1F9K
 
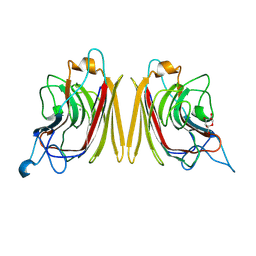 | | WINGED BEAN ACIDIC LECTIN COMPLEXED WITH METHYL-ALPHA-D-GALACTOSE | | Descriptor: | ACIDIC LECTIN, CALCIUM ION, MANGANESE (II) ION, ... | | Authors: | Manoj, N, Srinivas, V.R, Surolia, A, Vijayan, M, Suguna, K. | | Deposit date: | 2000-07-11 | | Release date: | 2001-07-11 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Carbohydrate specificity and salt-bridge mediated conformational change in acidic winged bean agglutinin.
J.Mol.Biol., 302, 2000
|
|
1FAY
 
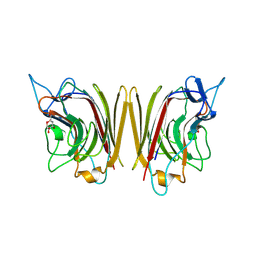 | | WINGED BEAN ACIDIC LECTIN COMPLEXED WITH METHYL-ALPHA-D-GALACTOSE (MONOCLINIC FORM) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ACIDIC LECTIN, CALCIUM ION, ... | | Authors: | Manoj, N, Srinivas, V.R, Surolia, A, Vijayan, M, Suguna, K. | | Deposit date: | 2000-07-14 | | Release date: | 2001-07-14 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Carbohydrate specificity and salt-bridge mediated conformational change in acidic winged bean agglutinin.
J.Mol.Biol., 302, 2000
|
|
1WBF
 
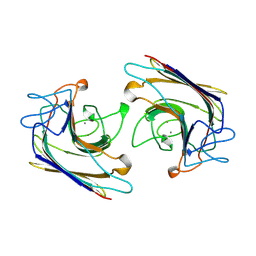 | | WINGED BEAN LECTIN, SACCHARIDE FREE FORM | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, MANGANESE (II) ION, ... | | Authors: | Manoj, N, Srinivas, V.R, Suguna, K. | | Deposit date: | 1998-12-16 | | Release date: | 1999-12-22 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of basic winged-bean lectin and a comparison with its saccharide-bound form.
Acta Crystallogr.,Sect.D, 55, 1999
|
|
1P9O
 
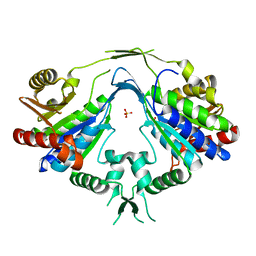 | | Crystal Structure of Phosphopantothenoylcysteine Synthetase | | Descriptor: | Phosphopantothenoylcysteine synthetase, SULFATE ION | | Authors: | Manoj, N, Strauss, E, Begley, T.P, Ealick, S.E. | | Deposit date: | 2003-05-12 | | Release date: | 2003-09-02 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of human phosphopantothenoylcysteine synthetase at 2.3 A resolution.
Structure, 11, 2003
|
|
1QZU
 
 | |
7BRF
 
 | |
7EFZ
 
 | |
5JIB
 
 | |
5GMA
 | |
5HFN
 | |
5ZCI
 
 | |
5ZCM
 
 | |
5FDF
 | |
6LDR
 
 | |
6LDS
 
 | |
6LDT
 
 | |
6M4Y
 
 | |
7BR4
 
 | |
7CTD
 
 | |
7CTL
 
 | |
7CTM
 
 | |
6KCX
 
 | |
6JY1
 
 | |
7EC9
 
 | |
2D3S
 
 | | Crystal Structure of basic winged bean lectin with Tn-antigen | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-galactopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, Basic agglutinin, ... | | Authors: | Kulkarni, K.A, Sinha, S, Katiyar, S, Surolia, A, Vijayan, M, Suguna, K. | | Deposit date: | 2005-10-01 | | Release date: | 2006-01-17 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural basis for the specificity of basic winged bean lectin for the Tn-antigen: a crystallographic, thermodynamic and modelling study
Febs Lett., 579, 2005
|
|
