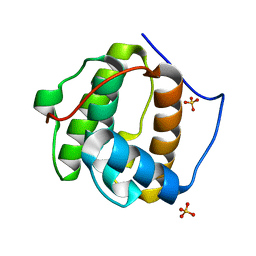
1PA7
 
 | |
2GW3
 
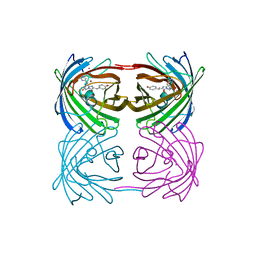 | | Crystal structure of stony coral fluorescent protein Kaede, green form | | Descriptor: | Kaede, NICKEL (II) ION | | Authors: | Hayashi, I, Mizuno, H, Miyawaki, A, Ikura, M. | | Deposit date: | 2006-05-03 | | Release date: | 2007-05-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystallographic evidence for water-assisted photo-induced peptide cleavage in the stony coral fluorescent protein Kaede.
J.Mol.Biol., 372, 2007
|
|
2GW4
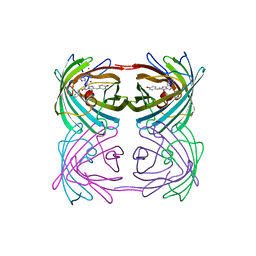 
 | | Crystal structure of stony coral fluorescent protein Kaede, red form | | Descriptor: | Kaede, NICKEL (II) ION | | Authors: | Hayashi, I, Mizuno, H, Miyawako, A, Ikura, M. | | Deposit date: | 2006-05-03 | | Release date: | 2007-05-08 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystallographic evidence for water-assisted photo-induced peptide cleavage in the stony coral fluorescent protein Kaede.
J.Mol.Biol., 372, 2007
|
|
2HQH
 
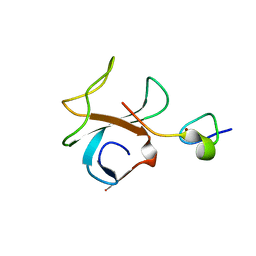 | | Crystal structure of p150Glued and CLIP-170 | | Descriptor: | Dynactin-1, Restin, ZINC ION | | Authors: | Hayashi, I, Ikura, M. | | Deposit date: | 2006-07-18 | | Release date: | 2007-08-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | CLIP170 autoinhibition mimics intermolecular interactions with p150Glued or EB1.
Nat.Struct.Mol.Biol., 14, 2007
|
|
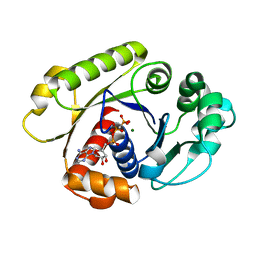
1G3R
 
 | |
1G3Q
 
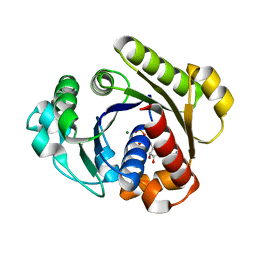 | |
7C7Y
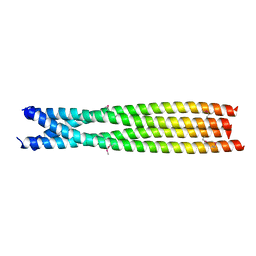 
 | | C-terminal domain of B. cereus TubY | | Descriptor: | Uncharacterized protein | | Authors: | Hayashi, I. | | Deposit date: | 2020-05-27 | | Release date: | 2020-10-28 | | Last modified: | 2021-03-17 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The C-terminal region of the plasmid partitioning protein TubY is a tetramer that can bind membranes and DNA.
J.Biol.Chem., 295, 2020
|
|
6AHT
 
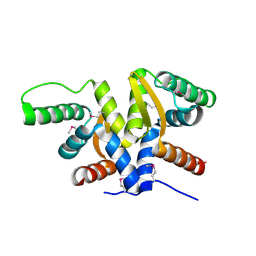 | | Plasmid partitioning protein TubR from Bacillus cereus | | Descriptor: | Conserved hypothetical plasmid protein | | Authors: | Hayashi, I. | | Deposit date: | 2018-08-20 | | Release date: | 2019-06-26 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Cooperative DNA Binding of the Plasmid Partitioning Protein TubR from the Bacillus cereus pXO1 Plasmid.
J.Mol.Biol., 430, 2018
|
|
1K40
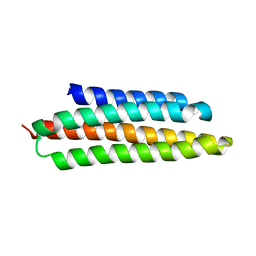 
 | |
1TXQ
 
