4ZM3
 
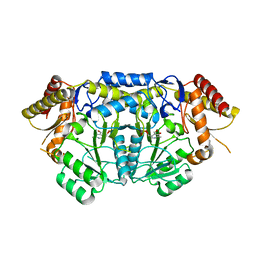 | | Crystal structure of PLP-Dependent 3-Aminobenzoate Synthase PctV wild-type | | Descriptor: | Aminotransferase, DI(HYDROXYETHYL)ETHER, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Hirayama, A, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2015-05-02 | | Release date: | 2015-10-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Mechanism-Based Trapping of the Quinonoid Intermediate by Using the K276R Mutant of PLP-Dependent 3-Aminobenzoate Synthase PctV in the Biosynthesis of Pactamycin.
Chembiochem, 16, 2015
|
|
4ZM4
 
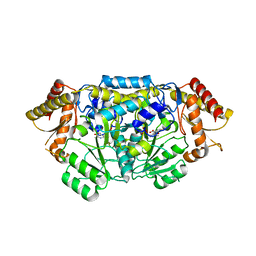 | | Complex structure of PctV K276R mutant with PMP and 3-dehydroshkimate | | Descriptor: | (3E,4R,5R)-4,5-dihydroxy-3-{[(Z)-{3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4(1H)-ylidene}methyl]imino}cyclohex-1-ene-1-carboxylic acid, Aminotransferase, PYRIDOXAL-5'-PHOSPHATE | | Authors: | Hirayama, A, Miyanaga, A, Kudo, F, Eguchi, T. | | Deposit date: | 2015-05-02 | | Release date: | 2015-10-14 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Mechanism-Based Trapping of the Quinonoid Intermediate by Using the K276R Mutant of PLP-Dependent 3-Aminobenzoate Synthase PctV in the Biosynthesis of Pactamycin.
Chembiochem, 16, 2015
|
|
6JW7
 
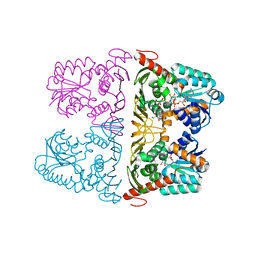 | | The crystal structure of KanD2 in complex with NADH and 3"-deamino-3"-hydroxykanamycin A | | Descriptor: | (2R,3S,4S,5R,6R)-2-(aminomethyl)-6-[(1R,2S,3S,4R,6S)-4,6-bis(azanyl)-3-[(2S,3R,4S,5S,6R)-6-(hydroxymethyl)-3,4,5-tris(oxidanyl)oxan-2-yl]oxy-2-oxidanyl-cyclohexyl]oxy-oxane-3,4,5-triol, 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, Dehydrogenase | | Authors: | Kudo, F, Kitayama, Y, Miyanaga, A, Hirayama, A, Eguchi, T. | | Deposit date: | 2019-04-18 | | Release date: | 2020-04-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Biochemical and structural analysis of a dehydrogenase, KanD2, and an aminotransferase, KanS2, that are responsible for the construction of the kanosamine moiety in kanamycin biosynthesis.
Biochemistry, 59, 2020
|
|
6JW6
 
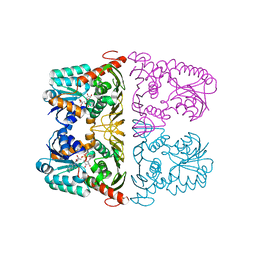 | | The crystal structure of KanD2 in complex with NAD | | Descriptor: | Dehydrogenase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Kudo, F, Kitayama, Y, Miyanaga, A, Hirayama, A, Eguchi, T. | | Deposit date: | 2019-04-18 | | Release date: | 2020-04-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Biochemical and structural analysis of a dehydrogenase, KanD2, and an aminotransferase, KanS2, that are responsible for the construction of the kanosamine moiety in kanamycin biosynthesis.
Biochemistry, 59, 2020
|
|
6JW8
 
 | | The crystal structure of KanD2 in complex with NADH and 3"-deamino-3"-hydroxykanamycin B | | Descriptor: | (2S,3R,4S,5S,6R)-2-[(1S,2S,3R,4S,6R)-3-[(2R,3R,4R,5S,6R)-6-(aminomethyl)-3-azanyl-4,5-bis(oxidanyl)oxan-2-yl]oxy-4,6-bis(azanyl)-2-oxidanyl-cyclohexyl]oxy-6-(hydroxymethyl)oxane-3,4,5-triol, 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, Dehydrogenase | | Authors: | Kudo, F, Kitayama, Y, Miyanaga, A, Hirayama, A, Eguchi, T. | | Deposit date: | 2019-04-18 | | Release date: | 2020-04-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Biochemical and structural analysis of a dehydrogenase, KanD2, and an aminotransferase, KanS2, that are responsible for the construction of the kanosamine moiety in kanamycin biosynthesis.
Biochemistry, 59, 2020
|
|
