6S41
 
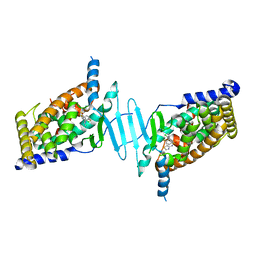 | | CRYSTAL STRUCTURE OF PXR IN COMPLEX WITH XPC-7455 | | Descriptor: | 4-[[(1~{S})-1-[2,5-bis(fluoranyl)phenyl]ethyl]amino]-5-chloranyl-2-fluoranyl-~{N}-(1,3-thiazol-4-yl)benzenesulfonamide, Nuclear receptor subfamily 1 group I member 2 | | Authors: | Focken, T, Maskos, K, Griessner, A, Krapp, S. | | Deposit date: | 2019-06-26 | | Release date: | 2019-10-02 | | Last modified: | 2019-11-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Identification of CNS-Penetrant Aryl Sulfonamides as Isoform-Selective NaV1.6 Inhibitors with Efficacy in Mouse Models of Epilepsy.
J.Med.Chem., 62, 2019
|
|
4F1L
 
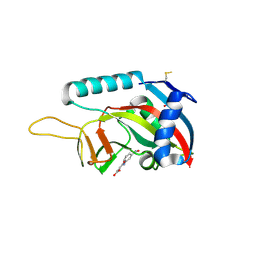 | | Human Artd8 (Parp14, Bal2) - catalytic domain in complex with inhibitor A16(Z) | | Descriptor: | (2Z)-4-[(3-carbamoylphenyl)amino]-4-oxobut-2-enoic acid, NITRATE ION, Poly [ADP-ribose] polymerase 14 | | Authors: | Karlberg, T, Andersson, C.D, Lindgren, A, Spjut, S, Thorsell, A.G, Ekblad, T, Weigelt, J, Elofsson, M, Linusson, A, Schuler, H. | | Deposit date: | 2012-05-07 | | Release date: | 2012-09-05 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Discovery of Ligands for ADP-Ribosyltransferases via Docking-Based Virtual Screening.
J.Med.Chem., 55, 2012
|
|
4F0E
 
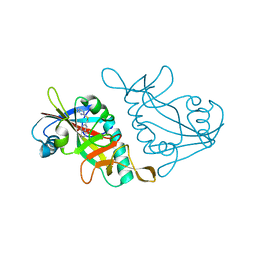 | | Human ADP-RIBOSYLTRANSFERASE 7 (ARTD7/PARP15), CATALYTIC DOMAIN IN COMPLEX WITH STO1102 | | Descriptor: | 8-methyl-2-[(pyrimidin-2-ylsulfanyl)methyl]quinazolin-4(1H)-one, Poly [ADP-ribose] polymerase 15 | | Authors: | Karlberg, T, Andersson, C.D, Lindgren, A, Thorsell, A.G, Ekblad, T, Spjut, S, Weigelt, J, Elofsson, M, Linusson, A, Schuler, H. | | Deposit date: | 2012-05-04 | | Release date: | 2012-09-05 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Discovery of Ligands for ADP-Ribosyltransferases via Docking-Based Virtual Screening.
J.Med.Chem., 55, 2012
|
|
4F1Q
 
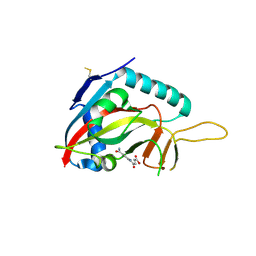 | | Human Artd8 (Parp14, Bal2) - catalytic domain in complex with A16(E) | | Descriptor: | (2E)-4-[(3-carbamoylphenyl)amino]-4-oxobut-2-enoic acid, NITRATE ION, Poly [ADP-ribose] polymerase 14 | | Authors: | Karlberg, T, Andersson, C.D, Lindgren, A, Spjut, S, Thorsell, A.G, Ekblad, T, Weigelt, J, Elofsson, M, Linusson, A, Schuler, H. | | Deposit date: | 2012-05-07 | | Release date: | 2012-09-05 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Discovery of Ligands for ADP-Ribosyltransferases via Docking-Based Virtual Screening.
J.Med.Chem., 55, 2012
|
|
4B78
 
