2J0T
 
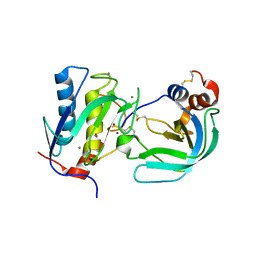 | | Crystal Structure of the Catalytic Domain of MMP-1 in Complex with the Inhibitory Domain of TIMP-1 | | Descriptor: | CALCIUM ION, INTERSTITIAL COLLAGENASE, METALLOPROTEINASE INHIBITOR 1, ... | | Authors: | Iyer, S, Wei, S, Brew, K, Acharya, K.R. | | Deposit date: | 2006-08-04 | | Release date: | 2006-10-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Crystal Structure of the Catalytic Domain of Matrix Metalloproteinase-1 in Complex with the Inhibitory Domain of Tissue Inhibitor of Metalloproteinase-1.
J.Biol.Chem., 282, 2007
|
|
1FZV
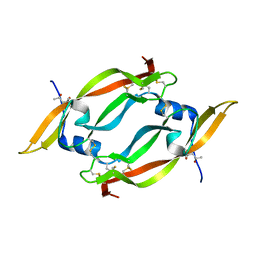 
 | | THE CRYSTAL STRUCTURE OF HUMAN PLACENTA GROWTH FACTOR-1 (PLGF-1), AN ANGIOGENIC PROTEIN AT 2.0A RESOLUTION | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PLACENTA GROWTH FACTOR | | Authors: | Iyer, S, Leonidas, D.D, Swaminathan, G.J, Maglione, D, Battisti, M, Tucci, M, Persico, M.G, Acharya, K.R. | | Deposit date: | 2000-10-04 | | Release date: | 2001-05-09 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of human placenta growth factor-1 (PlGF-1), an angiogenic protein, at 2.0 A resolution.
J.Biol.Chem., 276, 2001
|
|
3ZBV
 
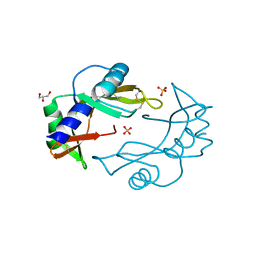 | | Crystal Structure of murine Angiogenin-2 | | Descriptor: | ANGIOGENIN-2, GLYCEROL, SULFATE ION, ... | | Authors: | Iyer, S, Holloway, D.E, Acharya, K.R. | | Deposit date: | 2012-11-13 | | Release date: | 2012-12-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Crystal Structures of Murine Angiogenin-2 and -3 - Probing 'Structure - Function' Relationships Amongst Angiogenin Homologues.
FEBS J., 280, 2013
|
|
3ZBW
 
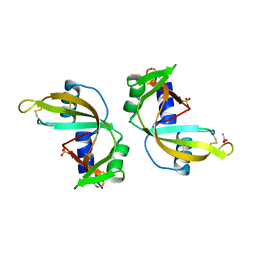 | | Crystal Structure of murine Angiogenin-3 | | Descriptor: | ACETIC ACID, ANGIOGENIN-3, SULFATE ION, ... | | Authors: | Iyer, S, Holloway, D.E, Acharya, K.R. | | Deposit date: | 2012-11-13 | | Release date: | 2012-12-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.801 Å) | | Cite: | Crystal Structures of Murine Angiogenin-2 and -3 - Probing 'Structure - Function' Relationships Amongst Angiogenin Homologues.
FEBS J., 280, 2013
|
|
4S2R
 
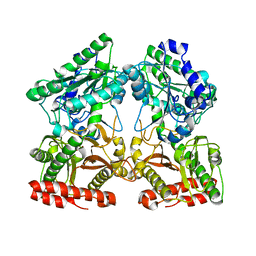 | | Crystal structure of X-prolyl aminopeptidase from Caenorhabditis elegans: a cytosolic enzyme with a di-nuclear active site | | Descriptor: | Protein APP-1, ZINC ION | | Authors: | Iyer, S, La-Borde, P, Payne, K.A.P, Parsons, M.R, Turner, A.J, Isaac, R.E, Acharya, K.R. | | Deposit date: | 2015-01-22 | | Release date: | 2015-04-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.949 Å) | | Cite: | Crystal structure of X-prolyl aminopeptidase from Caenorhabditis elegans: A cytosolic enzyme with a di-nuclear active site.
FEBS Open Bio, 5, 2015
|
|
4S2T
 
