2K7E
 | | NMR structure of the human tRNALys3 bound to the HIV genome Loop I | | Descriptor: | RNA (5'-R(*CP*UP*(SUR)P*UP*UP*AP*AP*(PSU)P*CP*UP*GP*C)-3'), RNA (5'-R(*GP*CP*GP*GP*UP*GP*UP*AP*AP*AP*AP*G)-3') | | Authors: | Bilbille, Y, Vendeix, F.P.A, Guenther, R, Malkiewicz, A, Ariza, X, Vilarrasa, J, Agris, P. | | Deposit date: | 2008-08-09 | | Release date: | 2008-10-07 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | The structure of the human tRNALys3 anticodon bound to the HIV genome is stabilized by modified nucleosides and adjacent mismatch base pairs.
Nucleic Acids Res., 37, 2009
|
|
229D
 
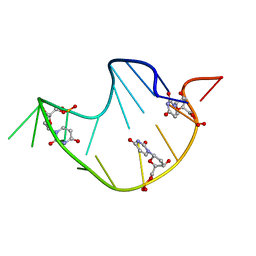 | | DNA ANALOG OF YEAST TRANSFER RNA PHE ANTICODON DOMAIN WITH MODIFIED BASES 5-METHYL CYTOSINE AND 1-METHYL GUANINE | | Descriptor: | DNA (5'-D(*CP*CP*AP*GP*AP*CP*(UMP)P*GP*AP*AP*(MG1)P*AP*(UMP)P*(5CM)P*(UMP)P*GP*G)-3') | | Authors: | Basti, M.M, Stuart, J.W, Lam, A.T, Guenther, R, Agris, P.F. | | Deposit date: | 1995-08-16 | | Release date: | 1995-12-07 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Design, biological activity and NMR-solution structure of a DNA analogue of yeast tRNA(Phe) anticodon domain.
Nat.Struct.Biol., 3, 1996
|
|
1KOS
 
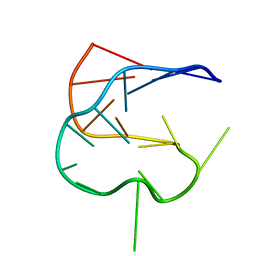 | | SOLUTION NMR STRUCTURE OF AN ANALOG OF THE YEAST TRNA PHE T STEM LOOP CONTAINING RIBOTHYMIDINE AT ITS NATURALLY OCCURRING POSITION | | Descriptor: | 5'-R(*CP*UP*GP*UP*GP*(5MU)P*UP*CP*GP*AP*UP*(CH)P*CP*AP*CP*AP*G)- 3' | | Authors: | Koshlap, K.M, Guenther, R, Sochacka, E, Malkiewicz, A, Agris, P.F. | | Deposit date: | 1999-05-03 | | Release date: | 1999-10-22 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | A distinctive RNA fold: the solution structure of an analogue of the yeast tRNAPhe T Psi C domain.
Biochemistry, 38, 1999
|
|
6MRM
 
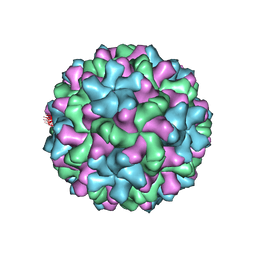 | | Red Clover Necrotic Mosaic Virus | | Descriptor: | CALCIUM ION, Capsid protein | | Authors: | Sherman, M.B, Smith, T.J. | | Deposit date: | 2018-10-14 | | Release date: | 2019-10-16 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Near-Atomic-Resolution Cryo-Electron Microscopy Structures of Cucumber Leaf Spot Virus and Red Clover Necrotic Mosaic Virus: Evolutionary Divergence at the Icosahedral Three-Fold Axes.
J.Virol., 94, 2020
|
|
1FEQ
 
 | | NMR SOLUTION STRUCTURE OF THE ANTICODON OF TRNA(LYS3) WITH T6A MODIFICATION AT POSITION 37 | | Descriptor: | 5'-R(*GP*CP*AP*GP*AP*CP*UP*UP*UP*UP*(T6A)P*AP*UP*CP*UP*GP*C)-3' | | Authors: | Stuart, J.W, Gdaniec, Z, Guenther, R.H, Marszalek, M, Sochacka, E, Malkiewicz, A, Agris, P.F. | | Deposit date: | 2000-07-21 | | Release date: | 2000-11-22 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Functional anticodon architecture of human tRNALys3 includes disruption of intraloop hydrogen bonding by the naturally occurring amino acid modification, t6A.
Biochemistry, 39, 2000
|
|
1LUU
 | |
1LUX
 | |
6MRL
 
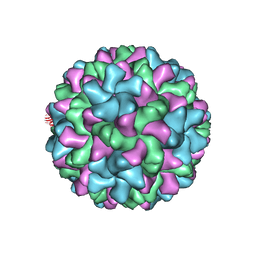 | | Cucumber Leaf Spot Virus | | Descriptor: | CALCIUM ION, p41 | | Authors: | Sherman, M.B, Smith, T.J. | | Deposit date: | 2018-10-14 | | Release date: | 2019-10-16 | | Last modified: | 2024-03-13 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Near-Atomic-Resolution Cryo-Electron Microscopy Structures of Cucumber Leaf Spot Virus and Red Clover Necrotic Mosaic Virus: Evolutionary Divergence at the Icosahedral Three-Fold Axes.
J.Virol., 94, 2020
|
|
