3KXC
 
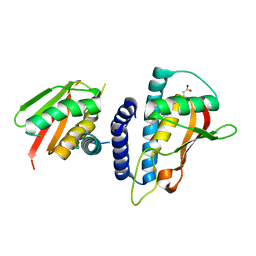 | | Mutant transport protein | | Descriptor: | PALMITIC ACID, Trafficking protein particle complex subunit 3, Trafficking protein particle complex subunit 6B | | Authors: | Kummel, D, Heinemann, U. | | Deposit date: | 2009-12-02 | | Release date: | 2010-04-21 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Characterization of the self-palmitoylation activity of the transport protein particle component Bet3
Cell.Mol.Life Sci., 67, 2010
|
|
2BJN
 
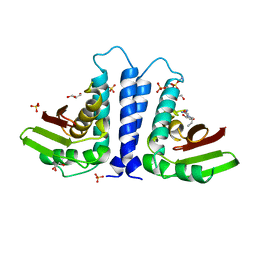 | | X-ray Structure of human TPC6 | | Descriptor: | GLYCEROL, SULFATE ION, TRAFFICKING PROTEIN PARTICLE COMPLEX SUBUNIT 6B | | Authors: | Kummel, D, Mueller, J.J, Roske, Y, Misselwitz, R, Bussow, K, Heinemann, U. | | Deposit date: | 2005-02-04 | | Release date: | 2005-07-20 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Structure of the Trapp Subunit Tpc6 Suggests a Model for a Trapp Subcomplex.
Embo Rep., 6, 2005
|
|
2CFH
 
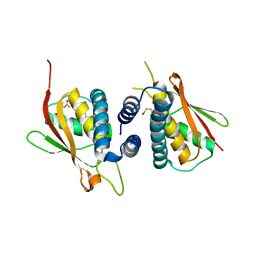 | | Structure of the Bet3-TPC6B core of TRAPP | | Descriptor: | PALMITIC ACID, TRAFFICKING PROTEIN PARTICLE COMPLEX SUBUNIT 3, TRAFFICKING PROTEIN PARTICLE COMPLEX SUBUNIT 6B | | Authors: | Kummel, D, Muller, J.J, Roske, Y, Henke, N, Heinemann, U. | | Deposit date: | 2006-02-21 | | Release date: | 2006-07-12 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the Bet3-Tpc6B Core of Trapp: Two Tpc6 Paralogs Form Trimeric Complexes with Bet3 and Mum2.
J.Mol.Biol., 361, 2006
|
|
5HIU
 
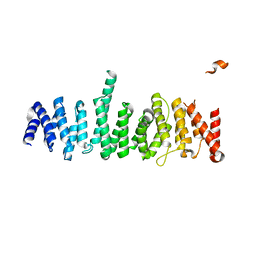 | | Structure of the TSC2 N-terminus | | Descriptor: | GTPase activator-like protein | | Authors: | Zech, R, Kiontke, S, Kummel, D. | | Deposit date: | 2016-01-12 | | Release date: | 2016-08-10 | | Last modified: | 2016-09-28 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the Tuberous Sclerosis Complex 2 (TSC2) N Terminus Provides Insight into Complex Assembly and Tuberous Sclerosis Pathogenesis.
J.Biol.Chem., 291, 2016
|
|
2W3C
 
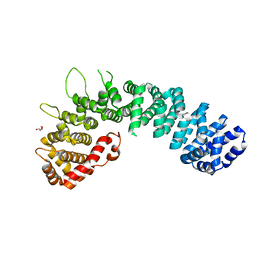 | | Globular head region of the human general vesicular transport factor p115 | | Descriptor: | DI(HYDROXYETHYL)ETHER, GENERAL VESICULAR TRANSPORT FACTOR P115 | | Authors: | Striegl, H, Roske, Y, Kummel, D, Heinemann, U. | | Deposit date: | 2008-11-11 | | Release date: | 2009-03-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Unusual Armadillo Fold in the Human General Vesicular Transport Factor P115
Plos One, 4, 2009
|
|
6SSH
 
