2KJ6
 | | NMR Solution Structure of a Tubulin folding cofactor B obtained from Arabidopsis thaliana: Northeast Structural Genomics Consortium target AR3436A | | Descriptor: | Tubulin folding cofactor B | | Authors: | Mani, R, Swapna, G.V.T, Shastry, R, Foote, E, Ciccosanti, C, Jiang, M, Xiao, R, Nair, R, Everett, J, Huang, Y.J, Acton, T, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-05-22 | | Release date: | 2009-07-21 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | NMR Solution Structure of Tbulin folding Cofactor B obtained from Arabidopsis thaliana: Northeast
To be Published
|
|
2KK8
 
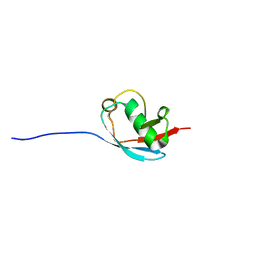 | | NMR Solution Structure of a Putative Uncharacterized Protein obtained from Arabidopsis thaliana: Northeast Structural Genomics Consortium target AR3449A | | Descriptor: | Uncharacterized protein AT4g05270 | | Authors: | Mani, R, Gurla, S.V.T, Shastry, R, Ciccosanti, C, Foote, E, Jiang, M, Xiao, R, Nair, R, Everett, J, Huang, Y, Acton, T, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-16 | | Release date: | 2009-06-30 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | NMR Solution Structure of a Putative Uncharacterized Protein obtained from Arabidopsis thaliana: Northeast Structural Genomics Consortium Target AR3449A
To be Published
|
|
2K5P
 
 | | NMR Solution Structure of a Thiamine Biosynthesis Protein from Geobacter Metallireducens: Northeast Structural Genomics Consortium Target GmR137 | | Descriptor: | thiamine-biosynthesis protein | | Authors: | Mani, R, Wang, H, Jiang, M, Magliaqui, M, Xiao, R, Nair, R, Baran, M.C, Gurla, S.V.T, Acton, T.B, Rost, B, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-30 | | Release date: | 2008-08-26 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | NMR Solution Structure of a Thiamine Biosynthesis Protein from Geobacter Metallireducens: Northeast Structural Genomics Consortium Target GmR137
To be Published
|
|
2L08
 | | Solution NMR Structure of Nonsense mRNA reducing factor 3A from H. Sapiens, Northeast Structural Genomics Consortium Target HR4714B | | Descriptor: | Regulator of nonsense transcripts 3A | | Authors: | Mani, R, Mao, L, Ciccosanti, C, Shastry, R, Acton, T.B, Xiao, R, Swapna, G.V.T, Everett, J.K, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-06-30 | | Release date: | 2010-08-11 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of Nonsense mRNA reducing factor 3A from H. Sapiens, Northeast Structural Genomics Consortium Target HR4714B
To be Published
|
|
2KRT
 | | Solution NMR Structure of a Conserved Hypothetical Membrane Lipoprotein Obtained from Ureaplasma parvum: Northeast Structural Genomics Consortium Target UuR17A (139-239) | | Descriptor: | Conserved hypothetical membrane lipoprotein | | Authors: | Mani, R, Swapna, G, Janjua, H, Ciccosanti, C, Huang, Y, Patel, D, Xiao, R, Acton, T, Everett, J, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-12-22 | | Release date: | 2010-01-05 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | NMR Solution Structure of a Conserved Hypothetical Membrane Lipoprotein obtained from Ureaplasma parvum : Northeast Structural Genomics Consortium Target UuR17A (139-239)
To be Published
|
|
1ZY6
 
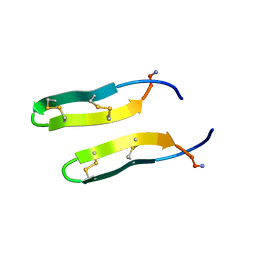 | | Membrane-bound dimer structure of Protegrin-1 (PG-1), a beta-Hairpin Antimicrobial Peptide in Lipid Bilayers from Rotational-Echo Double-Resonance Solid-State NMR | | Descriptor: | Protegrin 1 | | Authors: | Wu, X, Mani, R, Tang, M, Buffy, J.J, Waring, A.J, Sherman, M.A, Hong, M. | | Deposit date: | 2005-06-09 | | Release date: | 2006-06-13 | | Last modified: | 2022-03-02 | | Method: | SOLID-STATE NMR | | Cite: | Membrane-Bound Dimer Structure of a beta-Hairpin Antimicrobial Peptide from Rotational-Echo Double-Resonance Solid-State NMR.
Biochemistry, 45, 2006
|
|
6QTJ
 
