1H3H
 
 | | Structural Basis for Specific Recognition of an RxxK-containing SLP-76 peptide by the Gads C-terminal SH3 domain | | Descriptor: | GRB2-RELATED ADAPTOR PROTEIN 2, LYMPHOCYTE CYTOSOLIC PROTEIN 2 | | Authors: | Liu, Q, Berry, D, Nash, P, Pawson, T, McGlade, C.J, Li, S.S. | | Deposit date: | 2002-09-03 | | Release date: | 2003-03-06 | | Last modified: | 2013-07-24 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for Specific Binding of the Gads SH3 Domain to an Rxxk Motif-Containing Slp-76 Peptide: A Novel Mode of Peptide Recognition
Mol.Cell, 11, 2003
|
|
1SKA
 
 | | Crystallographic snapshots of Aspergillus fumigatus phytase revealing its enzymatic dynamics | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-phytase A | | Authors: | Liu, Q, Huang, Q, Lei, X.G, Hao, Q. | | Deposit date: | 2004-03-04 | | Release date: | 2004-09-28 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Crystallographic Snapshots of Aspergillus fumigatus Phytase, Revealing Its Enzymatic Dynamics
Structure, 12, 2004
|
|
1SK9
 
 | | Crystallographic snapshots of Aspergillus fumigatus phytase revealing its enzymatic dynamics | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-phytase A, PHOSPHATE ION | | Authors: | Liu, Q, Huang, Q, Lei, X.G, Hao, Q. | | Deposit date: | 2004-03-04 | | Release date: | 2004-09-28 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Crystallographic Snapshots of Aspergillus fumigatus Phytase, Revealing Its Enzymatic Dynamics
Structure, 12, 2004
|
|
1SKB
 
 | | Crystallographic snapshots of Aspergillus fumigatus phytase revealing its enzymatic dynamics | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-phytase A | | Authors: | Liu, Q, Huang, Q, Lei, X.G, Hao, Q. | | Deposit date: | 2004-03-04 | | Release date: | 2004-09-28 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Crystallographic Snapshots of Aspergillus fumigatus Phytase, Revealing Its Enzymatic Dynamics
Structure, 12, 2004
|
|
1MC2
 
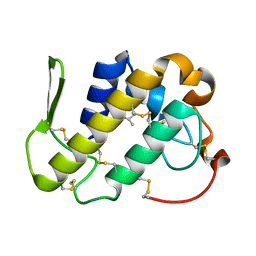 | | monomeric LYS-49 phospholipase A2 homologue purified from AG | | Descriptor: | Acutohaemonlysin, ISOPROPYL ALCOHOL | | Authors: | Liu, Q, Huang, Q.Q, Zhang, R.G, Weeks, C.M, Jelsch, C, Teng, M.K, Niu, L.W. | | Deposit date: | 2002-08-05 | | Release date: | 2002-08-21 | | Last modified: | 2018-02-14 | | Method: | X-RAY DIFFRACTION (0.85 Å) | | Cite: | The crystal structure of a novel, inactive, lysine 49 PLA2 from Agkistrodon acutus venom: an ultrahigh resolution, AB initio structure determination
J.Biol.Chem., 278, 2003
|
|
1YH3
 
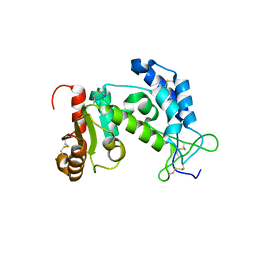 | | Crystal structure of human CD38 extracellular domain | | Descriptor: | ADP-ribosyl cyclase 1 | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Munshi, C, Lee, H.C, Hao, Q. | | Deposit date: | 2005-01-06 | | Release date: | 2005-09-27 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Crystal structure of human CD38 extracellular domain.
Structure, 13, 2005
|
|
1SK8
 
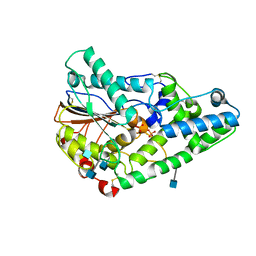 | | Crystallographic snapshots of Aspergillus fumigatus phytase revealing its enzymatic dynamics | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-phytase A, PHOSPHATE ION | | Authors: | Liu, Q, Huang, Q.Q, Lei, X.G, Hao, Q. | | Deposit date: | 2004-03-04 | | Release date: | 2005-02-08 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystallographic Snapshots of Aspergillus fumigatus Phytase, Revealing Its Enzymatic Dynamics
Structure, 12, 2004
|
|
2O3U
 