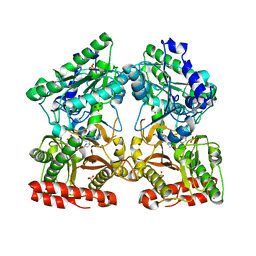 | | Crystal structure of X-prolyl aminopeptidase from Caenorhabditis elegans: a cytosolic enzyme with a di-nuclear active site | | Descriptor: | Protein APP-1, SULFATE ION, ZINC ION, ... | | Authors: | Iyer, S, La-Borde, P, Payne, K.A.P, Parsons, M.R, Turner, A.J, Isaac, R.E, Acharya, K.R. | | Deposit date: | 2015-01-22 | | Release date: | 2015-04-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal structure of X-prolyl aminopeptidase from Caenorhabditis elegans: A cytosolic enzyme with a di-nuclear active site.
FEBS Open Bio, 5, 2015
|
|
2XAC
 
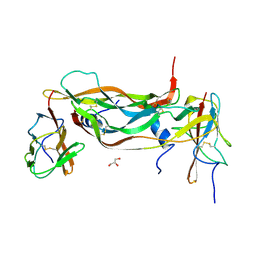 | |
2BEX
 
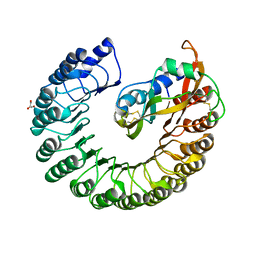 | | Crystal structure of Placental Ribonuclease Inhibitor in complex with Human Eosinophil Derived Neurotoxin at 2A resolution | | Descriptor: | ALPHA-KETOMALONIC ACID, GLYCEROL, NONSECRETORY RIBONUCLEASE, ... | | Authors: | Iyer, S, Holloway, D.E, Kumar, K, Shapiro, R, Acharya, K.R. | | Deposit date: | 2004-12-01 | | Release date: | 2005-03-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Molecular Recognition of Human Eosinophil-Derived Neurotoxin (Rnase 2) by Placental Ribonuclease Inhibitor
J.Mol.Biol., 347, 2005
|
|
2CLT
 
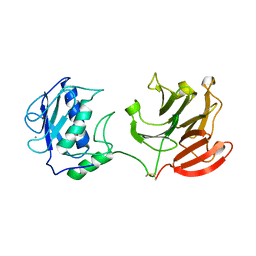 | | Crystal structure of the active form (full-length) of human fibroblast collagenase. | | Descriptor: | CALCIUM ION, INTERSTITIAL COLLAGENASE, ZINC ION | | Authors: | Iyer, S, Visse, R, Nagase, H, Acharya, K.R. | | Deposit date: | 2006-04-29 | | Release date: | 2006-08-09 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.67 Å) | | Cite: | Crystal Structure of an Active Form of Human Mmp-1.
J.Mol.Biol., 362, 2006
|
|
2C7W
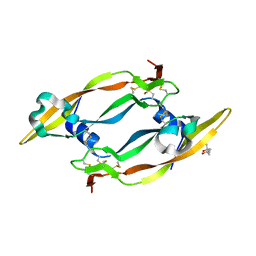 
 | | Crystal Structure of human vascular endothelial growth factor-B: Identification of amino acids important for angiogeninc activity | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, VASCULAR ENDOTHELIAL GROWTH FACTOR B PRECURSOR | | Authors: | Iyer, S, Scotney, P.D, Nash, A.D, Acharya, K.R. | | Deposit date: | 2005-11-29 | | Release date: | 2007-01-30 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Crystal Structure of Human Vascular Endothelial Growth Factor-B: Identification of Amino Acids Important for Receptor Binding
J.Mol.Biol., 359, 2006
|
|
6ULH
 
