2NEF
 
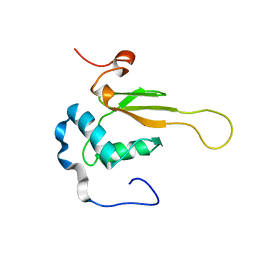 | | HIV-1 NEF (REGULATORY FACTOR), NMR, 40 STRUCTURES | | Descriptor: | NEGATIVE FACTOR (F-PROTEIN) | | Authors: | Grzesiek, S, Bax, A, Clore, G.M, Gronenborn, A.M, Hu, J.S, Kaufman, J, Palmer, I, Stahl, S.J, Tjandra, N, Wingfield, P.T. | | Deposit date: | 1997-02-12 | | Release date: | 1997-07-07 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Refined solution structure and backbone dynamics of HIV-1 Nef.
Protein Sci., 6, 1997
|
|
8AS3
 
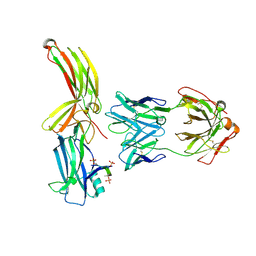 | | Structure of arrestin2 in complex with 6P CCR5 phosphopeptide and Fab30 | | Descriptor: | Beta-arrestin-1, C-C chemokine receptor type 5, Fab30 heavy chain, ... | | Authors: | Isaikina, P, Jakob, R.P, Maier, T, Grzesiek, S. | | Deposit date: | 2022-08-18 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | A key GPCR phosphorylation motif discovered in arrestin2⋅CCR5 phosphopeptide complexes.
Mol.Cell, 83, 2023
|
|
8AS4
 
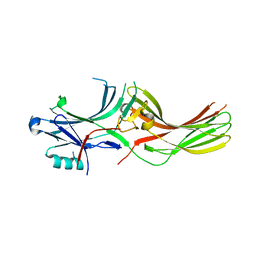 | |
8AS2
 
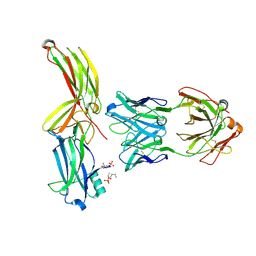 | | Structure of arrestin2 in complex with 4P CCR5 phosphopeptide and Fab30 | | Descriptor: | Beta-arrestin-1, C-C chemokine receptor type 5, Fab30 heavy chain, ... | | Authors: | Isaikina, P, Jakob, R.P, Maier, T, Grzesiek, S. | | Deposit date: | 2022-08-18 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | A key GPCR phosphorylation motif discovered in arrestin2⋅CCR5 phosphopeptide complexes.
Mol.Cell, 83, 2023
|
|
1BJ8
 
 | | THIRD N-TERMINAL DOMAIN OF GP130, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | GP130 | | Authors: | Kernebeck, T, Pflanz, S, Muller-Newen, G, Kurapkat, G, Scheek, R.M, Dijkstra, K, Heinrich, P.C, Wollmer, A, Grzesiek, S, Grotzinger, J. | | Deposit date: | 1998-07-02 | | Release date: | 1999-01-13 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | The signal transducer gp130: solution structure of the carboxy-terminal domain of the cytokine receptor homology region.
Protein Sci., 8, 1999
|
|
1DXW
 
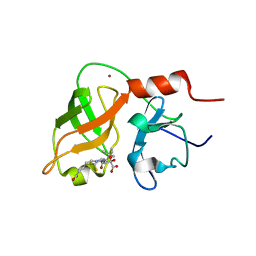 | | structure of hetero complex of non structural protein (NS) of hepatitis C virus (HCV) and synthetic peptidic compound | | Descriptor: | N-(tert-butoxycarbonyl)-L-alpha-glutamyl-N-[(1R)-1-(carboxycarbonyl)-3,3-difluoropropyl]-L-leucinamide, SERINE PROTEASE, ZINC ION | | Authors: | Barbato, G, Cicero, D.O, Cordier, F, Narjes, F, Gerlach, B, Sambucini, S, Grzesiek, S, Matassa, V.G, Defrancesco, R, Bazzo, R. | | Deposit date: | 2000-01-17 | | Release date: | 2001-01-12 | | Last modified: | 2020-01-15 | | Method: | SOLUTION NMR | | Cite: | Inhibitor Binding Induces Active Site Stabilisation of the Hcv Ns3 Protein Serine Protease Domain
Embo J., 19, 2000
|
|
1FOY
 
