1II6
 | | Crystal Structure of the Mitotic Kinesin Eg5 in Complex with Mg-ADP. | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, KINESIN-RELATED MOTOR PROTEIN Eg5, MAGNESIUM ION, ... | | Authors: | Turner, J, Anderson, R, Guo, J, Beraud, C, Sakowicz, R, Fletterick, R. | | Deposit date: | 2001-04-20 | | Release date: | 2001-07-18 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of the mitotic spindle kinesin Eg5 reveals a novel conformation of the neck-linker.
J.Biol.Chem., 276, 2001
|
|
1RY6
 
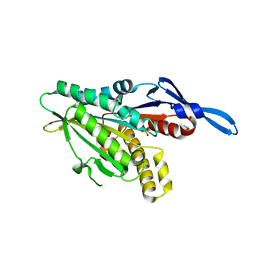 | | Crystal Structure of Internal Kinesin Motor Domain | | Descriptor: | INTERNAL KINESIN, SULFATE ION | | Authors: | Shipley, K, Hekmat-Nejad, M, Turner, J, Moores, C, Anderson, R, Milligan, R, Sakowicz, R, Fletterick, R. | | Deposit date: | 2003-12-19 | | Release date: | 2004-04-13 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure of a kinesin microtubule depolymerization machine.
Embo J., 23, 2004
|
|
1MKJ
 
 | | Human Kinesin Motor Domain With Docked Neck Linker | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, KINESIN HEAVY CHAIN, MAGNESIUM ION, ... | | Authors: | Sindelar, C.V, Budny, M.J, Rice, S, Naber, N, Fletterick, R, Cooke, R. | | Deposit date: | 2002-08-29 | | Release date: | 2002-10-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Two conformations in the human kinesin power stroke defined by X-ray crystallography and EPR spectroscopy.
Nat.Struct.Biol., 9, 2002
|
|
1AIM
 
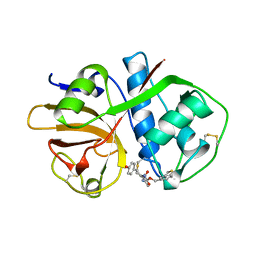 | |
1PYG
 
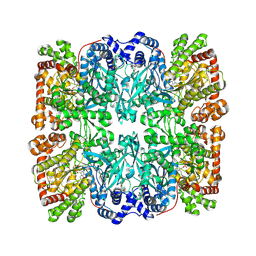 | |
3TX7
 
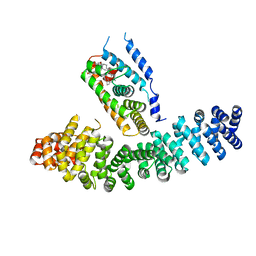 | | Crystal structure of LRH-1/beta-catenin complex | | Descriptor: | (2S)-3-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-2-[(6E)-HEXADEC-6-ENOYLOXY]PROPYL (8E)-OCTADEC-8-ENOATE, Catenin beta-1, Nuclear receptor subfamily 5 group A member 2 | | Authors: | Yumoto, F, Fletterick, R. | | Deposit date: | 2011-09-22 | | Release date: | 2011-12-14 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.76 Å) | | Cite: | Structural basis of coactivation of liver receptor homolog-1 by beta-catenin.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
1LOX
 
 | | RABBIT RETICULOCYTE 15-LIPOXYGENASE | | Descriptor: | (2E)-3-(2-OCT-1-YN-1-YLPHENYL)ACRYLIC ACID, 15-LIPOXYGENASE, FE (II) ION | | Authors: | Gillmor, S.A, Villasenor, A, Fletterick, R.J, Sigal, E, Browner, M.F. | | Deposit date: | 1997-10-06 | | Release date: | 1998-11-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The structure of mammalian 15-lipoxygenase reveals similarity to the lipases and the determinants of substrate specificity.
Nat.Struct.Biol., 4, 1997
|
|
3BKI
 
 | | Crystal Structure of the GluR2 ligand binding core (S1S2J) in complex with FQX at 1.87 Angstroms | | Descriptor: | Glutamate receptor 2, [1,2,5]oxadiazolo[3,4-g]quinoxaline-6,7(5H,8H)-dione 1-oxide | | Authors: | Cruz, L, Estebanez-Perpina, E, Pfaff, S, Borngraeber, S, Bao, N, Fletterick, R, England, P. | | Deposit date: | 2007-12-06 | | Release date: | 2008-09-16 | | Last modified: | 2019-09-04 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | 6-Azido-7-nitro-1,4-dihydroquinoxaline-2,3-dione (ANQX) forms an irreversible bond to the active site of the GluR2 AMPA receptor.
J.Med.Chem., 51, 2008
|
|
2QPY
 
 | | AR LBD with small molecule | | Descriptor: | 5-ALPHA-DIHYDROTESTOSTERONE, Androgen receptor, Coactivator peptide, ... | | Authors: | Estebanez-Perpina, E, Fletterick, R. | | Deposit date: | 2007-07-25 | | Release date: | 2007-09-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A surface on the androgen receptor that allosterically regulates coactivator binding.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
1FI8
 | | RAT GRANZYME B [N66Q] COMPLEXED TO ECOTIN [81-84 IEPD] | | Descriptor: | ECOTIN, NATURAL KILLER CELL PROTEASE 1 | | Authors: | Waugh, S.M, Harris, J.L, Fletterick, R.J, Craik, C.S. | | Deposit date: | 2000-08-03 | | Release date: | 2000-09-13 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The structure of the pro-apoptotic protease granzyme B reveals the molecular determinants of its specificity
Nat.Struct.Biol., 7, 2000
|
|
2P0M
 
 | | Revised structure of rabbit reticulocyte 15S-lipoxygenase | | Descriptor: | (2E)-3-(2-OCT-1-YN-1-YLPHENYL)ACRYLIC ACID, Arachidonate 15-lipoxygenase, FE (II) ION | | Authors: | Choi, J, Chon, J.K, Kim, S, Shin, W. | | Deposit date: | 2007-02-28 | | Release date: | 2007-10-09 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Conformational flexibility in mammalian 15S-lipoxygenase: Reinterpretation of the crystallographic data.
Proteins, 70, 2008
|
|
