1CCN
 
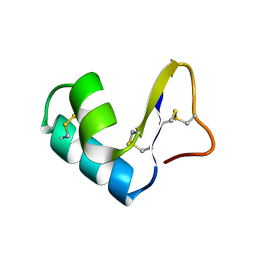 | | DIRECT NOE REFINEMENT OF CRAMBIN FROM 2D NMR DATA USING A SLOW-COOLING ANNEALING PROTOCOL | | Descriptor: | CRAMBIN | | Authors: | Bonvin, A.M.J.J, Rullmann, J.A.C, Lamerichs, R.M.J.N, Boelens, R, Kaptein, R. | | Deposit date: | 1993-04-14 | | Release date: | 1993-10-31 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Direct NOE refinement of biomolecular structures using 2D NMR data
J.Biomol.NMR, 1, 1991
|
|
1CCM
 
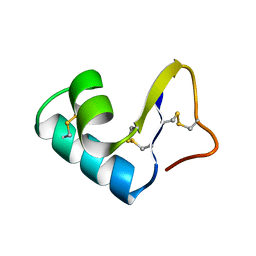 | | DIRECT NOE REFINEMENT OF CRAMBIN FROM 2D NMR DATA USING A SLOW-COOLING ANNEALING PROTOCOL | | Descriptor: | CRAMBIN | | Authors: | Bonvin, A.M.J.J, Rullmann, J.A.C, Lamerichs, R.M.J.N, Boelens, R, Kaptein, R. | | Deposit date: | 1993-04-14 | | Release date: | 1993-10-31 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | "Ensemble" iterative relaxation matrix approach: a new NMR refinement protocol applied to the solution structure of crambin.
Proteins, 15, 1993
|
|
2XY8
 
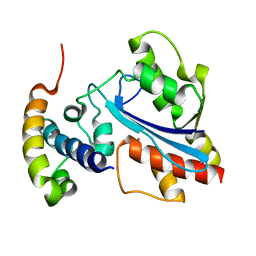 | |
2VDA
 
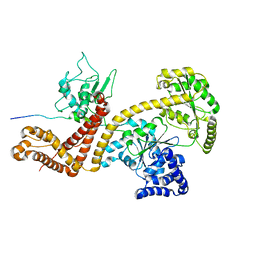 | | Solution structure of the SecA-signal peptide complex | | Descriptor: | MALTOPORIN, TRANSLOCASE SUBUNIT SECA | | Authors: | Gelis, I, Bonvin, A.M.J.J, Keramisanou, D, Koukaki, M, Gouridis, G, Karamanou, S, Economou, A, Kalodimos, C.G. | | Deposit date: | 2007-10-01 | | Release date: | 2007-11-27 | | Last modified: | 2018-01-24 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for Signal-Sequence Recognition by the Translocase Motor Seca as Determined by NMR
Cell(Cambridge,Mass.), 131, 2007
|
|
1WCO
 
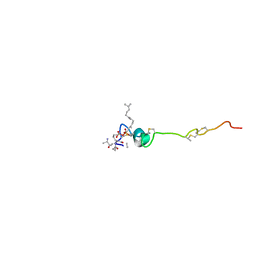 | | The solution structure of the nisin-lipid II complex | | Descriptor: | (2E,6E)-12-fluoro-11-(fluoromethyl)-3,7-dimethyldodeca-2,6,10-trien-1-yl trihydrogen diphosphate, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-N-acetyl-alpha-muramic acid, ALA-FGA-LYS-DAL-DAL PEPTIDE, ... | | Authors: | Hsu, S.-T.D, Breukink, E, Tischenko, E, Lutters, M.A.G, de Kruijff, B, Kaptein, R, Bonvin, A.M.J.J, van Nuland, N.A.J. | | Deposit date: | 2004-11-19 | | Release date: | 2005-03-07 | | Last modified: | 2020-07-29 | | Method: | SOLUTION NMR | | Cite: | The nisin-lipid II complex reveals a pyrophosphate cage that provides a blueprint for novel antibiotics.
Nat. Struct. Mol. Biol., 11, 2004
|
|
1SIY
 
