2XOW
 | | Structure of GlpG in complex with a mechanism-based isocoumarin inhibitor | | Descriptor: | 5-AMINO-2-(2-METHOXY-2-OXOETHYL)BENZOIC ACID, RHOMBOID PROTEASE GLPG, nonyl beta-D-glucopyranoside | | Authors: | Vinothkumar, K.R, Strisovsky, K, Andreeva, A, Christova, Y, Verhelst, S, Freeman, M. | | Deposit date: | 2010-08-24 | | Release date: | 2010-10-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | The Structural Basis for Catalysis and Substrate Specificity of a Rhomboid Protease
Embo J., 29, 2010
|
|
2XOV
 
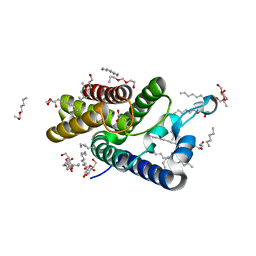 | | Crystal Structure of E.coli rhomboid protease GlpG, native enzyme | | Descriptor: | GLYCEROL, RHOMBOID PROTEASE GLPG, nonyl beta-D-glucopyranoside | | Authors: | Vinothkumar, K.R, Strisovsky, K, Andreeva, A, Christova, Y, Verhelst, S, Freeman, M. | | Deposit date: | 2010-08-24 | | Release date: | 2010-10-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | The structural basis for catalysis and substrate specificity of a rhomboid protease.
EMBO J., 29, 2010
|
|
3ZOT
 
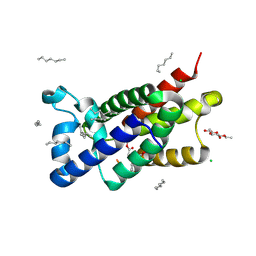 | | Structure of E.coli rhomboid protease GlpG in complex with monobactam L29 (data set 2) | | Descriptor: | CHLORIDE ION, RHOMBOID PROTEASE GLPG, nonyl beta-D-glucopyranoside, ... | | Authors: | Vinothkumar, K.R, Pierrat, O.A, Large, J.M, Freeman, M. | | Deposit date: | 2013-02-24 | | Release date: | 2013-05-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.399 Å) | | Cite: | Structure of Rhomboid Protease in Complex with Beta-Lactam Inhibitors Defines the S2' Cavity.
Structure, 21, 2013
|
|
3ZMH
 | | Structure of E.coli rhomboid protease GlpG in complex with monobactam L62 | | Descriptor: | CHLORIDE ION, CYCLOPENTYL 2-OXO-4-PHENYLAZETIDINE-1-CARBOXYLATE, RHOMBOID PROTEASE GLPG, ... | | Authors: | Vinothkumar, K.R, Pierrat, O.A, Large, J.M, Freeman, M. | | Deposit date: | 2013-02-11 | | Release date: | 2013-05-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of rhomboid protease in complex with beta-lactam inhibitors defines the S2' cavity.
Structure, 21, 2013
|
|
3ZMJ
 | | Structure of E.coli rhomboid protease GlpG in complex with monobactam L61 | | Descriptor: | 2-methylpropyl N-[(1R)-3-oxidanylidene-1-phenyl-propyl]carbamate, CHLORIDE ION, RHOMBOID PROTEASE GLPG, ... | | Authors: | Vinothkumar, K.R, Pierrat, O, Large, J.M, Freeman, M. | | Deposit date: | 2013-02-11 | | Release date: | 2013-05-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of Rhomboid Protease in Complex with Beta-Lactam Inhibitors Defines the S2' Cavity.
Structure, 21, 2013
|
|
3ZMI
 
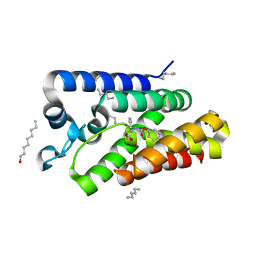 | | Structure of E.coli rhomboid protease GlpG in complex with monobactam L29 | | Descriptor: | RHOMBOID PROTEASE GLPG, nonyl beta-D-glucopyranoside, phenyl N-[(1R)-3-oxidanylidene-1-phenyl-propyl]carbamate | | Authors: | Vinothkumar, K.R, Pierrat, O.A, Large, J.M, Freeman, M. | | Deposit date: | 2013-02-11 | | Release date: | 2013-05-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of Rhomboid Protease in Complex with Beta-Lactam Inhibitors Defines the S2' Cavity.
Structure, 21, 2013
|
|
3KQ5
 