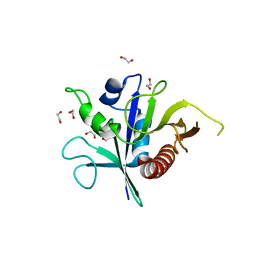 | | Structure of the TSC2 GAP domain | | Descriptor: | 1,2-ETHANEDIOL, GTPase activator-like protein | | Authors: | Hansmann, P, Kiontke, S, Kummel, D. | | Deposit date: | 2019-09-06 | | Release date: | 2020-05-27 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structure of the TSC2 GAP Domain: Mechanistic Insight into Catalysis and Pathogenic Mutations.
Structure, 28, 2020
|
|
7FSF
 
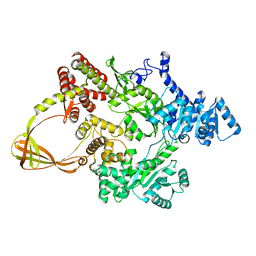 | | CRYSTAL STRUCTURE OF T. MARITIMA REVERSE GYRASE ACTIVE SITE VARIANT Y851F | | Descriptor: | Reverse gyrase, ZINC ION | | Authors: | Rasche, R, Kummel, D, Rudolph, M.G, Klostermeier, D. | | Deposit date: | 2023-01-04 | | Release date: | 2023-05-10 | | Last modified: | 2023-07-12 | | Method: | X-RAY DIFFRACTION (2.77 Å) | | Cite: | Structure of reverse gyrase with a minimal latch that supports ATP-dependent positive supercoiling without specific interactions with the topoisomerase domain.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
7FSE
 
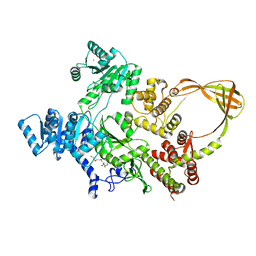 | | Crystal Structure of T. maritima reverse gyrase with a minimal latch | | Descriptor: | CHLORIDE ION, DODECAETHYLENE GLYCOL, Reverse gyrase, ... | | Authors: | Rasche, R, Kummel, D, Rudolph, M.G, Klostermeier, D. | | Deposit date: | 2023-01-04 | | Release date: | 2023-05-10 | | Last modified: | 2023-07-12 | | Method: | X-RAY DIFFRACTION (2.89 Å) | | Cite: | Structure of reverse gyrase with a minimal latch that supports ATP-dependent positive supercoiling without specific interactions with the topoisomerase domain.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
1TBX
 
 | | Crystal structure of SSV1 F-93 | | Descriptor: | Hypothetical 11.0 kDa protein | | Authors: | Kraft, P, Oeckinghaus, A, Kummel, D, Gauss, G.H, Wiedenheft, B, Young, M, Lawrence, C.M. | | Deposit date: | 2004-05-20 | | Release date: | 2004-07-13 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of F-93 from Sulfolobus spindle-shaped virus 1, a winged-helix DNA binding protein.
J.Virol., 78, 2004
|
|
8OFB
 
 | | Crystal Structure of T. maritima reverse gyrase with a minimal latch, hexagonal form | | Descriptor: | CHLORIDE ION, HEXAETHYLENE GLYCOL, Reverse gyrase, ... | | Authors: | Klostermeier, D, Rasche, R, Mhaindarkar, V, Kummel, D, Rudolph, M.G. | | Deposit date: | 2023-03-15 | | Release date: | 2023-04-26 | | Last modified: | 2023-06-07 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Structure of reverse gyrase with a minimal latch that supports ATP-dependent positive supercoiling without specific interactions with the topoisomerase domain.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
6ZFM
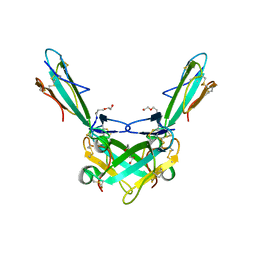 
 | | Structure of alpha-Cobratoxin with a peptide inhibitor | | Descriptor: | 3-[2-[2-[2-[2-[2-(2-azanylethoxy)ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]propan-1-ol, Alpha-cobratoxin, PENTAETHYLENE GLYCOL, ... | | Authors: | Kiontke, S, Kummel, D. | | Deposit date: | 2020-06-17 | | Release date: | 2020-12-02 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Peptide Inhibitors of the alpha-Cobratoxin-Nicotinic Acetylcholine Receptor Interaction.
J.Med.Chem., 63, 2020
|
|
8CPN
 
