2W7Q
 
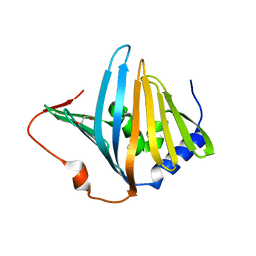 | | Structure of Pseudomonas aeruginosa LolA | | Descriptor: | GLYCEROL, OUTER-MEMBRANE LIPOPROTEIN CARRIER PROTEIN | | Authors: | Remans, K, Pauwels, K, van Ulsen, P, Savvides, S, Decanniere, K, Cornelis, P, Tommassen, J, Van Gelder, P. | | Deposit date: | 2008-12-23 | | Release date: | 2009-12-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Hydrophobic Surface Patches on Lola of Pseudomonas Aeruginosa are Essential for Lipoprotein Binding.
J.Mol.Biol., 401, 2010
|
|
4QAM
 
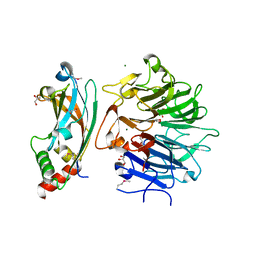 | | Crystal Structure of the RPGR RCC1-like domain in complex with the RPGR-interacting domain of RPGRIP1 | | Descriptor: | GLYCEROL, MAGNESIUM ION, X-linked retinitis pigmentosa GTPase regulator, ... | | Authors: | Remans, K, Buerger, M, Vetter, I.R, Wittinghofer, A. | | Deposit date: | 2014-05-05 | | Release date: | 2014-07-30 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | C2 domains as protein-protein interaction modules in the ciliary transition zone.
Cell Rep, 8, 2014
|
|
7A25
 
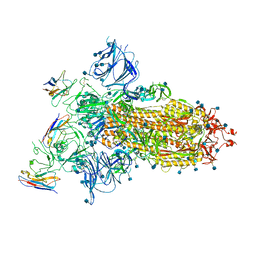 | |
7A29
 
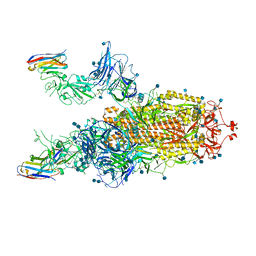 | |
7ZV5
 
 | |
7ZV8
 
 | | Crystal structure of SARS Cov-2 main protease in complex with an inhibitor 58 | | Descriptor: | 3C-like proteinase nsp5, DIMETHYL SULFOXIDE, OCTANOIC ACID (CAPRYLIC ACID), ... | | Authors: | Rahimova, R, Di Micco, S, Marquez, J.A. | | Deposit date: | 2022-05-13 | | Release date: | 2022-12-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.937 Å) | | Cite: | Rational design of the zonulin inhibitor AT1001 derivatives as potential anti SARS-CoV-2.
Eur.J.Med.Chem., 244, 2022
|
|
7ZV7
 
 | |