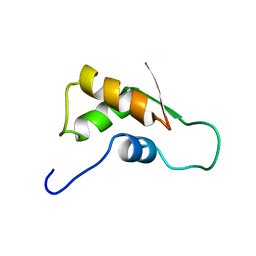 | | THE RNA BINDING DOMAIN OF RIBOSOMAL PROTEIN L11: THREE-DIMENSIONAL STRUCTURE OF THE RNA-BOUND FORM OF THE PROTEIN, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | RIBOSOMAL PROTEIN L11 | | Authors: | Hinck, A.P, Markus, M.A, Huang, S, Grzesiek, S, Kustanovich, I, Draper, D.E, Torchia, D.A. | | Deposit date: | 1997-05-26 | | Release date: | 1997-11-26 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | The RNA binding domain of ribosomal protein L11: three-dimensional structure of the RNA-bound form of the protein and its interaction with 23 S rRNA.
J.Mol.Biol., 274, 1997
|
|
1SP7
 
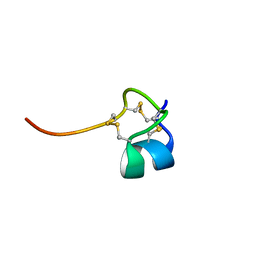 | | Structure of the Cys-rich C-terminal domain of Hydra minicollagen | | Descriptor: | mini-collagen | | Authors: | Meier, S, Haussinger, D, Pokidysheva, E, Bachinger, H.P, Grzesiek, S. | | Deposit date: | 2004-03-16 | | Release date: | 2004-05-18 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Determination of a high-precision NMR structure of the minicollagen cysteine rich domain from Hydra and characterization of its disulfide bond formation.
Febs Lett., 569, 2004
|
|
1NY9
 
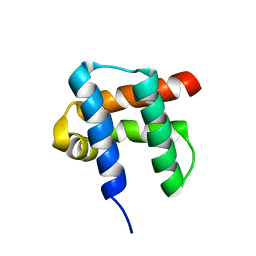 | | Antibiotic binding domain of a TipA-class multidrug resistance transcriptional regulator | | Descriptor: | Transcriptional activator tipA-S | | Authors: | Kahmann, J.D, Sass, H.J, Allan, M.G, Seto, H, Thompson, C.J, Grzesiek, S. | | Deposit date: | 2003-02-12 | | Release date: | 2003-04-15 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structural basis for antibiotic recognition by the TipA-class of
multidrug-resistance transcriptional regulators
Embo J., 22, 2003
|
|
1RFO
 
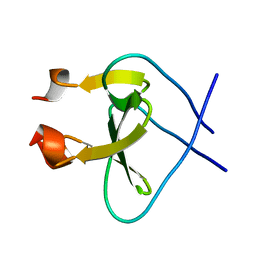 | | Trimeric Foldon of the T4 phagehead fibritin | | Descriptor: | whisker antigen control protein | | Authors: | Guthe, S, Kapinos, L, Moglich, A, Meier, S, Kiefhaber, T, Grzesiek, S. | | Deposit date: | 2003-11-10 | | Release date: | 2004-03-30 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Very fast folding and association of a trimerization domain from bacteriophage t4 fibritin.
J.Mol.Biol., 337, 2004
|
|
1U0P
 
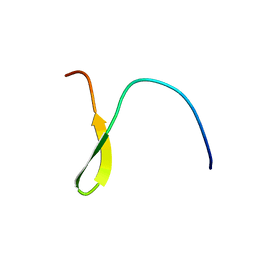 | | Stable A-state hairpin of T4 fibritin foldon | | Descriptor: | fibritin | | Authors: | Meier, S, Guthe, S, Kiefhaber, T, Grzesiek, S. | | Deposit date: | 2004-07-14 | | Release date: | 2005-02-22 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Foldon, the natural trimerization domain of T4 fibritin, dissociates into a monomeric A-state form containing a stable beta-hairpin: atomic details of trimer dissociation and
local beta-hairpin stability from residual dipolar couplings
J.Mol.Biol., 344, 2004
|
|
1CFD
 
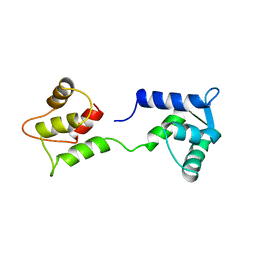 | | CALCIUM-FREE CALMODULIN | | Descriptor: | CALMODULIN | | Authors: | Kuboniwa, H, Tjandra, N, Grzesiek, S, Ren, H, Klee, C.B, Bax, A. | | Deposit date: | 1995-10-18 | | Release date: | 1995-12-07 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of calcium-free calmodulin.
Nat.Struct.Biol., 2, 1995
|
|
1CFC
 
