4GBM
 
 | |
4GOX
 
 | |
2EFJ
 
 | |
2EG5
 
 | | The structure of xanthosine methyltransferase | | Descriptor: | 9-[(2R,3R,4S,5R)-3,4-DIHYDROXY-5-(HYDROXYMETHYL)OXOLAN-2-YL]-3H-PURINE-2,6-DIONE, S-ADENOSYL-L-HOMOCYSTEINE, Xanthosine methyltransferase | | Authors: | McCarthy, A.A, McCarthy, J.G. | | Deposit date: | 2007-02-28 | | Release date: | 2007-05-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The structure of two N-methyltransferases from the caffeine biosynthetic pathway
Plant Physiol., 144, 2007
|
|
4G22
 
 | |
4G2M
 
 | |
4G0B
 
 | |
5IV8
 
 | | The LPS Transporter LptDE from Klebsiella pneumoniae, core complex | | Descriptor: | (HYDROXYETHYLOXY)TRI(ETHYLOXY)OCTANE, LPS biosynthesis protein, LPS-assembly lipoprotein LptE | | Authors: | Botos, I, McCarthy, J.G, Buchanan, S.K. | | Deposit date: | 2016-03-20 | | Release date: | 2016-05-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.938 Å) | | Cite: | Structural and Functional Characterization of the LPS Transporter LptDE from Gram-Negative Pathogens.
Structure, 24, 2016
|
|
5IV9
 
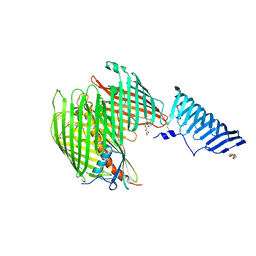 | | The LPS Transporter LptDE from Klebsiella pneumoniae, full-length | | Descriptor: | (HYDROXYETHYLOXY)TRI(ETHYLOXY)OCTANE, LPS-assembly lipoprotein LptE, LPS-assembly protein LptD | | Authors: | Botos, I, McCarthy, J.G, Buchanan, S.K. | | Deposit date: | 2016-03-20 | | Release date: | 2016-05-18 | | Last modified: | 2016-06-15 | | Method: | X-RAY DIFFRACTION (4.369 Å) | | Cite: | Structural and Functional Characterization of the LPS Transporter LptDE from Gram-Negative Pathogens.
Structure, 24, 2016
|
|
5IXM
 
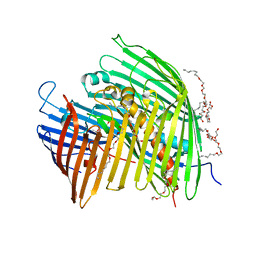 | | The LPS Transporter LptDE from Yersinia pestis, core complex | | Descriptor: | (HYDROXYETHYLOXY)TRI(ETHYLOXY)OCTANE, LPS-assembly lipoprotein LptE, LPS-assembly protein LptD | | Authors: | Botos, I, Mayclin, S.J, McCarthy, J.G, Buchanan, S.K. | | Deposit date: | 2016-03-23 | | Release date: | 2016-05-18 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.746 Å) | | Cite: | Structural and Functional Characterization of the LPS Transporter LptDE from Gram-Negative Pathogens.
Structure, 24, 2016
|
|
5IVA
 
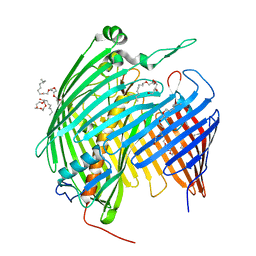 | |