 | | Crystal structure of the PolB16_OarG intein variant S1A, N183A | | Descriptor: | IODIDE ION, PolB16 intein | | Authors: | Kattelmann, S, Pasch, T, Mootz, H.D, Kummel, D. | | Deposit date: | 2023-03-03 | | Release date: | 2023-05-17 | | Last modified: | 2023-05-31 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural and biochemical analysis of a novel atypically split intein reveals a conserved histidine specific to cysteine-less inteins.
Chem Sci, 14, 2023
|
|
3TW8
 
 | | GEF domain of DENND 1B in complex with Rab GTPase Rab35 | | Descriptor: | DENN domain-containing protein 1B, Ras-related protein Rab-35 | | Authors: | Wu, X.D, Kummel, D, Reinisch, K.M. | | Deposit date: | 2011-09-21 | | Release date: | 2011-11-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Insights regarding guanine nucleotide exchange from the structure of a DENN-domain protein complexed with its Rab GTPase substrate.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3RK3
 
 | | Truncated SNARE complex with complexin | | Descriptor: | Complexin-1, SNAP25, Syntaxin 1a, ... | | Authors: | Kuemmel, D, Reinisch, K.M. | | Deposit date: | 2011-04-17 | | Release date: | 2011-07-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Complexin cross-links prefusion SNAREs into a zigzag array.
Nat.Struct.Mol.Biol., 18, 2011
|
|
3RL0
 
 | | Truncated SNARE complex with complexin (P1) | | Descriptor: | Complexin-1, Synaptosomal-associated protein 25, Syntaxin-1A, ... | | Authors: | Kuemmel, D, Reinisch, K.M. | | Deposit date: | 2011-04-19 | | Release date: | 2011-07-27 | | Last modified: | 2011-08-24 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | Complexin cross-links prefusion SNAREs into a zigzag array.
Nat.Struct.Mol.Biol., 18, 2011
|
|
3RK2
 
 | | Truncated SNARE complex | | Descriptor: | CALCIUM ION, Synaptosomal-associated protein 25, Syntaxin-1A, ... | | Authors: | Kuemmel, D, Reinisch, K.M. | | Deposit date: | 2011-04-17 | | Release date: | 2011-07-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Complexin cross-links prefusion SNAREs into a zigzag array.
Nat.Struct.Mol.Biol., 18, 2011
|
|
4UUY
 
 | | Structural Identification of the Vps18 beta-propeller reveals a critical role in the HOPS complex stability and function. | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, SULFATE ION, ... | | Authors: | Behrmann, H, Gohlke, U, Heinemann, U. | | Deposit date: | 2014-08-01 | | Release date: | 2014-10-29 | | Last modified: | 2014-12-17 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Structural Identification of the Vps18 Beta-Propeller Reveals a Critical Role in the Hops Complex Stability and Function.
J.Biol.Chem., 289, 2014
|
|
7ZTY
 
 | |
5LDD
 
 | |
7P9D
 
 | | Crystal structure of Chlamydomonas reinhardtii NADPH Dependent Thioredoxin Reductase 1 domain | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Thioredoxin reductase | | Authors: | Singh, R.K, Marchetti, G.M, Hippler, M, Kuemmel, D. | | Deposit date: | 2021-07-27 | | Release date: | 2022-01-12 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Structural analysis revealed a novel conformation of the NTRC reductase domain from Chlamydomonas reinhardtii.
J.Struct.Biol., 214, 2021
|
|
7ZU0
 
