODECYL-BETA-D-MALTOSIDE, ... | | Authors: | Evans, R, Brooke, E.J, Wehlin, S.A, Nomerotskaia, E, Sargent, F, Carr, S.B, Phillips, S.E.V, Armstrong, F.A. | | Deposit date: | 2015-08-24 | | Release date: | 2015-11-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Mechanism of hydrogen activation by [NiFe] hydrogenases.
Nat. Chem. Biol., 12, 2016
|
|
2XOL
 
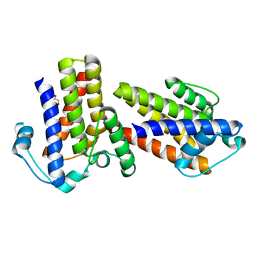 | | High resolution structure of TtrD from Archaeoglobus fulgidus | | Descriptor: | 1,2-ETHANEDIOL, CHAPERONE PROTEIN TTRD | | Authors: | Dawson, A, Coulthurst, S.J, Sargent, F, Hunter, W.N. | | Deposit date: | 2010-08-18 | | Release date: | 2011-09-07 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Conserved Signal Peptide Recognition Systems Across the Prokaryotic Domains.
Biochemistry, 51, 2012
|
|
2YJM
 | | Structure of TtrD from Archaeoglobus fulgidus | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, TTRD | | Authors: | Dawson, A, Coulthurst, S.J, Hunter, W.N, Sargent, F. | | Deposit date: | 2011-05-20 | | Release date: | 2012-02-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Conserved Signal Peptide Recognition Systems Across the Prokaryotic Domains.
Biochemistry, 51, 2012
|
|
2Y6Y
 
 | | Crystal structure of TtrD from Archaeoglobus fulgidus | | Descriptor: | CHAPERONE PROTEIN TTRD, CHLORIDE ION | | Authors: | Dawson, A, Coulthurst, S.J, Sargent, F, Hunter, W.N. | | Deposit date: | 2011-01-27 | | Release date: | 2012-02-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Conserved Signal Peptide Recognition Systems Across the Prokaryotic Domains.
Biochemistry, 51, 2012
|
|
5OPZ
 
 | | Crystal structure of Serratia marcescens L-Ala D-Glu endopeptidase ChiX | | Descriptor: | CHLORIDE ION, ChiX, ZINC ION | | Authors: | Owen, R.A, Fyfe, P.K, Lodge, A, Biboy, J, Vollmer, W, Hunter, W.N, Sargent, F. | | Deposit date: | 2017-08-10 | | Release date: | 2018-01-17 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Structure and activity of ChiX: a peptidoglycan hydrolase required for chitinase secretion by Serratia marcescens.
Biochem. J., 475, 2018
|
|
5OQ1
 
 | | Crystal structure of Serratia marcescens ChiX (used as MR model for superior PDB 5OPZ) | | Descriptor: | CHLORIDE ION, ChiX, ZINC ION | | Authors: | Owen, R.A, Fyfe, P.K, Lodge, A, Biboy, J, Vollmer, W, Hunter, W.N, Sargent, F. | | Deposit date: | 2017-08-10 | | Release date: | 2018-01-17 | | Last modified: | 2018-02-07 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Structure and activity of ChiX: a peptidoglycan hydrolase required for chitinase secretion by Serratia marcescens.
Biochem. J., 475, 2018
|
|
4GD3
 
 | | Structure of E. coli hydrogenase-1 in complex with cytochrome b | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, CHLORIDE ION, DODECYL-BETA-D-MALTOSIDE, ... | | Authors: | Volbeda, A, Fontecilla-Camps, J.C, Darnault, C. | | Deposit date: | 2012-07-31 | | Release date: | 2013-01-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Crystal Structure of the O(2)-Tolerant Membrane-Bound Hydrogenase 1 from Escherichia coli in Complex with Its Cognate Cytochrome b.
Structure, 21, 2013
|
|
5A4M
 
 | | Mechanism of Hydrogen activation by NiFe-hydrogenases | | Descriptor: | CHLORIDE ION, FE3-S4 CLUSTER, FE4-S3 CLUSTER, ... | | Authors: | Evans, R.M, Brooke, E.J, Wehlin, S.A.M, Nomerotskaia, E, Sergent, F, Carr, S.B, Philips, S.E.V, Armstrong, F.A. | | Deposit date: | 2015-06-10 | | Release date: | 2015-11-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Mechanism of Hydrogen Activation by [Nife] Hydrogenases.
Nat.Chem.Biol., 12, 2016
|
|
6EHS
 
 | | E. coli Hydrogenase-2 chemically reduced structure | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, DITHIONITE, FE3-S4 CLUSTER, ... | | Authors: | Carr, S.B, Evans, R.M, Beaton, S.E, Armstrong, F.A. | | Deposit date: | 2017-09-15 | | Release date: | 2018-04-18 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The structure of hydrogenase-2 fromEscherichia coli: implications for H2-driven proton pumping.
Biochem. J., 475, 2018
|
|
6EN9
 | | E. coli Hydrogenase-2 (hydrogen reduced form) | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, Hydrogenase-2 large chain, ... | | Authors: | Carr, S.B, Beaton, S.E, Evans, R.M, Armstrong, F.A. | | Deposit date: | 2017-10-04 | | Release date: | 2018-04-18 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The structure of hydrogenase-2 fromEscherichia coli: implications for H2-driven proton pumping.
Biochem. J., 475, 2018
|
|
6EHQ
 | | E. coli Hydrogenase-2 (as isolated form). | | Descriptor: | CARBONMONOXIDE-(DICYANO) IRON, FE3-S4 CLUSTER, Hydrogenase-2 large chain, ... | | Authors: | Carr, S.B, Beaton, S.E, Evans, R.M, Armstrong, F.A. | | Deposit date: | 2017-09-14 | | Release date: | 2018-04-18 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The structure of hydrogenase-2 fromEscherichia coli: implications for H2-driven proton pumping.
Biochem. J., 475, 2018
|
|
6G94
 | |
