6FXP
 
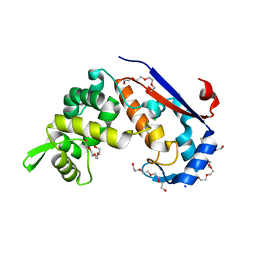 | | Crystal structure of S. aureus glucosaminidase B | | Descriptor: | CHLORIDE ION, SODIUM ION, TETRAETHYLENE GLYCOL, ... | | Authors: | Pintar, S, Turk, D. | | Deposit date: | 2018-03-09 | | Release date: | 2019-03-20 | | Last modified: | 2020-09-30 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Domain sliding of two Staphylococcus aureus N-acetylglucosaminidases enables their substrate-binding prior to its catalysis.
Commun Biol, 3, 2020
|
|
6FXO
 
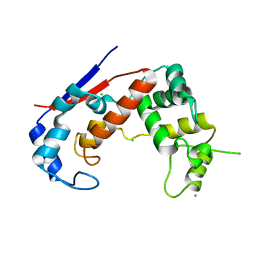 | | Crystal structure of Major Bifunctional Autolysin | | Descriptor: | Bifunctional autolysin, CHLORIDE ION | | Authors: | Pintar, S, Turk, D. | | Deposit date: | 2018-03-09 | | Release date: | 2019-03-20 | | Last modified: | 2020-09-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Domain sliding of two Staphylococcus aureus N-acetylglucosaminidases enables their substrate-binding prior to its catalysis.
Commun Biol, 3, 2020
|
|
8BV3
 
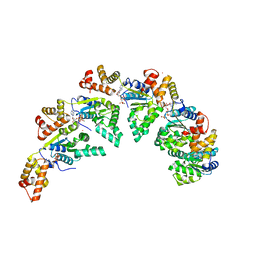 | | Bacillus subtilis DnaA domain III structure | | Descriptor: | Chromosomal replication initiator protein DnaA, MAGNESIUM ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Pintar, S, Hubbard, J.A. | | Deposit date: | 2022-12-01 | | Release date: | 2023-12-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | The bacterial replication origin BUS promotes nucleobase capture.
Nat Commun, 14, 2023
|
|
8BTG
 
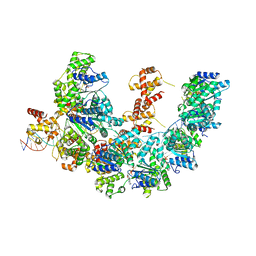 | | Cryo-EM structure of the bacterial replication origin opening basal unwinding system | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Chromosomal replication initiator protein DnaA, DNA (41-MER), ... | | Authors: | Pelliciari, S, Bodet-Lefevre, S, Murray, H, Ilangovan, A. | | Deposit date: | 2022-11-28 | | Release date: | 2024-02-14 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The bacterial replication origin BUS promotes nucleobase capture.
Nat Commun, 14, 2023
|
|
