2RR3
 
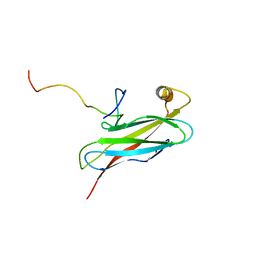 | | Solution structure of the complex between human VAP-A MSP domain and human OSBP FFAT motif | | Descriptor: | Oxysterol-binding protein 1, Vesicle-associated membrane protein-associated protein A | | Authors: | Furuita, K, Jee, J, Fukada, H, Mishima, M, Kojima, C. | | Deposit date: | 2010-03-09 | | Release date: | 2010-03-23 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Electrostatic interaction between oxysterol-binding protein and VAMP-associated protein A revealed by NMR and mutagenesis studies
J.Biol.Chem., 285, 2010
|
|
1VBF
 
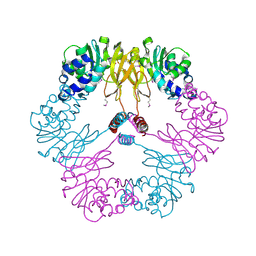 | | Crystal structure of protein L-isoaspartate O-methyltransferase homologue from Sulfolobus tokodaii | | Descriptor: | 231aa long hypothetical protein-L-isoaspartate O-methyltransferase | | Authors: | Tanaka, Y, Tsumoto, K, Yasutake, Y, Umetsu, M, Yao, M, Tanaka, I, Fukada, H, Kumagai, I. | | Deposit date: | 2004-02-25 | | Release date: | 2004-08-10 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | How Oligomerization Contributes to the Thermostability of an Archaeon Protein: PROTEIN L-ISOASPARTYL-O-METHYLTRANSFERASE FROM SULFOLOBUS TOKODAII
J.Biol.Chem., 279, 2004
|
|
5Y90
 
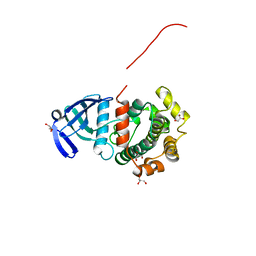 | | MAP2K7 mutant -C218S | | Descriptor: | Dual specificity mitogen-activated protein kinase kinase 7, GLYCEROL | | Authors: | Kinoshita, T, Hashimoto, T, Sogabe, Y, Matsumoto, T, Sawa, M, Fukada, H. | | Deposit date: | 2017-08-22 | | Release date: | 2017-10-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | High-resolution structure discloses the potential for allosteric regulation of mitogen-activated protein kinase kinase 7
Biochem. Biophys. Res. Commun., 493, 2017
|
|
4TV7
 
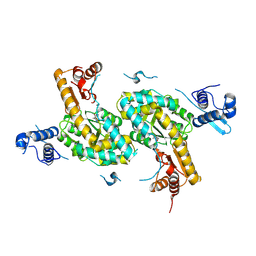 | |
5Y52
 
 | |
5Y2P
 
 | |
5HXV
 
 | |
4XFP
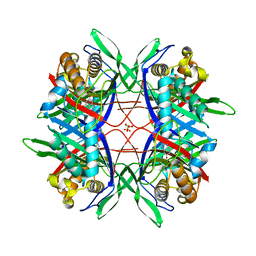 
 | | Crystal Structure of Highly Active Mutant of Bacillus sp. TB-90 Urate Oxidase | | Descriptor: | 8-AZAXANTHINE, CHLORIDE ION, SULFATE ION, ... | | Authors: | Hibi, T, Hayashi, Y, Kawamura, A, Itoh, T. | | Deposit date: | 2014-12-28 | | Release date: | 2016-01-13 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Glycine Substitution of Surface Proline 287 Involves Entropic Enhancement of Bacillus sp. TB-90 Uricase Activity
To be published
|
|
5AYJ
 
 | | Hyperthermostable mutant of Bacillus sp. TB-90 Urate Oxidase - R298C | | Descriptor: | 9-METHYL URIC ACID, HEXAETHYLENE GLYCOL, SULFATE ION, ... | | Authors: | Hibi, T, Kume, A, Kawamura, A, Itoh, T, Nishiya, Y. | | Deposit date: | 2015-08-21 | | Release date: | 2016-01-20 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Hyperstabilization of Tetrameric Bacillus sp. TB-90 Urate Oxidase by Introducing Disulfide Bonds through Structural Plasticity
Biochemistry, 55, 2016
|
|
7V92
 
