1JRA
 
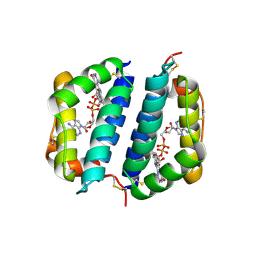 | | Crystal Structure of Erv2p | | Descriptor: | ERV2 PROTEIN, MITOCHONDRIAL, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Gross, E, Sevier, C.S, Vala, A, Kaiser, C.A, Fass, D. | | Deposit date: | 2001-08-13 | | Release date: | 2001-12-28 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A new FAD-binding fold and intersubunit disulfide shuttle in the thiol oxidase Erv2p.
Nat.Struct.Biol., 9, 2002
|
|
1JR8
 
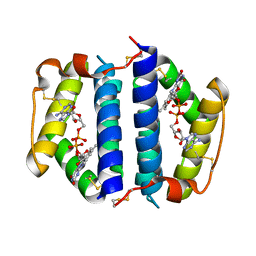 | | Crystal Structure of Erv2p | | Descriptor: | Erv2 PROTEIN, mitochondrial, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Gross, E, Sevier, C.S, Vala, A, Kaiser, C.A, Fass, D. | | Deposit date: | 2001-08-13 | | Release date: | 2001-12-28 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | A new FAD-binding fold and intersubunit disulfide shuttle in the thiol oxidase Erv2p.
Nat.Struct.Biol., 9, 2002
|
|
1RP4
 
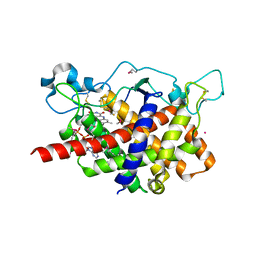 | | Structure of Ero1p, Source of Disulfide Bonds for Oxidative Protein Folding in the Cell | | Descriptor: | 1-ETHYL-PYRROLIDINE-2,5-DIONE, CADMIUM ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Gross, E, Kastner, D.B, Kaiser, C.A, Fass, D. | | Deposit date: | 2003-12-03 | | Release date: | 2004-06-08 | | Last modified: | 2018-01-31 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of ero1p, source of disulfide bonds for oxidative protein folding in the cell.
Cell(Cambridge,Mass.), 117, 2004
|
|
1RQ1
 
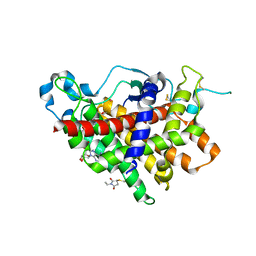 | | Structure of Ero1p, Source of Disulfide Bonds for Oxidative Protein Folding in the Cell | | Descriptor: | 1-ETHYL-PYRROLIDINE-2,5-DIONE, CADMIUM ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Gross, E, Kastner, D.B, Kaiser, C.A, Fass, D. | | Deposit date: | 2003-12-04 | | Release date: | 2004-06-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of ero1p, source of disulfide bonds for oxidative protein folding in the cell.
Cell(Cambridge,Mass.), 117, 2004
|
|
5GLG
 
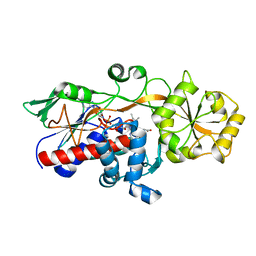 | |
5ZYN
 
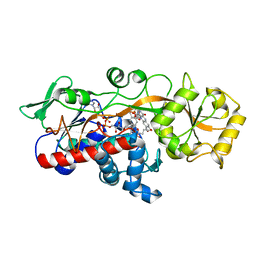 | | Fumarate reductase | | Descriptor: | FLAVIN MONONUCLEOTIDE, FLAVIN-ADENINE DINUCLEOTIDE, Fumarate reductase 2, ... | | Authors: | Park, H.H, Kim, C.M. | | Deposit date: | 2018-05-25 | | Release date: | 2018-10-03 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Molecular basis of maintaining an oxidizing environment under anaerobiosis by soluble fumarate reductase.
Nat Commun, 9, 2018
|
|
2HJ3
 
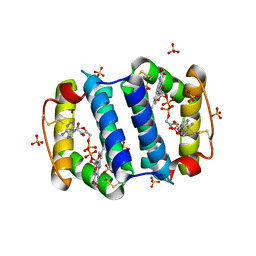 | | Structure of the Arabidopsis Thaliana Erv1 Thiol Oxidase | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, SULFATE ION, Sulfhydryl oxidase Erv1p | | Authors: | Vitu, E, Fass, D. | | Deposit date: | 2006-06-30 | | Release date: | 2006-08-22 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Gain of Function in an ERV/ALR Sulfhydryl Oxidase by Molecular Engineering of the Shuttle Disulfide.
J.Mol.Biol., 362, 2006
|
|
3ED3
 
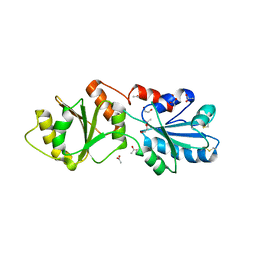 | | Crystal Structure of the Yeast Dithiol/Disulfide Oxidoreductase Mpd1p | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Protein disulfide-isomerase MPD1 | | Authors: | Vitu, E, Greenblatt, H.M, Fass, D. | | Deposit date: | 2008-09-02 | | Release date: | 2008-11-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Yeast Mpd1p reveals the structural diversity of the protein disulfide isomerase family
J.Mol.Biol., 384, 2008
|
|
3LGO
 
 | | Structure of Gse1p, member of the GSE/EGO complex | | Descriptor: | Protein SLM4 | | Authors: | Kogan, K, Fass, D. | | Deposit date: | 2010-01-21 | | Release date: | 2010-08-04 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structural conservation of components in the amino acid sensing branch of the TOR pathway in yeast and mammals.
J.Mol.Biol., 402, 2010
|
|