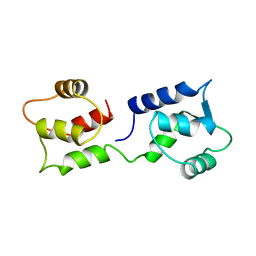 | | CALCIUM-FREE CALMODULIN | | Descriptor: | CALMODULIN | | Authors: | Kuboniwa, H, Tjandra, N, Grzesiek, S, Ren, H, Klee, C.B, Bax, A. | | Deposit date: | 1995-08-02 | | Release date: | 1995-12-07 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of calcium-free calmodulin.
Nat.Struct.Biol., 2, 1995
|
|
7O7F
 
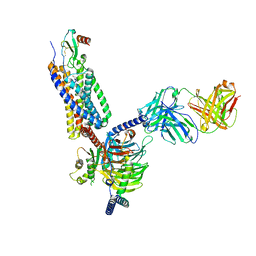 | | Structural basis of the activation of the CC chemokine receptor 5 by a chemokine agonist | | Descriptor: | C-C chemokine receptor type 5, C-C motif chemokine 5, Fab antibody fragment heavy chain, ... | | Authors: | Isaikina, P, Tsai, C.-J, Dietz, N.B, Pamula, F, Goldie, K.N, Schertler, G.F.X, Maier, T, Stahlberg, H, Deupi, X, Grzesiek, S. | | Deposit date: | 2021-04-13 | | Release date: | 2021-06-30 | | Last modified: | 2022-03-16 | | Method: | ELECTRON MICROSCOPY (3.15 Å) | | Cite: | Structural basis of the activation of the CC chemokine receptor 5 by a chemokine agonist.
Sci Adv, 7, 2021
|
|
1H95
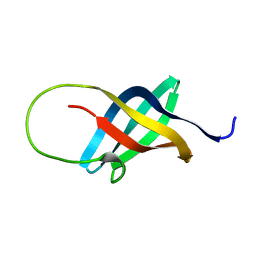 
 | | Solution structure of the single-stranded DNA-binding Cold Shock Domain (CSD) of human Y-box protein 1 (YB1) determined by NMR (10 lowest energy structures) | | Descriptor: | Y-BOX BINDING PROTEIN | | Authors: | Kloks, C.P.A.M, Spronk, C.A.E.M, Hoffmann, A, Vuister, G.W, Grzesiek, S, Hilbers, C.W. | | Deposit date: | 2001-02-23 | | Release date: | 2002-02-21 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | The Solution Structure and DNA-Binding Properties of the Cold-Shock Domain of the Human Y-Box Protein Yb-1.
J.Mol.Biol., 316, 2002
|
|
6SFT
 
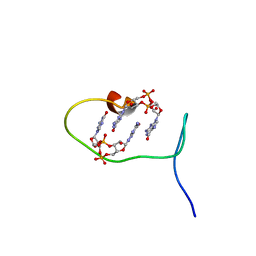 | | Solution structure of protein ARR_CleD in complex with c-di-GMP | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), Two-component receiver protein CleD | | Authors: | Habazettl, J, Hee, C.S, Jenal, U, Schirmer, T, Grzesiek, S. | | Deposit date: | 2019-08-02 | | Release date: | 2020-06-10 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Intercepting second-messenger signaling by rationally designed peptides sequestering c-di-GMP.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
1W1N
 
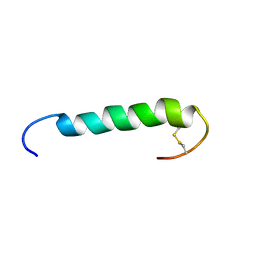 | | The solution structure of the FATC Domain of the Protein Kinase TOR1 from yeast | | Descriptor: | PHOSPHATIDYLINOSITOL 3-KINASE TOR1 | | Authors: | Dames, S.A, Mulet, J.M, Rathgeb-Szabo, K, Hall, M.N, Grzesiek, S. | | Deposit date: | 2004-06-23 | | Release date: | 2005-03-16 | | Last modified: | 2018-05-02 | | Method: | SOLUTION NMR | | Cite: | The solution structure of the FATC domain of the protein kinase target of rapamycin suggests a role for redox-dependent structural and cellular stability.
J. Biol. Chem., 280, 2005
|
|
2HM3
 