 | | HOPS tethering complex from yeast | | Descriptor: | E3 ubiquitin-protein ligase PEP5, Vacuolar membrane protein PEP3, Vacuolar morphogenesis protein 6, ... | | Authors: | Shvarev, D, Schoppe, J, Koenig, C, Perz, A, Fuellbrunn, N, Kiontke, S, Langemeyer, L, Januliene, D, Schnelle, K, Kuemmel, D, Froehlich, F, Moeller, A, Ungermann, C. | | Deposit date: | 2022-05-11 | | Release date: | 2022-09-28 | | Last modified: | 2022-10-12 | | Method: | ELECTRON MICROSCOPY (4.4 Å) | | Cite: | Structure of the HOPS tethering complex, a lysosomal membrane fusion machinery.
Elife, 11, 2022
|
|
6LTV
 
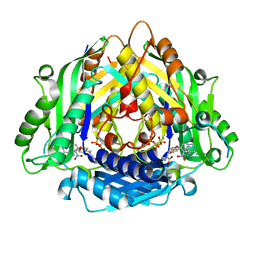 | | Crystal Structure of I122A/I330A variant of S-adenosylmethionine synthetase from Cryptosporidium hominis in complex with ONB-SAM (2-nitro benzyme S-adenosyl-methionine) | | Descriptor: | MAGNESIUM ION, S-adenosylmethionine synthase, TRIPHOSPHATE, ... | | Authors: | Singh, R.K, Michailidou, F, Rentmeister, A, Kuemmel, D. | | Deposit date: | 2020-01-23 | | Release date: | 2020-10-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Engineered SAM Synthetases for Enzymatic Generation of AdoMet Analogs with Photocaging Groups and Reversible DNA Modification in Cascade Reactions.
Angew.Chem.Int.Ed.Engl., 60, 2021
|
|
6LTW
 
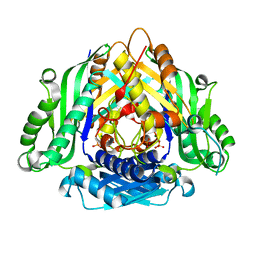 | | Crystal structure of Apo form of I122A/I330A variant of S-adenosylmethionine synthetase from Cryptosporidium hominis | | Descriptor: | MAGNESIUM ION, PHOSPHATE ION, S-adenosylmethionine synthase | | Authors: | Singh, R.K, Michailidou, F, Rentmeister, A, Kuemmel, D. | | Deposit date: | 2020-01-23 | | Release date: | 2020-10-21 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Engineered SAM Synthetases for Enzymatic Generation of AdoMet Analogs with Photocaging Groups and Reversible DNA Modification in Cascade Reactions.
Angew.Chem.Int.Ed.Engl., 60, 2021
|
|
7P9E
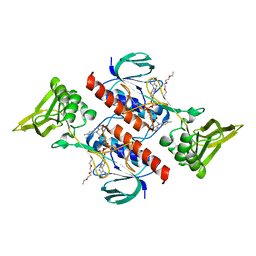 
 | |
7QLA
 
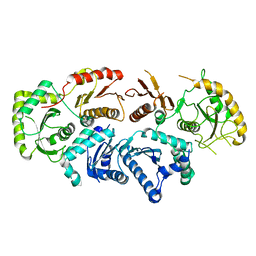 | | Structure of the Rab GEF complex Mon1-Ccz1 | | Descriptor: | Ccz1, Vacuolar fusion protein MON1 | | Authors: | Klink, B.U, Herrmann, E, Antoni, C, Langemeyer, L, Kiontke, S, Gatsogiannis, C, Ungermann, C, Raunser, S, Kuemmel, D. | | Deposit date: | 2021-12-20 | | Release date: | 2022-02-09 | | Last modified: | 2022-02-16 | | Method: | ELECTRON MICROSCOPY (3.85 Å) | | Cite: | Structure of the Mon1-Ccz1 complex reveals molecular basis of membrane binding for Rab7 activation.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
