4ZO1
 
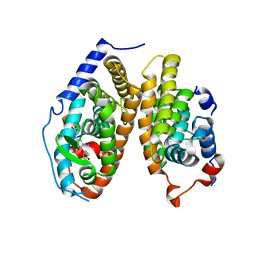 | | Crystal Structure of the T3-bound TR-beta Ligand-binding Domain in complex with RXR-alpha | | Descriptor: | 3,5,3'TRIIODOTHYRONINE, Nuclear receptor coactivator 2, Retinoic acid receptor RXR-alpha, ... | | Authors: | Bruning, J.B, Kojetin, D.J, Matta-Camacho, E, Hughes, T.S, Srinivasan, S, Nwachukwu, J.C, Cavett, V, Nowak, J, Chalmers, M.J, Marciano, D.P, Kamenecka, T.M, Rance, M, Shulman, A.I, Mangelsdorf, D.J, Griffin, P.R, Nettles, K.W. | | Deposit date: | 2015-05-05 | | Release date: | 2015-09-02 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.221 Å) | | Cite: | Structural mechanism for signal transduction in RXR nuclear receptor heterodimers.
Nat Commun, 6, 2015
|
|
2XSE
 
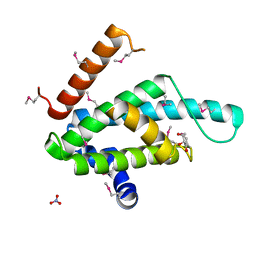 | | The structural basis for recognition of J-base containing DNA by a novel DNA-binding domain in JBP1 | | Descriptor: | GLYCEROL, NITRATE ION, THYMINE DIOXYGENASE JBP1 | | Authors: | Heidebrecht, T, Christodoulou, E, Chalmers, M.J, Jan, S, ter Riete, B, Grover, R.K, Joosten, R.P, Littler, D, vanLuenen, H, Griffin, P.R, Wentworth, P, Borst, P, Perrakis, A. | | Deposit date: | 2010-09-28 | | Release date: | 2011-03-30 | | Last modified: | 2011-08-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Structural Basis for Recognition of Base J Containing DNA by a Novel DNA Binding Domain in Jbp1.
Nucleic Acids Res., 39, 2011
|
|
5VB9
 
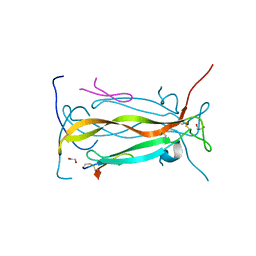 | | IL-17A in complex with peptide | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Interleukin-17A, ... | | Authors: | Antonysamy, S, Russell, M, Zhang, A, Groshong, C, Manglicmot, D, Lu, F, Benach, J, Wasserman, S.R, Zhang, F, Afshar, S, Bina, H, Broughton, H, Chalmers, M, Dodge, J, Espada, A, Jones, S, Ting, J.P, Woodman, M. | | Deposit date: | 2017-03-28 | | Release date: | 2018-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Utilization of peptide phage display to investigate hotspots on IL-17A and what it means for drug discovery.
PLoS ONE, 13, 2018
|
|
4K4J
 
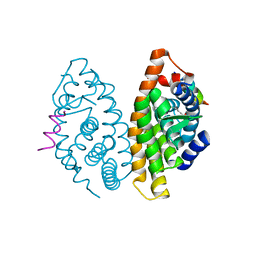 | | Crystal structure of human Retinoid X Receptor alpha-ligand binding domain complex with 9cUAB30 and the coactivator peptide GRIP-1 | | Descriptor: | (2E,4E,6Z,8E)-8-(3,4-dihydronaphthalen-1(2H)-ylidene)-3,7-dimethylocta-2,4,6-trienoic acid, Nuclear receptor coactivator 2 peptide, Retinoic acid receptor RXR-alpha | | Authors: | Xia, G, Smith, C.D, Muccio, D.D. | | Deposit date: | 2013-04-12 | | Release date: | 2013-11-13 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Defining the Communication between Agonist and Coactivator Binding in the Retinoid X Receptor alpha Ligand Binding Domain.
J.Biol.Chem., 289, 2014
|
|
4K6I
 