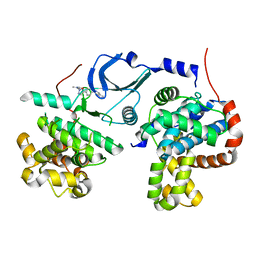 | | Crystal structure of human CDK8/CYCC in complex with BI 919811 | | Descriptor: | Cyclin-C, Cyclin-dependent kinase 8, ~{N},~{N}-dimethyl-2-[4-[4-(2,6-naphthyridin-4-yl)phenyl]pyrazol-1-yl]ethanamide | | Authors: | Boettcher, J. | | Deposit date: | 2019-02-25 | | Release date: | 2020-03-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Selective and Potent CDK8/19 Inhibitors Enhance NK-Cell Activity and Promote Tumor Surveillance.
Mol.Cancer Ther., 19, 2020
|
|
6QTG
 
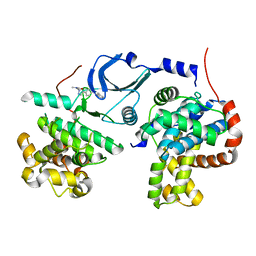 | | Crystal structure of human CDK8/CYCC in complex with BI-1347 | | Descriptor: | 2-[4-(4-isoquinolin-4-ylphenyl)pyrazol-1-yl]-~{N},~{N}-dimethyl-ethanamide, Cyclin-C, Cyclin-dependent kinase 8 | | Authors: | Boettcher, J. | | Deposit date: | 2019-02-25 | | Release date: | 2020-03-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Selective and Potent CDK8/19 Inhibitors Enhance NK-Cell Activity and Promote Tumor Surveillance.
Mol.Cancer Ther., 19, 2020
|
|
6R3S
 
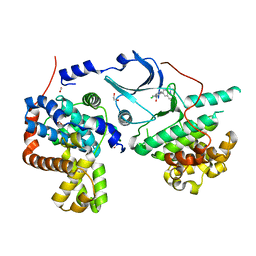 | | CRYSTAL STRUCTURE OF CDK8-CycC IN COMPLEX WITH COMPOUND 1 | | Descriptor: | 1,2-ETHANEDIOL, 6-[5-chloranyl-4-[(1~{S})-1-oxidanylethyl]pyridin-3-yl]-3,4-dihydro-2~{H}-1,8-naphthyridine-1-carboxamide, Cyclin-C, ... | | Authors: | Boettcher, J. | | Deposit date: | 2019-03-21 | | Release date: | 2020-04-08 | | Last modified: | 2020-04-22 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Selective and Potent CDK8/19 Inhibitors Enhance NK-Cell Activity and Promote Tumor Surveillance.
Mol.Cancer Ther., 19, 2020
|
|
4GMX
 
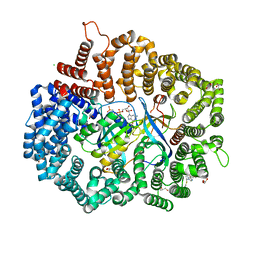 | |
2KAD
 
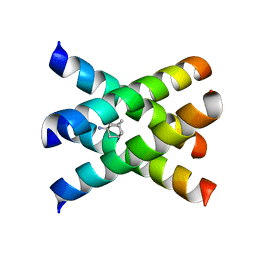 | | Magic-Angle-Spinning Solid-State NMR Structure of Influenza A M2 Transmembrane Domain | | Descriptor: | (3S,5S,7S)-tricyclo[3.3.1.1~3,7~]decan-1-amine, Transmembrane peptide of Matrix protein 2 | | Authors: | Hong, M, Cady, S.D, Mishanina, T.V. | | Deposit date: | 2008-11-04 | | Release date: | 2008-11-18 | | Last modified: | 2021-10-20 | | Method: | SOLID-STATE NMR | | Cite: | Structure of amantadine-bound M2 transmembrane peptide of influenza A in lipid bilayers from magic-angle-spinning solid-state NMR: the role of Ser31 in amantadine binding.
J.Mol.Biol., 385, 2009
|
|