 | | Aminoimidazoles as BACE-1 Inhibitors: From De Novo Design to Ab- lowering in Brain | | Descriptor: | (3R,5R)-3-methoxy-5-(4-methoxyphenyl)-5-(3-pyridin-3-ylphenyl)-3,4-dihydropyrrol-2-amine, BETA-SECRETASE 1 | | Authors: | Gravenfors, Y, Blid, J, Ginman, T, Karlstrom, S, Kihlstrom, J, Kolmodin, K, Lindstrom, J, Berg, S, Kieseritzky, F, Slivo, C, Swahn, B, Viklund, J, Olsson, L, Johansson, P, Eketjall, S, Falting, J, Jeppsson, F, Stromberg, K, Janson, J, Rahm, F. | | Deposit date: | 2012-08-16 | | Release date: | 2013-06-26 | | Last modified: | 2013-07-03 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Core Refinement Toward Permeable Beta-Secretase (Bace-1) Inhibitors with Low Herg Activity.
J.Med.Chem., 56, 2013
|
|
4B72
 
 | | Aminoimidazoles as BACE-1 Inhibitors: From De Novo Design to Ab- lowering in Brain | | Descriptor: | (6R)-6-(4-methoxyphenyl)-2-methyl-6-(3-pyrimidin-5-ylphenyl)pyrrolo[3,4-d][1,3]thiazol-4-amine, BETA-SECRETASE 1 | | Authors: | Gravenfors, Y, Blid, J, Ginman, T, Karlstrom, S, Kihlstrom, J, Kolmodin, K, Lindstrom, J, Berg, S, Kieseritzky, F, Slivo, C, Swahn, B, Viklund, J, Olsson, L, Johansson, P, Eketjall, S, Falting, J, Jeppsson, F, Stromberg, K, Janson, J, Rahm, F. | | Deposit date: | 2012-08-16 | | Release date: | 2013-06-26 | | Last modified: | 2013-07-03 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Core Refinement Toward Permeable Beta-Secretase (Bace-1) Inhibitors with Low Herg Activity.
J.Med.Chem., 56, 2013
|
|
4B70
 
 | | Aminoimidazoles as BACE-1 Inhibitors: From De Novo Design to Ab- lowering in Brain | | Descriptor: | (2S)-2-[3-(3-chlorophenyl)phenyl]-2-methyl-5,6-dihydro-1,3-oxazin-4-amine, BETA-SECRETASE 1 | | Authors: | Gravenfors, Y, Blid, J, Ginman, T, Karlstrom, S, Kihlstrom, J, Kolmodin, K, Lindstrom, J, Berg, S, Kieseritzky, F, Slivo, C, Swahn, B, Viklund, J, Olsson, L, Johansson, P, Eketjall, S, Falting, J, Jeppsson, F, Stromberg, K, Janson, J, Rahm, F. | | Deposit date: | 2012-08-16 | | Release date: | 2013-06-26 | | Last modified: | 2013-07-03 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Core Refinement Toward Permeable Beta-Secretase (Bace-1) Inhibitors with Low Herg Activity.
J.Med.Chem., 56, 2013
|
|
4B77
 
 | | Aminoimidazoles as BACE-1 Inhibitors: From De Novo Design to Ab- lowering in Brain | | Descriptor: | (5R)-5-(4-methoxyphenyl)-5-(3-pyrimidin-5-ylphenyl)-3,4-dihydropyrrol-2-amine, BETA-SECRETASE 1, DIMETHYL SULFOXIDE | | Authors: | Gravenfors, Y, Blid, J, Ginman, T, Karlstrom, S, Kihlstrom, J, Kolmodin, K, Lindstrom, J, Berg, S, Kieseritzky, F, Slivo, C, Swahn, B, Viklund, J, Olsson, L, Johansson, P, Eketjall, S, Falting, J, Jeppsson, F, Stromberg, K, Janson, J, Rahm, F. | | Deposit date: | 2012-08-16 | | Release date: | 2013-06-26 | | Last modified: | 2013-07-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Core Refinement Toward Permeable Beta-Secretase (Bace-1) Inhibitors with Low Herg Activity.
J.Med.Chem., 56, 2013
|
|