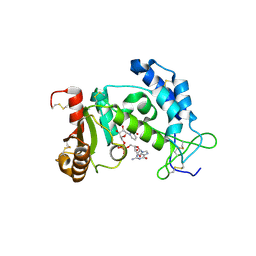 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | 3-(AMINOCARBONYL)-1-[(2R,3R,4S,5R)-5-({[(S)-{[(S)-{[(2R,3S,4R,5R)-5-(2-AMINO-6-OXO-1,6-DIHYDRO-9H-PURIN-9-YL)-3,4-DIHYD ROXYTETRAHYDROFURAN-2-YL]METHOXY}(HYDROXY)PHOSPHORYL]OXY}(HYDROXY)PHOSPHORYL]OXY}METHYL)-3,4-DIHYDROXYTETRAHYDROFURAN-2- YL]PYRIDINIUM, ADP-ribosyl cyclase 1 | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3S
 
 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | ADP-ribosyl cyclase 1, CYCLIC ADENOSINE DIPHOSPHATE-RIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3T
 
 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | ADP-ribosyl cyclase 1, CYCLIC GUANOSINE DIPHOSPHATE-RIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3Q
 
 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | ADP-ribosyl cyclase 1, CYCLIC ADENOSINE DIPHOSPHATE-RIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3R
 
 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | ADP-ribosyl cyclase 1, CYCLIC ADENOSINE DIPHOSPHATE-RIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2PGL
 
 | | Catalysis associated conformational changes revealed by human CD38 complexed with a non-hydrolyzable substrate analog | | Descriptor: | ADP-ribosyl cyclase 1, N1-CYCLIC INOSINE 5'-DIPHOSPHORIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Moreau, C, Graeff, R, Potter, B.V.L, Lee, H.C, Hao, Q. | | Deposit date: | 2007-04-09 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Catalysis-associated Conformational Changes Revealed by Human CD38 Complexed with a Non-hydrolyzable Substrate Analog
J.Biol.Chem., 282, 2007
|
|
2PGJ
 
 | | Catalysis associated conformational changes revealed by human cd38 complexed with a non-hydrolyzable substrate analog | | Descriptor: | ADP-ribosyl cyclase 1, N1-CYCLIC INOSINE 5'-DIPHOSPHORIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Moreau, C, Graeff, R, Potter, B.V.L, Lee, H.C, Hao, Q. | | Deposit date: | 2007-04-09 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Catalysis-associated Conformational Changes Revealed by Human CD38 Complexed with a Non-hydrolyzable Substrate Analog
J.Biol.Chem., 282, 2007
|
|
4TKQ
 
 | | Native-SAD phasing for YetJ from Bacillus Subtilis | | Descriptor: | CALCIUM ION, CHLORIDE ION, Uncharacterized protein YetJ | | Authors: | Liu, Q, Chang, Y, Hendrickson, W.A, New York Consortium on Membrane Protein Structure (NYCOMPS) | | Deposit date: | 2014-05-27 | | Release date: | 2014-06-18 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8025 Å) | | Cite: | Multi-crystal native SAD analysis at 6 keV.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
3DZH
 
 | | Crystal structure of human CD38 extracellular domain, GTP complex | | Descriptor: | ADP-ribosyl cyclase 1, GUANOSINE-5'-TRIPHOSPHATE | | Authors: | Liu, Q, Kriksunov, I.A, Jiang, H, Graeff, R, Lin, H, Lee, H.C, Hao, Q. | | Deposit date: | 2008-07-29 | | Release date: | 2008-11-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Covalent and Noncovalent Intermediates of an NAD Utilizing Enzyme, Human CD38.
Chem.Biol., 15, 2008
|
|
3DZJ
 
 | | Crystal structure of human CD38 extracellular domain E226Q mutant, NMN complex | | Descriptor: | ADP-ribosyl cyclase 1, BETA-NICOTINAMIDE RIBOSE MONOPHOSPHATE | | Authors: | Liu, Q, Kriksunov, I.A, Jiang, H, Graeff, R, Lin, H, Lee, H.C, Hao, Q. | | Deposit date: | 2008-07-29 | | Release date: | 2008-11-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Covalent and Noncovalent Intermediates of an NAD Utilizing Enzyme, Human CD38.
Chem.Biol., 15, 2008
|
|
3DZK
 
 | | Crystal structure of human CD38 extracellular domain, NMN complex | | Descriptor: | ADP-ribosyl cyclase 1, BETA-NICOTINAMIDE RIBOSE MONOPHOSPHATE | | Authors: | Liu, Q, Kriksunov, I.A, Jiang, H, Graeff, R, Lin, H, Lee, H.C, Hao, Q. | | Deposit date: | 2008-07-29 | | Release date: | 2008-11-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Covalent and Noncovalent Intermediates of an NAD Utilizing Enzyme, Human CD38.
Chem.Biol., 15, 2008
|
|
3DZF
 