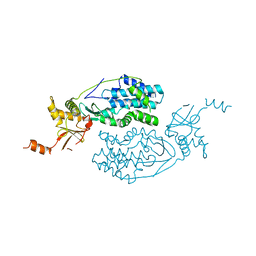 | | Crystal structure of an uncharacterized protein from Coxiella burnetii | | Descriptor: | Hypothetical cytosolic protein | | Authors: | Bonanno, J.B, Freeman, M, Bain, K.T, Chang, S, Ozyurt, S, Wasserman, S, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-11-17 | | Release date: | 2009-12-08 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of an uncharacterized protein from Coxiella burnetii
To be Published
|
|
6Y5M
 
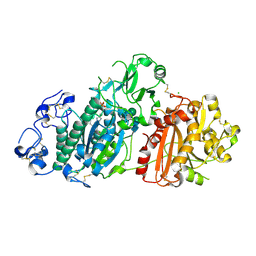 | | Crystal structure of mouse Autotaxin in complex with compound 1a | | Descriptor: | (~{E})-3-[4-chloranyl-2-[(5-methyl-1,2,3,4-tetrazol-2-yl)methyl]phenyl]-1-[(2~{R})-4-[(4-fluorophenyl)methyl]-2-methyl-piperazin-1-yl]prop-2-en-1-one, 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | Authors: | Faller, M, Zink, F. | | Deposit date: | 2020-02-25 | | Release date: | 2020-11-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.011 Å) | | Cite: | Development of autotaxin inhibitors: A series of tetrazole cinnamides.
Bioorg.Med.Chem.Lett., 31, 2021
|
|
7OEX
 
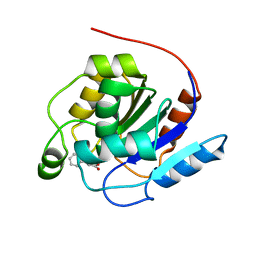 | |
4ZPV
 
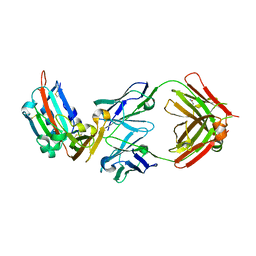 | | Structure of MERS-Coronavirus Spike Receptor-binding Domain (England1 Strain) in Complex with Vaccine-Elicited Murine Neutralizing Antibody D12 (Crystal Form 2) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, D12 Fab Heavy chain, D12 Fab light chain, ... | | Authors: | Joyce, M.G, Mascola, J.R, Graham, B.S, Kwong, P.D. | | Deposit date: | 2015-05-08 | | Release date: | 2015-10-21 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Evaluation of candidate vaccine approaches for MERS-CoV.
Nat Commun, 6, 2015
|
|
4ZPW
 
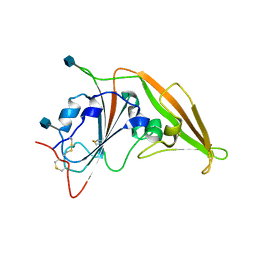 | | Structure of unbound MERS-CoV spike receptor-binding domain (England1 strain). | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, PHOSPHATE ION, Spike glycoprotein | | Authors: | Joyce, M.G, Mascola, J.R, Graham, B.S, Kwong, P.D. | | Deposit date: | 2015-05-08 | | Release date: | 2015-08-12 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.023 Å) | | Cite: | Evaluation of candidate vaccine approaches for MERS-CoV.
Nat Commun, 6, 2015
|
|
2I1Y
 
 | | Crystal structure of the phosphatase domain of human PTP IA-2 | | Descriptor: | GLYCEROL, Receptor-type tyrosine-protein phosphatase | | Authors: | Faber-Barata, J, Patskovsky, Y, Alvarado, J, Smith, D, Koss, J, Wasserman, S.R, Ozyurt, S, Atwell, S, Powell, A, Kearins, M.C, Maletic, M, Rooney, I, Bain, K.T, Freeman, M, Russell, J.C, Thompson, D.A, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2006-08-15 | | Release date: | 2006-08-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Structural genomics of protein phosphatases
J.STRUCT.FUNCT.GENOM., 8, 2007
|
|