 | |
6UMS
 
 | |
6P5H
 
 | |
6P5B
 
 | | Crystal Structure of MavC in Complex with Ub-UbE2N | | Descriptor: | MavC, Ubiquitin, Ubiquitin-conjugating enzyme E2 N | | Authors: | Puvar, K, Iyer, S, Negron Teron, K.I, Das, C. | | Deposit date: | 2019-05-30 | | Release date: | 2020-05-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.099 Å) | | Cite: | Legionella effector MavC targets the Ube2N~Ub conjugate for noncanonical ubiquitination.
Nat Commun, 11, 2020
|
|
2VWE
 
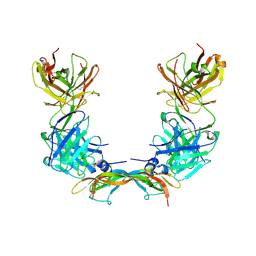 | | Crystal Structure of Vascular Endothelial Growth Factor-B in Complex with a Neutralizing Antibody Fab Fragment | | Descriptor: | ANTI-VEGF-B MONOCLONAL ANTIBODY, VASCULAR ENDOTHELIAL GROWTH FACTOR B | | Authors: | Leonard, P, Scotney, P.D, Jabeen, T, Iyer, S, Fabri, L.J, Nash, A.D, Acharya, K.R. | | Deposit date: | 2008-06-23 | | Release date: | 2008-11-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Crystal Structure of Vascular Endothelial Growth Factor-B in Complex with a Neutralising Antibody Fab Fragment.
J.Mol.Biol., 384, 2008
|
|
2WK5
 
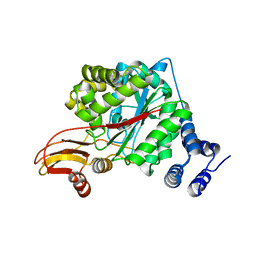 | | Structural features of native human thymidine phosphorylase and in complex with 5-iodouracil | | Descriptor: | GLYCEROL, THYMIDINE PHOSPHORYLASE | | Authors: | Mitsiki, E, Papageorgiou, A.C, Iyer, S, Thiyagarajan, N, Prior, S.H, Sleep, D, Finnis, C, Acharya, K.R. | | Deposit date: | 2009-06-05 | | Release date: | 2009-07-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | Structures of Native Human Thymidine Phosphorylase and in Complex with 5-Iodouracil.
Biochem.Biophys.Res.Commun., 386, 2009
|
|
2WK6
 
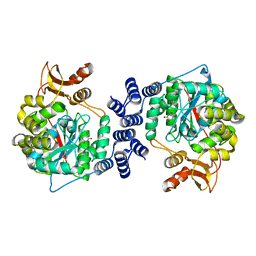 | | Structural features of native human thymidine phosphorylase and in complex with 5-iodouracil | | Descriptor: | 5-IODOURACIL, THYMIDINE PHOSPHORYLASE | | Authors: | Mitsiki, E, Papageorgiou, A.C, Iyer, S, Thiyagarajan, N, Prior, S.H, Sleep, D, Finnis, C, Acharya, K.R. | | Deposit date: | 2009-06-05 | | Release date: | 2009-07-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structures of Native Human Thymidine Phosphorylase and in Complex with 5-Iodouracil.
Biochem.Biophys.Res.Commun., 386, 2009
|
|
6UMP
 
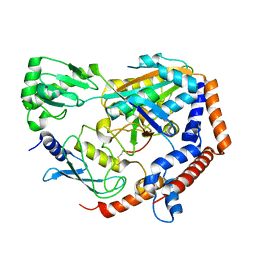 | | Crystal structure of MavC in complex with substrate mimic in P65 space group | | Descriptor: | MavC, Ubiquitin, Ubiquitin-conjugating enzyme E2 N | | Authors: | Puvar, K, Iyer, S, Luo, Z.Q, Das, C. | | Deposit date: | 2019-10-10 | | Release date: | 2020-05-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Legionella effector MavC targets the Ube2N~Ub conjugate for noncanonical ubiquitination.
Nat Commun, 11, 2020
|
|
7LK4
 