 | | Crystal structure of human CD38 extracellular domain complexed with a covalent intermediate, ara-F-ribose-5'-phosphate | | Descriptor: | 2-deoxy-2-fluoro-5-O-phosphono-alpha-D-arabinofuranose, ADP-ribosyl cyclase 1 | | Authors: | Liu, Q, Kriksunov, I.A, Jiang, H, Graeff, R, Lin, H, Lee, H.C, Hao, Q. | | Deposit date: | 2008-07-29 | | Release date: | 2008-11-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Covalent and Noncovalent Intermediates of an NAD Utilizing Enzyme, Human CD38.
Chem.Biol., 15, 2008
|
|
3DZI
 
 | | Crystal structure of human CD38 extracellular domain, ribose-5'-phosphate intermediate/GTP complex | | Descriptor: | 2-amino-9-{5-O-[(R)-hydroxy{[(R)-hydroxy(phosphonooxy)phosphoryl]oxy}phosphoryl]-beta-D-ribofuranosyl}-9H-purin-6-yl 5-O-phosphono-beta-D-ribofuranoside, ADP-ribosyl cyclase 1, ANY 5'-MONOPHOSPHATE NUCLEOTIDE, ... | | Authors: | Liu, Q, Kriksunov, I.A, Jiang, H, Graeff, R, Lin, H, Lee, H.C, Hao, Q. | | Deposit date: | 2008-07-29 | | Release date: | 2008-11-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Covalent and Noncovalent Intermediates of an NAD Utilizing Enzyme, Human CD38.
Chem.Biol., 15, 2008
|
|
3DZG
 
 | | Crystal structure of human CD38 extracellular domain, ara-F-ribose-5'-phosphate/nicotinamide complex | | Descriptor: | 2-deoxy-2-fluoro-5-O-phosphono-alpha-D-arabinofuranose, ADP-ribosyl cyclase 1, NICOTINAMIDE | | Authors: | Liu, Q, Kriksunov, I.A, Jiang, H, Graeff, R, Lin, H, Lee, H.C, Hao, Q. | | Deposit date: | 2008-07-29 | | Release date: | 2008-11-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Covalent and Noncovalent Intermediates of an NAD Utilizing Enzyme, Human CD38.
Chem.Biol., 15, 2008
|
|
3F6Y
 
 | | Conformational Closure of the Catalytic Site of Human CD38 Induced by Calcium | | Descriptor: | ADP-ribosyl cyclase 1, CALCIUM ION | | Authors: | Liu, Q, Graeff, R, Kriksunov, I.A, Lam, C.M.C, Lee, H.C, Hao, Q. | | Deposit date: | 2008-11-07 | | Release date: | 2008-11-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Conformational Closure of the Catalytic Site of Human CD38 Induced by Calcium
Biochemistry, 47, 2008
|
|
3QIC
 
 | | The structure of human glucokinase E339K mutation | | Descriptor: | GLYCEROL, Glucokinase, alpha-D-glucopyranose | | Authors: | Liu, Q, Liu, S, Liu, J. | | Deposit date: | 2011-01-27 | | Release date: | 2011-06-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of E339K mutated human glucokinase reveals changes in the ATP binding site.
Febs Lett., 585, 2011
|
|
3I9N
 
 | | Crystal structure of human CD38 complexed with an analog ribo-2'F-ADP ribose | | Descriptor: | ADP-ribosyl cyclase 1, [(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxytetrahydrofuran-2-yl]methyl [(2R,3R,4S)-4-fluoro-3-hydroxytetrahydrofuran-2-yl]methyl dihydrogen diphosphate | | Authors: | Liu, Q, Graeff, R, Kriksunov, I.A, Jiang, H, Zhang, B, Oppenheimer, N, Lin, H, Potter, B.V.L, Lee, H.C, Hao, Q. | | Deposit date: | 2009-07-12 | | Release date: | 2009-07-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Structural basis for enzymatic evolution from a dedicated ADP-ribosyl cyclase to a multifunctional NAD hydrolase
J.Biol.Chem., 284, 2009
|
|
3I9K
 
 | | Crystal structure of ADP ribosyl cyclase complexed with substrate NAD | | Descriptor: | ADP-ribosyl cyclase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Liu, Q, Graeff, R, Kriksunov, I.A, Jiang, H, Zhang, B, Oppenheimer, N, Lin, H, Potter, B.V.L, Lee, H.C, Hao, Q. | | Deposit date: | 2009-07-12 | | Release date: | 2009-07-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Structural basis for enzymatic evolution from a dedicated ADP-ribosyl cyclase to a multifunctional NAD hydrolase
J.Biol.Chem., 284, 2009
|
|