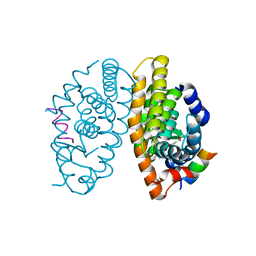 | | Crystal structure of human retinoid X receptor alpha-ligand binding domain complex with Targretin and the coactivator peptide GRIP-1 | | Descriptor: | 4-[1-(3,5,5,8,8-pentamethyl-5,6,7,8-tetrahydronaphthalen-2-yl)ethenyl]benzoic acid, Nuclear receptor coactivator 2, Retinoic acid receptor RXR-alpha | | Authors: | Xia, G, Smith, C.D, Muccio, D.D. | | Deposit date: | 2013-04-15 | | Release date: | 2013-11-13 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Defining the Communication between Agonist and Coactivator Binding in the Retinoid X Receptor alpha Ligand Binding Domain.
J.Biol.Chem., 289, 2014
|
|
5V39
 
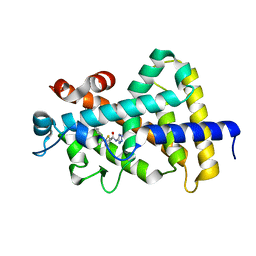 | |
3GZ0
 
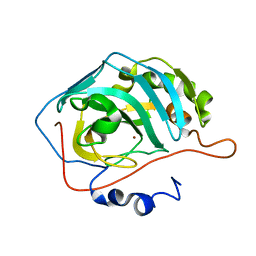 | |
2Q6S
 
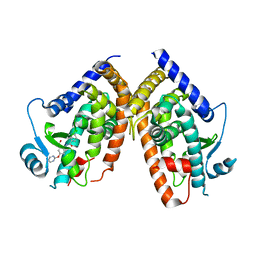 | |
2Q5P
 
 | | Crystal Structure of PPARgamma bound to partial agonist MRL24 | | Descriptor: | (2S)-2-(3-{[1-(4-METHOXYBENZOYL)-2-METHYL-5-(TRIFLUOROMETHOXY)-1H-INDOL-3-YL]METHYL}PHENOXY)PROPANOIC ACID, Peroxisome Proliferator-Activated Receptor gamma | | Authors: | Bruning, J.B, Nettles, K.W. | | Deposit date: | 2007-06-01 | | Release date: | 2007-10-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Partial Agonists Activate PPARgamma Using a Helix 12 Independent Mechanism
Structure, 15, 2007
|
|
2Q59
 
 | | Crystal Structure of PPARgamma LBD bound to full agonist MRL20 | | Descriptor: | (2S)-2-(2-{[1-(4-METHOXYBENZOYL)-2-METHYL-5-(TRIFLUOROMETHOXY)-1H-INDOL-3-YL]METHYL}PHENOXY)PROPANOIC ACID, Peroxisome Proliferator-Activated Receptor gamma | | Authors: | Bruning, J.B, Nettles, K.W. | | Deposit date: | 2007-05-31 | | Release date: | 2007-10-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Partial Agonists Activate PPARgamma Using a Helix 12 Independent Mechanism
Structure, 15, 2007
|
|
2Q6R
 
 | |
2Q5S
 
 | |
2Q61
 
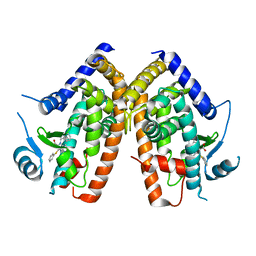 | |
2R6W
 
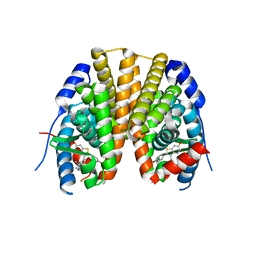 | | Estrogen receptor alpha ligand-binding domain complexed to a SERM | | Descriptor: | Estrogen receptor, [6-hydroxy-2-(4-hydroxyphenyl)-1-benzothien-3-yl]{4-[2-(4-methylpiperidin-1-yl)ethoxy]phenyl}methanone | | Authors: | Wang, Y. | | Deposit date: | 2007-09-06 | | Release date: | 2008-04-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Prediction of the tissue-specificity of selective estrogen receptor modulators by using a single biochemical method.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
2R6Y
 
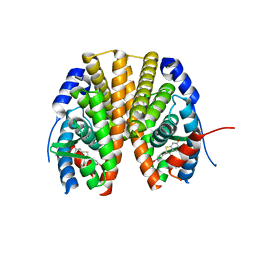 | | Estrogen receptor alpha ligand-binding domain in complex with a SERM | | Descriptor: | Estrogen receptor, [6-hydroxy-2-(4-hydroxyphenyl)-1-benzothien-3-yl][4-(2-pyrrolidin-1-ylethoxy)phenyl]methanone | | Authors: | Wang, Y. | | Deposit date: | 2007-09-06 | | Release date: | 2008-04-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Prediction of the tissue-specificity of selective estrogen receptor modulators by using a single biochemical method.
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
3D6D
 
