1D8D
 
 | | CO-CRYSTAL STRUCTURE OF RAT PROTEIN FARNESYLTRANSFERASE COMPLEXED WITH A K-RAS4B PEPTIDE SUBSTRATE AND FPP ANALOG AT 2.0A RESOLUTION | | Descriptor: | ACETATE ION, K-RAS4B PEPTIDE SUBSTRATE, ZINC ION, ... | | Authors: | Long, S.B, Casey, P.J, Beese, L.S. | | Deposit date: | 1999-10-22 | | Release date: | 2000-02-09 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The basis for K-Ras4B binding specificity to protein farnesyltransferase revealed by 2 A resolution ternary complex structures.
Structure Fold.Des., 8, 2000
|
|
1D8E
 
 | | Zinc-depleted FTase complexed with K-RAS4B peptide substrate and FPP analog. | | Descriptor: | ACETATE ION, FARNESYLTRANSFERASE (ALPHA SUBUNIT), FARNESYLTRANSFERASE (BETA SUBUNIT), ... | | Authors: | Long, S.B, Casey, P.J, Beese, L.S. | | Deposit date: | 1999-10-22 | | Release date: | 2000-03-20 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The basis for K-Ras4B binding specificity to protein farnesyltransferase revealed by 2 A resolution ternary complex structures.
Structure Fold.Des., 8, 2000
|
|
3DD2
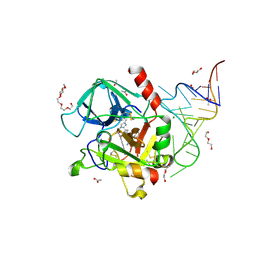 
 | | Crystal structure of an RNA aptamer bound to human thrombin | | Descriptor: | ACETIC ACID, D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Long, S.B, Sullenger, B.A. | | Deposit date: | 2008-06-04 | | Release date: | 2008-11-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of an RNA aptamer bound to thrombin.
Rna, 14, 2008
|
|
1KZO
 
 | | PROTEIN FARNESYLTRANSFERASE COMPLEXED WITH FARNESYLATED K-RAS4B PEPTIDE PRODUCT AND FARNESYL DIPHOSPHATE SUBSTRATE BOUND SIMULTANEOUSLY | | Descriptor: | ACETIC ACID, FARNESYL, FARNESYL DIPHOSPHATE, ... | | Authors: | Long, S.B, Casey, P.J, Beese, L.S. | | Deposit date: | 2002-02-07 | | Release date: | 2002-10-16 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Reaction Path of Protein Farnesyltransferase at Atomic Resolution
Nature, 419, 2002
|
|
1KZP
 
 | | PROTEIN FARNESYLTRANSFERASE COMPLEXED WITH A FARNESYLATED K-RAS4B PEPTIDE PRODUCT | | Descriptor: | ACETIC ACID, FARNESYL, Farnesylated K-Ras4B peptide product, ... | | Authors: | Long, S.B, Casey, P.J, Beese, L.S. | | Deposit date: | 2002-02-07 | | Release date: | 2002-10-16 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Reaction path of protein farnesyltransferase at atomic resolution
Nature, 419, 2002
|
|
1JCS
 
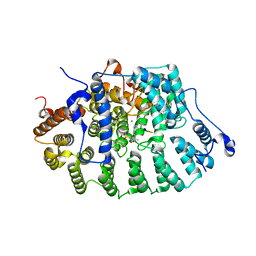 | | CRYSTAL STRUCTURE OF RAT PROTEIN FARNESYLTRANSFERASE COMPLEXED WITH THE PEPTIDE SUBSTRATE TKCVFM AND AN ANALOG OF FARNESYL DIPHOSPHATE | | Descriptor: | ACETIC ACID, PROTEIN FARNESYLTRANSFERASE, ALPHA SUBUNIT, ... | | Authors: | Long, S.B, Casey, P.J, Beese, L.S. | | Deposit date: | 2001-06-11 | | Release date: | 2001-11-02 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The crystal structure of human protein farnesyltransferase reveals the basis for inhibition by CaaX tetrapeptides and their mimetics.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1JCR
 
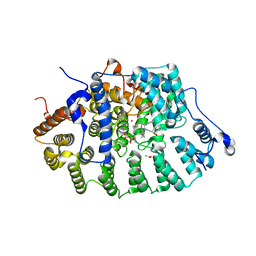 | | CRYSTAL STRUCTURE OF RAT PROTEIN FARNESYLTRANSFERASE COMPLEXED WITH THE NON-SUBSTRATE TETRAPEPTIDE INHIBITOR CVFM AND FARNESYL DIPHOSPHATE SUBSTRATE | | Descriptor: | ACETIC ACID, FARNESYL DIPHOSPHATE, PROTEIN FARNESYLTRANSFERASE, ... | | Authors: | Long, S.B, Casey, P.J, Beese, L.S. | | Deposit date: | 2001-06-11 | | Release date: | 2001-11-02 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of human protein farnesyltransferase reveals the basis for inhibition by CaaX tetrapeptides and their mimetics.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1JCQ
 
