2Y8C
 
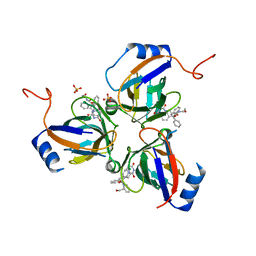 | | Plasmodium falciparum dUTPase in complex with a trityl ligand | | Descriptor: | (2S)-2-[(2,4-dioxopyrimidin-1-yl)methyl]-N-(2-hydroxyethyl)-4-trityloxy-butanamide, DEOXYURIDINE 5'-TRIPHOSPHATE NUCLEOTIDOHYDROLASE, SULFATE ION | | Authors: | Baragana, B, McCarthy, O, Sanchez, P, Bosch, C, Kaiser, M, Brun, R, Whittingham, J.L, Roberts, S, Zhou, X.-X, Wilson, K.S, Johansson, N.G, Gonzalez-Pacanowska, D, Gilbert, I.H. | | Deposit date: | 2011-02-04 | | Release date: | 2012-03-07 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Beta-Branched Acyclic Nucleoside Analogues as Inhibitors of Plasmodium Falciparum Dutpase
Bioorg.Med.Chem., 19, 2011
|
|
8SPK
 
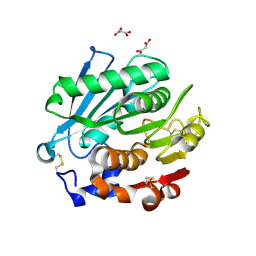 | | Crystal structure of Antarctic PET-degrading enzyme | | Descriptor: | Lipase 1, MALONATE ION | | Authors: | Furtado, A.A, Blazquez-Sanchez, P, Grinen, A, Vargas, J.A, Leonardo, D.A, Sculaccio, S.A, Pereira, H.M, Diez, B, Garratt, R.C, Ramirez-Sarmiento, C.A. | | Deposit date: | 2023-05-03 | | Release date: | 2023-08-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Engineering the catalytic activity of an Antarctic PET-degrading enzyme by loop exchange.
Protein Sci., 32, 2023
|
|
8F4X
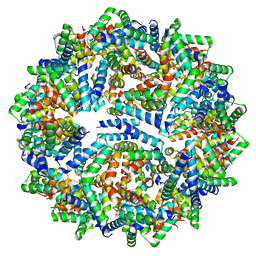 
 | |
8F53
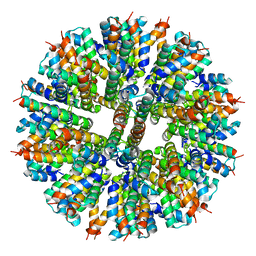 
 | |
8F54
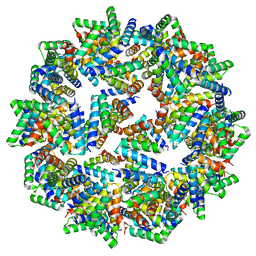 
 | |
7NEI
 
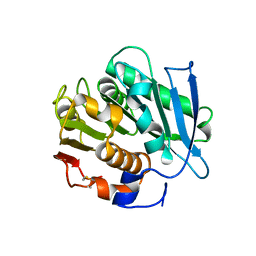 | |
4MPI
 
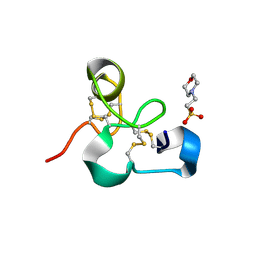 | | Crystal structure of the chitin-binding module (CBM18) of a chitinase-like protein from Hevea brasiliensis | | Descriptor: | 1,4-DIETHYLENE DIOXIDE, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Class I chitinase | | Authors: | Martinez-Caballero, C.S, Hermoso, J.A, Rodriguez-Romero, A. | | Deposit date: | 2013-09-12 | | Release date: | 2014-08-27 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.602 Å) | | Cite: | Comparative study of two GH19 chitinase-like proteins from Hevea brasiliensis, one exhibiting a novel carbohydrate-binding domain.
Febs J., 281, 2014
|
|
4MST
 
 | |
8BRB
 
 | |
8BRA
 
 | |
5VBK
 
 | |
2KY3
 
 | |
7M1E
 
 | |
7M1D
 
 | |
6AX2
 
 | |
6BB6
 
 | |
6BAM
 
 | |
6B9W
 
 | |
5IPO
 
 | |
5JYH
 
 | |
6OGX
 
 | | Ternary complex of OX40R (TNFRSF4) bound to Fab1 and Fab2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Fab 1 Heavy Chain, Fab1 Light Chain, ... | | Authors: | Ultsch, M.H, Boenig, G, Harris, S.F. | | Deposit date: | 2019-04-03 | | Release date: | 2019-07-10 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.77 Å) | | Cite: | Tetravalent biepitopic targeting enables intrinsic antibody agonism of tumor necrosis factor receptor superfamily members.
Mabs, 11, 2019
|
|
6OKM
 
 | | Human OX40R (TNFRSF4) bound to Fab 3C8 | | Descriptor: | Fab 3C8 Heavy Chain, Fab 3C8 light chain, Tumor necrosis factor receptor superfamily member 4 | | Authors: | Boenig, G, Ultsch, M.H, Harris, S.F. | | Deposit date: | 2019-04-14 | | Release date: | 2019-08-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Tetravalent biepitopic targeting enables intrinsic antibody agonism of tumor necrosis factor receptor superfamily members.
Mabs, 11, 2019
|
|
6OKN
 
 | | OX40R (TNFRSF4) bound to Fab 1A7 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Fab 1A7 heavy chain, Fab 1A7 light chain, ... | | Authors: | Ultsch, M.H, Boenig, G, Harris, S.F. | | Deposit date: | 2019-04-14 | | Release date: | 2019-07-10 | | Last modified: | 2019-08-14 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Tetravalent biepitopic targeting enables intrinsic antibody agonism of tumor necrosis factor receptor superfamily members.
Mabs, 11, 2019
|
|
7PTJ
 
 | |
7PTH
 
 | |