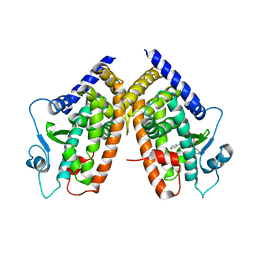 | | Crystal Structure of the complex between PPARgamma LBD and the LT175(R-enantiomer) | | Descriptor: | (2S)-2-(biphenyl-4-yloxy)-3-phenylpropanoic acid, Peroxisome proliferator-activated receptor gamma | | Authors: | Pochetti, G, Montanari, R. | | Deposit date: | 2008-05-19 | | Release date: | 2008-12-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of the Peroxisome Proliferator-Activated Receptor gamma (PPARgamma) Ligand Binding Domain Complexed with a Novel Partial Agonist: A New Region of the Hydrophobic Pocket Could Be Exploited for Drug Design
J.Med.Chem., 51, 2008
|
|
3U9Q
 
 | |
3UC4
 
 | | The crystal structure of Snf1-related kinase 2.6 | | Descriptor: | Serine/threonine-protein kinase SRK2E | | Authors: | Zhou, X.E, Ng, L.-M, Soon, F.-F, Kovach, A, Suino-Powell, K.M, Li, J, Melcher, K, Xu, H.E. | | Deposit date: | 2011-10-26 | | Release date: | 2011-12-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis for basal activity and autoactivation of abscisic acid (ABA) signaling SnRK2 kinases.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3UC3
 
 | | The crystal structure of Snf1-related kinase 2.3 | | Descriptor: | COBALT (II) ION, Serine/threonine-protein kinase SRK2I | | Authors: | Zhou, X.E, Ng, L.-M, Soon, F.-F, Kovach, A, Suino-Powell, K.M, Li, J, Melcher, K, Xu, H.E. | | Deposit date: | 2011-10-25 | | Release date: | 2011-12-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis for basal activity and autoactivation of abscisic acid (ABA) signaling SnRK2 kinases.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3UJG
 
 | | Crystal structure of SnRK2.6 in complex with HAB1 | | Descriptor: | MAGNESIUM ION, Protein phosphatase 2C 16, SULFATE ION, ... | | Authors: | Zhou, X.E, Soon, F.-F, Ng, L.-M, Kovach, A, Tan, M.H.E, Suino-Powell, K.M, He, Y, Xu, Y, Brunzelle, J.S, Li, J, Melcher, K, Xu, H.E. | | Deposit date: | 2011-11-07 | | Release date: | 2012-02-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Molecular mimicry regulates ABA signaling by SnRK2 kinases and PP2C phosphatases.
Science, 335, 2012
|
|
3UJL
 
 | | Crystal structure of abscisic acid bound PYL2 in complex with type 2C protein phosphatase ABI2 | | Descriptor: | (2Z,4E)-5-[(1S)-1-hydroxy-2,6,6-trimethyl-4-oxocyclohex-2-en-1-yl]-3-methylpenta-2,4-dienoic acid, Abscisic acid receptor PYL2, MAGNESIUM ION, ... | | Authors: | Zhou, X.E, Soon, F.-F, Ng, L.-M, Kovach, A, Tan, M.H.E, Suino-Powell, K.M, He, Y, Xu, Y, Brunzelle, J.S, Li, J, Melcher, K, Xu, H.E. | | Deposit date: | 2011-11-07 | | Release date: | 2012-02-15 | | Last modified: | 2012-03-28 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Molecular mimicry regulates ABA signaling by SnRK2 kinases and PP2C phosphatases.
Science, 335, 2012
|
|
3UJK
 
 | | Crystal structure of protein phosphatase ABI2 | | Descriptor: | MAGNESIUM ION, Protein phosphatase 2C 77 | | Authors: | Zhou, X.E, Soon, F.-F, Ng, L.-M, Kovach, A, Tan, M.H.E, Suino-Powell, K.M, He, Y, Xu, Y, Brunzelle, J.S, Li, J, Melcher, K, Xu, H.E. | | Deposit date: | 2011-11-07 | | Release date: | 2012-02-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular mimicry regulates ABA signaling by SnRK2 kinases and PP2C phosphatases.
Science, 335, 2012
|
|