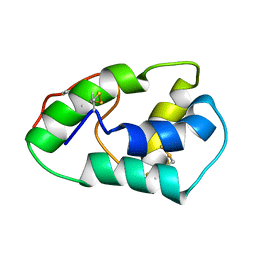 | | NMR structure of mung bean non-specific lipid transfer protein 1 | | Descriptor: | Nonspecific lipid-transfer protein 1 | | Authors: | Lin, K.F, Liu, Y.N, Hsu, S.T.D, Samuel, D, Cheng, C.S, Bonvin, A.M.J.J, Lyu, P.C. | | Deposit date: | 2004-03-02 | | Release date: | 2005-04-05 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Characterization and Structural Analyses of Nonspecific Lipid Transfer Protein 1 from Mung Bean
Biochemistry, 44, 2005
|
|
2BJC
 
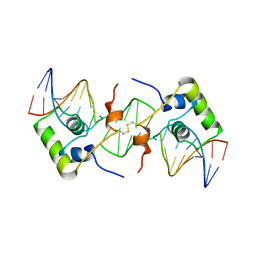 | | NMR structure of a protein-DNA complex of an altered specificity mutant of the lac repressor headpiece that mimics the gal repressor | | Descriptor: | 5'-D(*GP*AP*AP*TP*TP*GP*TP*AP*AP*GP *CP*GP*CP*TP*TP*AP*CP*AP*AP*TP*TP*C)-3', LACTOSE OPERON REPRESSOR | | Authors: | Salinas, R.K, Folkers, G.E, Bonvin, A.M.J.J, Das, D, Boelens, R, Kaptein, R. | | Deposit date: | 2005-02-01 | | Release date: | 2005-10-18 | | Last modified: | 2020-01-15 | | Method: | SOLUTION NMR | | Cite: | Altered Specificity in DNA Binding by the Lac Repressor: A Mutant Lac Headpiece that Mimics the Gal Repressor
Chembiochem, 6, 2005
|
|
2BGF
 
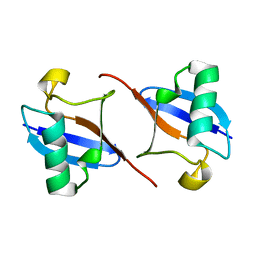 | |
2C06
 
 | | NMR-based model of the complex of the toxin Kid and a 5-nucleotide substrate RNA fragment (AUACA) | | Descriptor: | 5'-R(*AP*UP*AP*CP*AP)-3', KID TOXIN PROTEIN | | Authors: | Kamphuis, M.B, Bonvin, A.M.J.J, Monti, M.C, Lemonnier, M, Munoz-Gomez, A, Van Den Heuvel, R.H.H, Diaz-Orejas, R, Boelens, R. | | Deposit date: | 2005-08-25 | | Release date: | 2006-02-08 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Model for RNA Binding and the Catalytic Site of the Rnase Kid of the Bacterial Pard Toxin-Antitoxin System.
J.Mol.Biol., 357, 2006
|
|
1KFT
 
 | | Solution Structure of the C-Terminal domain of UvrC from E-coli | | Descriptor: | Excinuclease ABC subunit C | | Authors: | Singh, S, Folkers, G.E, Bonvin, A.M.J.J, Boelens, R, Wechselberger, R, Niztayev, A, Kaptein, R. | | Deposit date: | 2001-11-23 | | Release date: | 2002-11-20 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure and DNA-binding properties of the C-terminal domain of UvrC from E.coli
EMBO J., 21, 2002
|
|
1UR6
 
 | | NMR based structural model of the UbcH5B-CNOT4 complex | | Descriptor: | POTENTIAL TRANSCRIPTIONAL REPRESSOR NOT4HP, UBIQUITIN-CONJUGATING ENZYME E2-17 KDA 2, ZINC ION | | Authors: | Dominguez, C, Bonvin, A.M.J.J, Winkler, G.S, Van Schaik, F.M.A, Timmers, H.Th.M, Boelens, R. | | Deposit date: | 2003-10-27 | | Release date: | 2004-05-07 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR, THEORETICAL MODEL | | Cite: | Structural Model of the Ubch5B/Cnot4 Complex Revealed by Combining NMR, Mutagenesis, and Docking Approaches.
Structure, 12, 2004
|
|
1W4U
 
 | | NMR solution structure of the ubiquitin conjugating enzyme UbcH5B | | Descriptor: | UBIQUITIN-CONJUGATING ENZYME E2-17 KDA 2 | | Authors: | Houben, K, Dominguez, C, Van Schaik, F.M.A, Timmers, H.T.M, Bonvin, A.M.J.J, Boelens, R. | | Deposit date: | 2004-07-29 | | Release date: | 2004-11-10 | | Last modified: | 2017-04-19 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Ubiquitin-Conjugating Enzyme Ubch5B
J.Mol.Biol., 344, 2004
|
|
2K6G
 