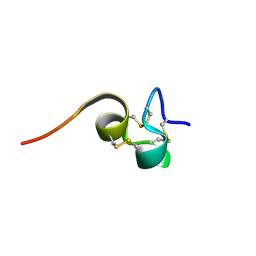 | | Nematocyst outer wall antigen, cysteine rich domain NW1 | | Descriptor: | Nematocyst outer wall antigen | | Authors: | Meier, S, Jensen, P.R, Grzesiek, S, Oezbek, S. | | Deposit date: | 2006-07-11 | | Release date: | 2007-02-06 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Sequence-Structure and Structure-Function Analysis in Cysteine-rich Domains Forming the Ultrastable Nematocyst Wall.
J.Mol.Biol., 368, 2007
|
|
2HM4
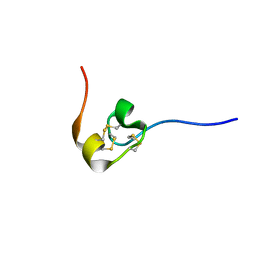 
 | | Nematocyst Outer Wall Antigen, NW1 K21P | | Descriptor: | Nematocyst outer wall antigen | | Authors: | Meier, S, Jensen, P.R, Grzesiek, S, Oezbek, S. | | Deposit date: | 2006-07-11 | | Release date: | 2007-02-06 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Continuous molecular evolution of protein-domain structures by single amino Acid changes.
Curr.Biol., 17, 2007
|
|
2HM6
 
 | | Nematocyst outer wall antigen, NW1 G11V K21P | | Descriptor: | Nematocyst outer wall antigen | | Authors: | Meier, S, Jensen, P.R, Grzesiek, S, Oezbek, S. | | Deposit date: | 2006-07-11 | | Release date: | 2007-02-06 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Continuous molecular evolution of protein-domain structures by single amino Acid changes.
Curr.Biol., 17, 2007
|
|
2HM5
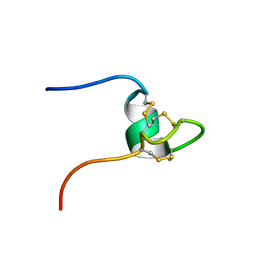 
 | | NW1, K21P, Structural Species II | | Descriptor: | Nematocyst outer wall antigen | | Authors: | Meier, S, Jensen, P.R, Grzesiek, S, Oezbek, S. | | Deposit date: | 2006-07-11 | | Release date: | 2007-02-06 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Continuous molecular evolution of protein-domain structures by single amino Acid changes.
Curr.Biol., 17, 2007
|
|
2V37
 
 | | Solution structure of the N-terminal extracellular domain of human T- cadherin | | Descriptor: | CADHERIN-13 | | Authors: | Dames, S.A, Bang, E.J, Ahrens, T, Haeussinger, D, Grzesiek, S. | | Deposit date: | 2007-06-13 | | Release date: | 2008-06-10 | | Last modified: | 2018-03-28 | | Method: | SOLUTION NMR | | Cite: | Insights into the low adhesive capacity of human T-cadherin from the NMR structure of Its N-terminal extracellular domain.
J. Biol. Chem., 283, 2008
|
|
2FOW
 
 | | THE RNA BINDING DOMAIN OF RIBOSOMAL PROTEIN L11: THREE-DIMENSIONAL STRUCTURE OF THE RNA-BOUND FORM OF THE PROTEIN, NMR, 26 STRUCTURES | | Descriptor: | RIBOSOMAL PROTEIN L11 | | Authors: | Hinck, A.P, Markus, M.A, Huang, S, Grzesiek, S, Kustanovich, I, Draper, D.E, Torchia, D.A. | | Deposit date: | 1997-05-26 | | Release date: | 1997-11-26 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | The RNA binding domain of ribosomal protein L11: three-dimensional structure of the RNA-bound form of the protein and its interaction with 23 S rRNA.
J.Mol.Biol., 274, 1997
|
|
2GLO
 
 | | Solution structure of the Brinker DNA binding domain in complex with the omb enhancer | | Descriptor: | 5'-D(*GP*TP*TP*GP*AP*CP*GP*CP*CP*TP*CP*A)-3', 5'-D(*TP*GP*AP*GP*GP*CP*GP*TP*CP*AP*AP*C)-3', brinker CG9653-PA | | Authors: | Cordier, F, Hartmann, B, Rogowski, M, Affolter, M, Grzesiek, S. | | Deposit date: | 2006-04-05 | | Release date: | 2006-08-29 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | DNA recognition by the brinker repressor - an extreme case of coupling between binding and folding
J.Mol.Biol., 361, 2006
|
|
2OQ9
 
 | |