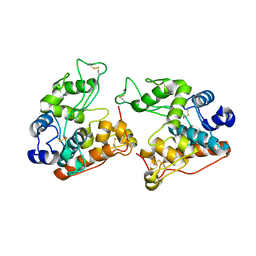 | |
7V91
 
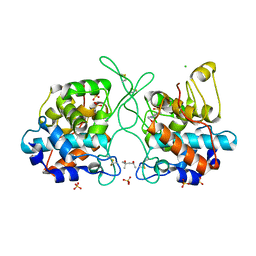 | | Crystal Structure of the Catalytic Domain of a Family GH19 Chitinase from Gazyumaru, Ficus microcarpa | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CHLORIDE ION, ... | | Authors: | Kozome, D, Kubota, T, Ishikawa, K. | | Deposit date: | 2021-08-24 | | Release date: | 2022-04-27 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural Analysis and Construction of a Thermostable Antifungal Chitinase.
Appl.Environ.Microbiol., 88, 2022
|
|
5B5S
 
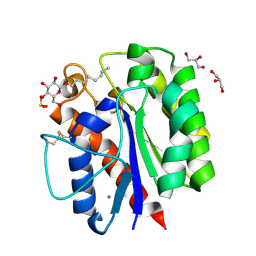 | |
2LAA
 
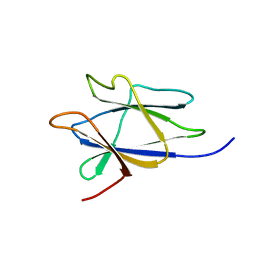 | |
3AT4
 
 | | Crystal structure of CK2alpha with pyradine derivertive | | Descriptor: | Casein kinase II subunit alpha, [1-(6-{6-[(1-methylethyl)amino]-1H-indazol-1-yl}pyrazin-2-yl)-1H-pyrrol-3-yl]acetic acid | | Authors: | Kinoshita, T. | | Deposit date: | 2010-12-23 | | Release date: | 2011-11-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A detailed thermodynamic profile of cyclopentyl and isopropyl derivatives binding to CK2 kinase
Mol.Cell.Biochem., 356, 2011
|
|
3AT3
 
 | | Crystal structure of CK2alpha with pyradine derivative | | Descriptor: | (1-{6-[6-(cyclopentylamino)-1H-indazol-1-yl]pyrazin-2-yl}-1H-pyrrol-3-yl)acetic acid, Casein kinase II subunit alpha | | Authors: | Kinoshita, T. | | Deposit date: | 2010-12-23 | | Release date: | 2011-11-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A detailed thermodynamic profile of cyclopentyl and isopropyl derivatives binding to CK2 kinase
Mol.Cell.Biochem., 356, 2011
|
|
3AT2
 
 | | Crystal structure of CK2alpha | | Descriptor: | 1,2-ETHANEDIOL, Casein kinase II subunit alpha | | Authors: | Kinoshita, T. | | Deposit date: | 2010-12-23 | | Release date: | 2011-11-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | A detailed thermodynamic profile of cyclopentyl and isopropyl derivatives binding to CK2 kinase
Mol.Cell.Biochem., 356, 2011
|
|
2LAB
 
 | |
5YJA
 
 | |
5Z2B
 
 | |
5Z27
 
 | |
5YJ2
 
 | |
5Y8U
 
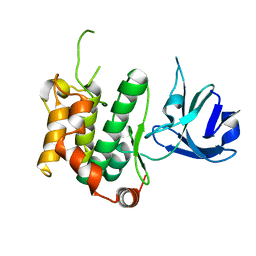 | | Crystal structure of the C276S mutant of MAP2K7 | | Descriptor: | Dual specificity mitogen-activated protein kinase kinase 7 | | Authors: | Kinoshita, T. | | Deposit date: | 2017-08-21 | | Release date: | 2017-10-11 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.92 Å) | | Cite: | High-resolution structure discloses the potential for allosteric regulation of mitogen-activated protein kinase kinase 7
Biochem. Biophys. Res. Commun., 493, 2017
|
|
3AKO
 
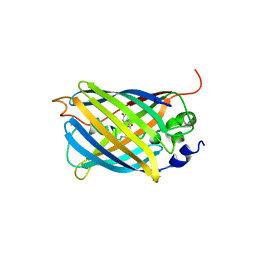 | | Crystal Structure of the Reassembled Venus | | Descriptor: | 3,6,9,12,15,18,21-HEPTAOXATRICOSANE-1,23-DIOL, SULFATE ION, Venus | | Authors: | Isogai, M, Tada, T. | | Deposit date: | 2010-07-15 | | Release date: | 2011-08-03 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure and characteristics of reassembled fluorescent protein, a new insight into the reassembly mechanisms
Bioorg.Med.Chem.Lett., 21, 2011
|
|
7CMQ
 
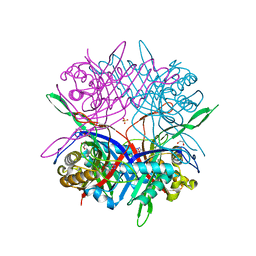 | |
7CUF
 
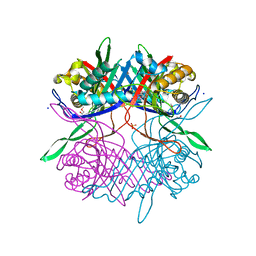 | |