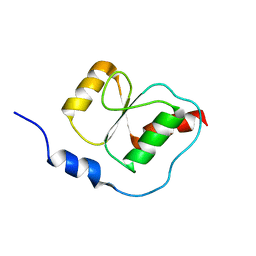 | |
2KXI
 
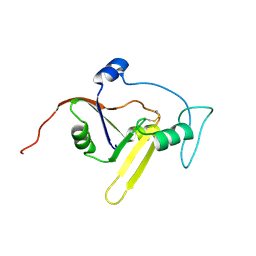 | | Solution NMR structure of the apoform of NarE (NMB1343) | | Descriptor: | Uncharacterized protein | | Authors: | Koehler, C, Carlier, L, Veggi, D, Soriani, M, Pizza, M, Boelens, R, Bonvin, A.M.J.J. | | Deposit date: | 2010-05-06 | | Release date: | 2011-03-02 | | Last modified: | 2018-08-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the apoform of NarE (NMB1343)
To be Published
|
|
1ARR
 
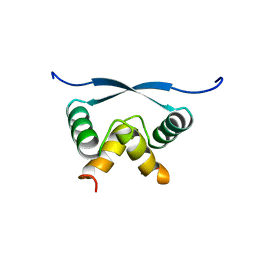 | | RELAXATION MATRIX REFINEMENT OF THE SOLUTION STRUCTURE OF THE ARC REPRESSOR | | Descriptor: | ARC REPRESSOR | | Authors: | Bonvin, A.M.J.J, Vis, H, Burgering, M.J.M, Breg, J.N, Boelens, R, Kaptein, R. | | Deposit date: | 1993-08-24 | | Release date: | 1994-01-31 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Nuclear magnetic resonance solution structure of the Arc repressor using relaxation matrix calculations.
J.Mol.Biol., 236, 1994
|
|
1ARQ
 
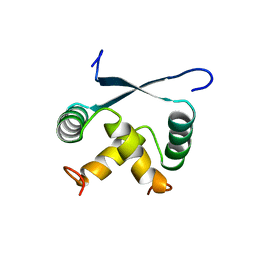 | | RELAXATION MATRIX REFINEMENT OF THE SOLUTION STRUCTURE OF THE ARC REPRESSOR | | Descriptor: | ARC REPRESSOR | | Authors: | Bonvin, A.M.J.J, Vis, H, Burgering, M.J.M, Breg, J.N, Boelens, R, Kaptein, R. | | Deposit date: | 1993-08-24 | | Release date: | 1994-01-31 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Nuclear magnetic resonance solution structure of the Arc repressor using relaxation matrix calculations.
J.Mol.Biol., 236, 1994
|
|
1CJG
 
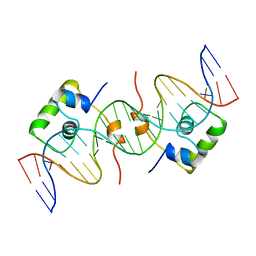 | | NMR STRUCTURE OF LAC REPRESSOR HP62-DNA COMPLEX | | Descriptor: | DNA (5'-D(*GP*AP*AP*TP*TP*GP*TP*GP*AP*GP*CP*GP*CP*TP*CP*AP*CP*AP*AP*TP*TP*C)-3'), PROTEIN (LAC REPRESSOR) | | Authors: | Spronk, C.A.E.M, Bonvin, A.M.J.J, Radha, P.K, Melacini, G, Boelens, R, Kaptein, R. | | Deposit date: | 1999-04-14 | | Release date: | 2000-01-01 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The solution structure of Lac repressor headpiece 62 complexed to a symmetrical lac operator.
Structure Fold.Des., 7, 1999
|
|
6YFY
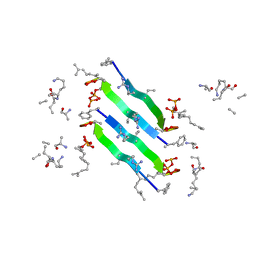 
 | | Solid-state NMR structure of the D-Arg4,L10-teixobactin - Lipid II complex in lipid bilayers. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-N-acetyl-alpha-muramic acid, 3-methylbut-2-en-1-ol, D-Arg4,Leu10-Teixobactin, ... | | Authors: | Weingarth, M.H, Shukla, R. | | Deposit date: | 2020-03-26 | | Release date: | 2020-06-10 | | Last modified: | 2023-11-15 | | Method: | SOLID-STATE NMR | | Cite: | Mode of action of teixobactins in cellular membranes.
Nat Commun, 11, 2020
|
|
1EFA
 