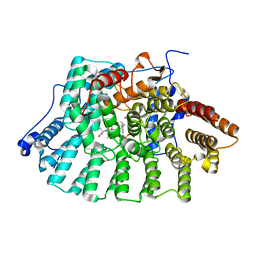 | | CRYSTAL STRUCTURE OF HUMAN PROTEIN FARNESYLTRANSFERASE COMPLEXED WITH FARNESYL DIPHOSPHATE AND THE PEPTIDOMIMETIC INHIBITOR L-739,750 | | Descriptor: | 2(S)-{2(S)-[2(R)-AMINO-3-MERCAPTO]PROPYLAMINO-3(S)-METHYL}PENTYLOXY-3-PHENYLPROPIONYLMETHIONINE SULFONE, ACETIC ACID, FARNESYL DIPHOSPHATE, ... | | Authors: | Long, S.B, Casey, P.J, Beese, L.S. | | Deposit date: | 2001-06-11 | | Release date: | 2001-11-02 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The crystal structure of human protein farnesyltransferase reveals the basis for inhibition by CaaX tetrapeptides and their mimetics.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
2R9R
 
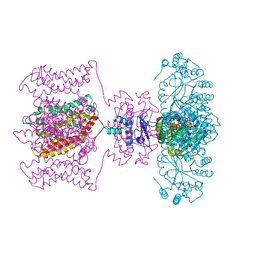 | | Shaker family voltage dependent potassium channel (kv1.2-kv2.1 paddle chimera channel) in association with beta subunit | | Descriptor: | (1R)-2-{[(S)-{[(2S)-2,3-dihydroxypropyl]oxy}(hydroxy)phosphoryl]oxy}-1-[(hexadecanoyloxy)methyl]ethyl (9Z)-octadec-9-enoate, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, POTASSIUM ION, ... | | Authors: | Long, S.B, Tao, X, Campbell, E.B, Mackinnon, R. | | Deposit date: | 2007-09-13 | | Release date: | 2007-11-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Atomic structure of a voltage-dependent K+ channel in a lipid membrane-like environment.
Nature, 450, 2007
|
|
2A79
 
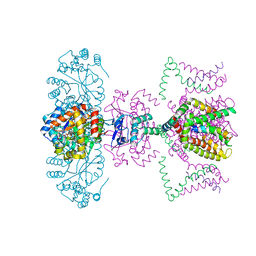 | | Mammalian Shaker Kv1.2 potassium channel- beta subunit complex | | Descriptor: | NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, POTASSIUM ION, Potassium voltage-gated channel subfamily A member 2, ... | | Authors: | Long, S.B, Campbell, E.B, MacKinnon, R. | | Deposit date: | 2005-07-05 | | Release date: | 2005-07-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of a mammalian voltage-dependent Shaker family K+ channel.
Science, 309, 2005
|
|
6XQO
 
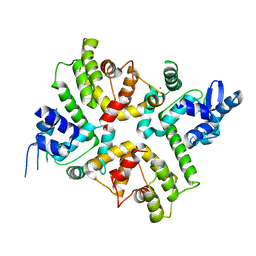 | | Structure of the human MICU1-MICU2 heterodimer, calcium bound, in association with a lipid nanodisc | | Descriptor: | CALCIUM ION, Calcium uptake protein 1, mitochondrial, ... | | Authors: | Long, S.B, Wang, C, Baradaran, R, Jacewicz, A, Delgado, B. | | Deposit date: | 2020-07-09 | | Release date: | 2020-07-29 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Structures reveal gatekeeping of the mitochondrial Ca 2+ uniporter by MICU1-MICU2.
Elife, 9, 2020
|
|
7SQH
 
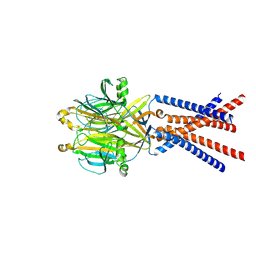 | |
7SQF
 
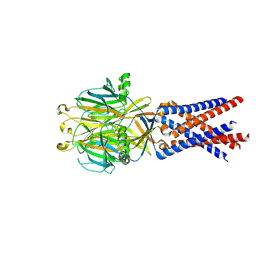 | |
7SQG
 
 | |
7MHY
 
 | |
7MHZ
 
 | |
1FT2
 
 | |
5T5N
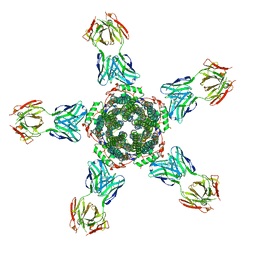 
 | | Calcium-activated chloride channel bestrophin-1 (BEST1), triple mutant: I76A, F80A, F84A; in complex with an Fab antibody fragment, chloride, and calcium | | Descriptor: | CALCIUM ION, CHLORIDE ION, Fab antibody fragment, ... | | Authors: | Long, S.B, Vaisey, G. | | Deposit date: | 2016-08-31 | | Release date: | 2016-11-30 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Distinct regions that control ion selectivity and calcium-dependent activation in the bestrophin ion channel.
Proc. Natl. Acad. Sci. U.S.A., 113, 2016
|
|
7KR5
 
