8SUZ
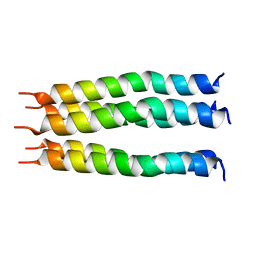 
 | | Open State of the SARS-CoV-2 Envelope Protein Transmembrane Domain, Determined by Solid-State NMR | | Descriptor: | Envelope small membrane protein | | Authors: | Medeiros-Silva, J, Dregni, A.J, Somberg, N.H, Hong, M. | | Deposit date: | 2023-05-14 | | Release date: | 2023-10-25 | | Method: | SOLID-STATE NMR | | Cite: | Atomic structure of the open SARS-CoV-2 E viroporin.
Sci Adv, 9, 2023
|
|
7SFQ
 
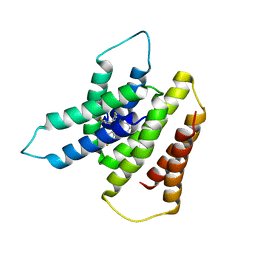 | | EmrE S64V Mutant Bound to tetra(4-fluorophenyl)phosphonium at pH 8.0 | | Descriptor: | Multidrug transporter EmrE, tetrakis(4-fluorophenyl)phosphanium | | Authors: | Shcherbakov, A.A, Spreacker, P.J, Dregni, A.J, Henzler-Wildman, K.A, Hong, M. | | Deposit date: | 2021-10-04 | | Release date: | 2022-03-02 | | Method: | SOLID-STATE NMR | | Cite: | High-pH structure of EmrE reveals the mechanism of proton-coupled substrate transport.
Nat Commun, 13, 2022
|
|
6NZN
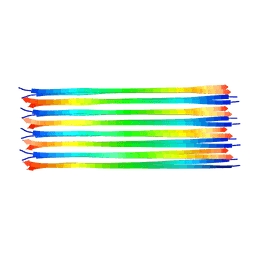 
 | | Dimer-of-dimer amyloid fibril structure of glucagon | | Descriptor: | Glucagon | | Authors: | Gelenter, M.D, Smith, K.J, Liao, S.Y, Mandala, V.S, Dregni, A.J, Lamm, M.S, Tian, Y, Wei, X, Pochan, D.J, Tucker, T.J, Su, Y, Hong, M. | | Deposit date: | 2019-02-14 | | Release date: | 2019-06-05 | | Last modified: | 2023-06-14 | | Method: | SOLID-STATE NMR | | Cite: | The peptide hormone glucagon forms amyloid fibrils with two coexisting beta-strand conformations.
Nat.Struct.Mol.Biol., 26, 2019
|
|
7K3G
 
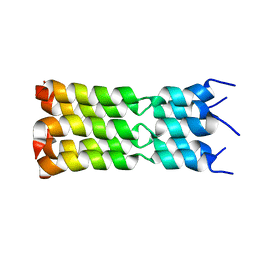 | | SARS-CoV-2 Envelope Protein Transmembrane Domain: Pentameric Structure Determined by Solid-State NMR | | Descriptor: | Envelope small membrane protein | | Authors: | Mandala, V.S, Hong, M, McKay, M.J, Shcherbakov, A.S, Dregni, A.J. | | Deposit date: | 2020-09-11 | | Release date: | 2020-09-30 | | Last modified: | 2023-06-14 | | Method: | SOLID-STATE NMR | | Cite: | Structure and drug binding of the SARS-CoV-2 envelope protein transmembrane domain in lipid bilayers.
Nat.Struct.Mol.Biol., 27, 2020
|
|
8G55
 | |
8G54
 | |
8G58
 | |