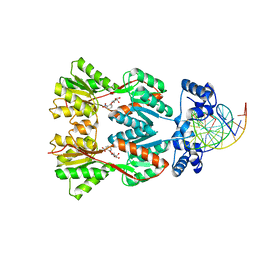 | |
3FID
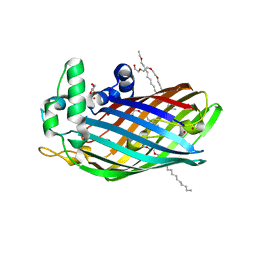 
 | | LpxR from Salmonella typhimurium | | Descriptor: | GLYCEROL, PENTAETHYLENE GLYCOL MONODECYL ETHER, Putative outer membrane protein (LpxR), ... | | Authors: | Rutten, L, Gros, P. | | Deposit date: | 2008-12-11 | | Release date: | 2009-02-10 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Active-site architecture and catalytic mechanism of the lipid A deacylase LpxR of Salmonella typhimurium
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
7QGV
 
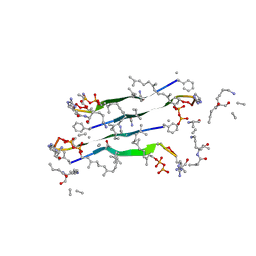 | | Solid-state NMR structure of Teixobactin-Lipid II. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-N-acetyl-alpha-muramic acid, 3-methylbut-2-en-1-ol, Lipid II, ... | | Authors: | Weingarth, M.H, Shukla, R. | | Deposit date: | 2021-12-10 | | Release date: | 2022-08-03 | | Last modified: | 2023-11-15 | | Method: | SOLID-STATE NMR | | Cite: | Teixobactin kills bacteria by a two-pronged attack on the cell envelope.
Nature, 608, 2022
|
|
1OSL
 
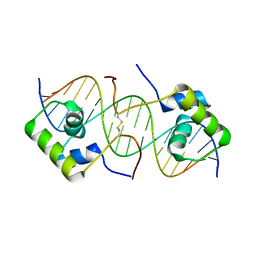 | | Solution structure of a dimeric lactose DNA-binding domain complexed to a nonspecific DNA sequence | | Descriptor: | 5'-D(*CP*GP*AP*TP*AP*AP*GP*AP*TP*AP*TP*CP*TP*TP*AP*TP*CP*G)-3', Lactose operon repressor | | Authors: | Kalodimos, C.G, Bonvin, A.M.J.J, Boelens, R, Kaptein, R. | | Deposit date: | 2003-03-20 | | Release date: | 2004-05-04 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | Structure and flexibility adaptation in nonspecific and specific protein-DNA complexes.
Science, 305, 2004
|
|
1RGD
 
 | | STRUCTURE REFINEMENT OF THE GLUCOCORTICOID RECEPTOR-DNA BINDING DOMAIN FROM NMR DATA BY RELAXATION MATRIX CALCULATIONS | | Descriptor: | GLUCOCORTICOID RECEPTOR, ZINC ION | | Authors: | Van Tilborg, M.A.A, Bonvin, A.M.J.J, Hard, K, Davis, A, Maler, B, Boelens, R, Yamamoto, K.R, Kaptein, R. | | Deposit date: | 1995-01-06 | | Release date: | 1995-02-14 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Structure refinement of the glucocorticoid receptor-DNA binding domain from NMR data by relaxation matrix calculations.
J.Mol.Biol., 247, 1995
|
|
1W9N
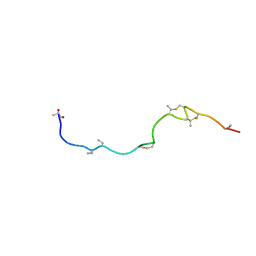 
 | | Isolation and characterization of epilancin 15X, a novel antibiotic from a clinical strain of Staphylococcus epidermidis | | Descriptor: | EPILANCIN 15X | | Authors: | Ekkelenkamp, M, Hanssen, M.G.M, Hsu, S.-T.D, de Jong, A, Milatovic, D, Verhoef, J, van Nuland, N.A.J. | | Deposit date: | 2004-10-14 | | Release date: | 2005-04-01 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Isolation and structural characterization of epilancin 15X, a novel lantibiotic from a clinical strain of Staphylococcus epidermidis.
FEBS Lett., 579, 2005
|
|
5U3A
 
 | |