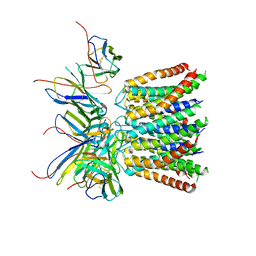 | | Cryo-EM structure of the CRAC channel Orai in an open conformation; H206A gain-of-function mutation in complex with an antibody | | Descriptor: | 19B5 Fab heavy chain, 19B5 Fab light chain, CALCIUM ION, ... | | Authors: | Long, S.B, Hou, X, Outhwaite, I.R. | | Deposit date: | 2020-11-18 | | Release date: | 2020-12-09 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Cryo-EM structure of the calcium release-activated calcium channel Orai in an open conformation.
Elife, 9, 2020
|
|
6X4S
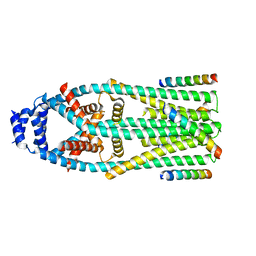 
 | | MCU-EMRE complex of a metazoan mitochondrial calcium uniporter | | Descriptor: | CALCIUM ION, Calcium uniporter protein,Protein EMRE homolog, mitochondrial-like Protein fusion | | Authors: | Long, S.B, Wang, C, Baradaran, R. | | Deposit date: | 2020-05-22 | | Release date: | 2020-09-02 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structure and Reconstitution of an MCU-EMRE Mitochondrial Ca 2+ Uniporter Complex.
J.Mol.Biol., 432, 2020
|
|
6XQN
 
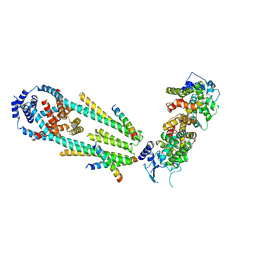 | | Structure of a mitochondrial calcium uniporter holocomplex (MICU1, MICU2, MCU, EMRE) in low Ca2+ | | Descriptor: | CALCIUM ION, Calcium uniporter protein, Calcium uptake protein 1, ... | | Authors: | Long, S.B, Wang, C, Baradaran, R, Jacewicz, A, Delgado, B. | | Deposit date: | 2020-07-09 | | Release date: | 2020-07-29 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Structures reveal gatekeeping of the mitochondrial Ca 2+ uniporter by MICU1-MICU2.
Elife, 9, 2020
|
|
1S63
 
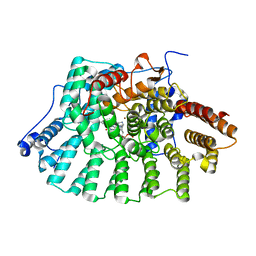 | | Human protein farnesyltransferase complexed with L-778,123 and FPP | | Descriptor: | 4-[(5-{[4-(3-CHLOROPHENYL)-3-OXOPIPERAZIN-1-YL]METHYL}-1H-IMIDAZOL-1-YL)METHYL]BENZONITRILE, FARNESYL DIPHOSPHATE, Protein farnesyltransferase beta subunit, ... | | Authors: | Long, S.B, Casey, P.J, Beese, L.S. | | Deposit date: | 2004-01-22 | | Release date: | 2004-07-27 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystallographic Analysis Reveals that Anticancer Clinical Candidate L-778,123 Inhibits Protein Farnesyltransferase and Geranylgeranyltransferase-I by Different Binding Modes.
Biochemistry, 43, 2004
|
|
4HKR
 
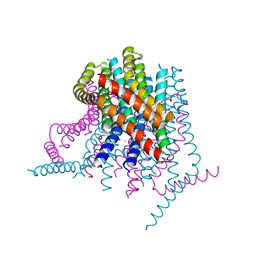 | | Calcium release-activated calcium (CRAC) channel ORAI | | Descriptor: | BARIUM ION, Calcium release-activated calcium channel protein 1, FE (III) ION, ... | | Authors: | Long, S.B, Hou, X. | | Deposit date: | 2012-10-15 | | Release date: | 2012-11-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.3511 Å) | | Cite: | Crystal structure of the calcium release-activated calcium channel Orai.
Science, 338, 2012
|
|
4HKS
 
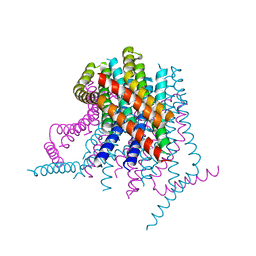 | |
6DNF
 
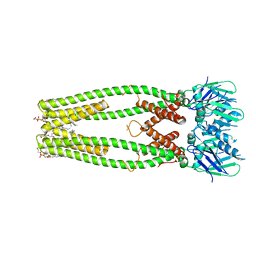 | |