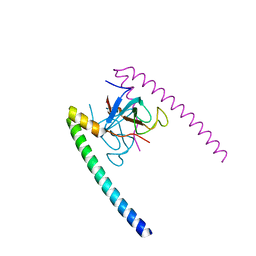 | |
1UEG
 
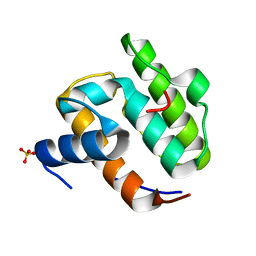 | |
8HBR
 
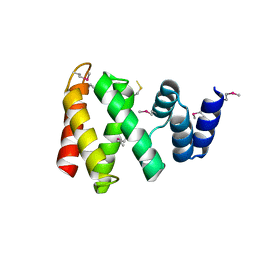 | | The C-terminal domain of Spiral2 | | Descriptor: | TOG domain-containing protein | | Authors: | Hayashi, I. | | Deposit date: | 2022-10-30 | | Release date: | 2023-01-18 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the C-terminal domain of the plant-specific microtubule-associated protein Spiral2.
Acta Crystallogr.,Sect.F, 79, 2023
|
|
4EI7
 
 | | Crystal structure of Bacillus cereus TubZ, GDP-form | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Plasmid replication protein RepX | | Authors: | Hayashi, I, Hoshino, S. | | Deposit date: | 2012-04-05 | | Release date: | 2012-08-15 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Filament formation of the FtsZ/tubulin-like protein TubZ from the Bacillus cereus pXO1 plasmid.
J.Biol.Chem., 287, 2012
|
|
4EI9
 
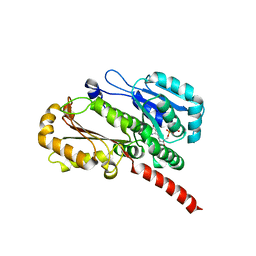 | | Crystal structure of Bacillus cereus TubZ, GTP-form | | Descriptor: | 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, GUANOSINE-5'-DIPHOSPHATE, Plasmid replication protein RepX | | Authors: | Hayashi, I, Hoshino, S. | | Deposit date: | 2012-04-05 | | Release date: | 2012-08-15 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Filament formation of the FtsZ/tubulin-like protein TubZ from the Bacillus cereus pXO1 plasmid.
J.Biol.Chem., 287, 2012
|
|
4EI8
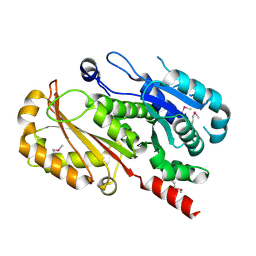 
 | |
3WOY
 
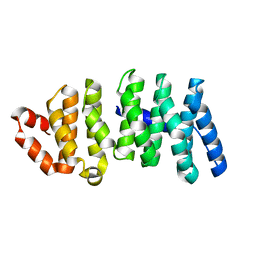 | | Crystal structure of CLASP2 TOG domain (TOG2) | | Descriptor: | CLIP-associating protein 2 | | Authors: | Hayashi, I, Maki, T. | | Deposit date: | 2014-01-06 | | Release date: | 2015-05-27 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | CLASP2 Has Two Distinct TOG Domains That Contribute Differently to Microtubule Dynamics
J. Mol. Biol., 427, 2015
|
|
3WOZ
 
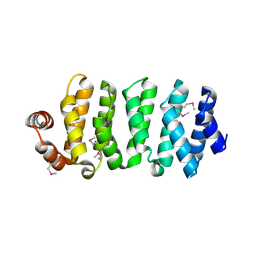 | | Crystal structure of CLASP2 TOG domain (TOG3) | | Descriptor: | CLIP-associating protein 2 | | Authors: | Hayashi, I, Maki, T. | | Deposit date: | 2014-01-06 | | Release date: | 2015-05-27 | | Last modified: | 2018-10-17 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | CLASP2 Has Two Distinct TOG Domains That Contribute Differently to Microtubule Dynamics
J. Mol. Biol., 427, 2015
|
|