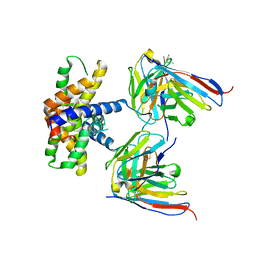 | |
6EB6
 
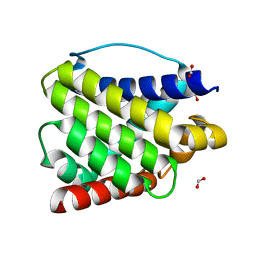 | | Crystal structure of BAX W139A monomer | | Descriptor: | Apoptosis regulator BAX, FORMIC ACID | | Authors: | Robin, A.Y. | | Deposit date: | 2018-08-05 | | Release date: | 2019-04-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.023 Å) | | Cite: | BAX Activation: Mutations Near Its Proposed Non-canonical BH3 Binding Site Reveal Allosteric Changes Controlling Mitochondrial Association.
Cell Rep, 27, 2019
|
|
6OAM
 
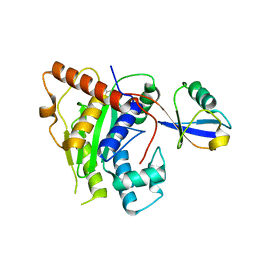 | | Crystal Structure of ChlaDUB2 DUB domain | | Descriptor: | Deubiquitinase and deneddylase Dub2, Ubiquitin | | Authors: | Hausman, J.M, Das, C. | | Deposit date: | 2019-03-17 | | Release date: | 2020-04-22 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.503 Å) | | Cite: | The Two Deubiquitinating Enzymes fromChlamydia trachomatisHave Distinct Ubiquitin Recognition Properties.
Biochemistry, 59, 2020
|
|
7VKE
 
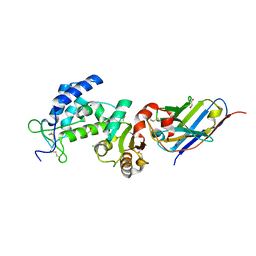 | | Crystal structure of human CD38 ECD in complex with UniDab(TM) F11A | | Descriptor: | 1,2-ETHANEDIOL, ADP-ribosyl cyclase/cyclic ADP-ribose hydrolase 1, CHLORIDE ION, ... | | Authors: | Schooten, W.V, Schellenberger, U, Ugamraj, H.S, Manicka, S, Bijpuria, S, Gondu, R.K. | | Deposit date: | 2021-09-29 | | Release date: | 2022-08-03 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | TNB-738, a biparatopic antibody, boosts intracellular NAD+ by inhibiting CD38 ecto-enzyme activity.
Mabs, 14, 2022
|
|
6MRN
 
 | | Crystal Structure of ChlaDUB2 DUB domain | | Descriptor: | Deubiquitinase and deneddylase Dub2 | | Authors: | Hausman, J.M, Das, C. | | Deposit date: | 2018-10-15 | | Release date: | 2019-10-30 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | The Two Deubiquitinating Enzymes fromChlamydia trachomatisHave Distinct Ubiquitin Recognition Properties.
Biochemistry, 59, 2020
|
|
7K02
 
 | |
6WTG
 
 | | SdeA DUB Domain in complex with Ubiquitin | | Descriptor: | Ubiquitin, Ubiquitinating/deubiquitinating enzyme SdeA | | Authors: | Kenny, S, Sheedlo, M, Das, C. | | Deposit date: | 2020-05-02 | | Release date: | 2021-03-03 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Insights into Ubiquitin Product Release in Hydrolysis Catalyzed by the Bacterial Deubiquitinase SdeA.
Biochemistry, 60, 2021
|
|
