4EXV
 
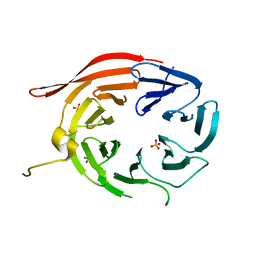 | | Structure of Kluyveromyces lactis Hsv2p | | Descriptor: | SULFATE ION, SVP1-like protein 2 | | Authors: | Baskaran, S, Hurley, J.H. | | Deposit date: | 2012-05-01 | | Release date: | 2012-07-04 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Two-Site Recognition of Phosphatidylinositol 3-Phosphate by PROPPINs in Autophagy.
Mol.Cell, 47, 2012
|
|
3NB0
 
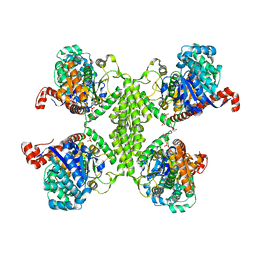 | |
3NAZ
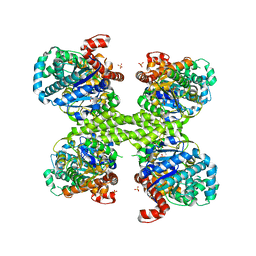 
 | | Basal state form of Yeast Glycogen Synthase | | Descriptor: | Glycogen [starch] synthase isoform 2, PEPTIDE, SULFATE ION | | Authors: | Baskaran, S, Hurley, T.D. | | Deposit date: | 2010-06-02 | | Release date: | 2010-10-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural basis for glucose-6-phosphate activation of glycogen synthase.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3NCH
 
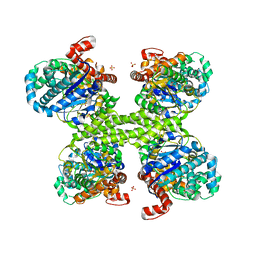 | |
3O3C
 
 | | Glycogen synthase basal state UDP complex | | Descriptor: | Glycogen [starch] synthase isoform 2, SULFATE ION, URIDINE-5'-DIPHOSPHATE | | Authors: | Baskaran, S, Hurley, T.D. | | Deposit date: | 2010-07-23 | | Release date: | 2010-10-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.512 Å) | | Cite: | Structural basis for glucose-6-phosphate activation of glycogen synthase.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3RSZ
 
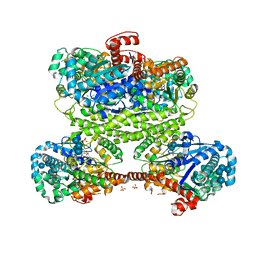 | | Maltodextran bound basal state conformation of yeast glycogen synthase isoform 2 | | Descriptor: | Glycogen [starch] synthase isoform 2, SULFATE ION, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Baskaran, S, Hurley, T.D. | | Deposit date: | 2011-05-02 | | Release date: | 2011-08-10 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.009 Å) | | Cite: | Multiple Glycogen-binding Sites in Eukaryotic Glycogen Synthase Are Required for High Catalytic Efficiency toward Glycogen.
J.Biol.Chem., 286, 2011
|
|
3RT1
 
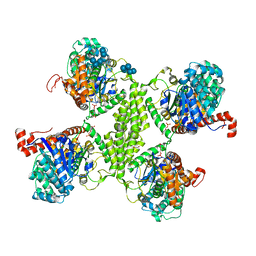 | |
5SUK
 
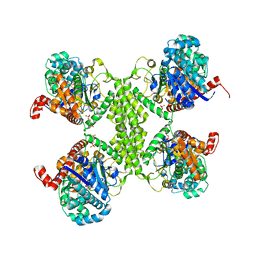 | |
3D14
 
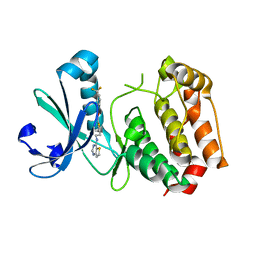 | | Crystal structure of mouse Aurora A (Asn186->Gly, Lys240->Arg, Met302->Leu) in complex with 1-{5-[2-(thieno[3,2-d]pyrimidin-4-ylamino)-ethyl]- thiazol-2-yl}-3-(3-trifluoromethyl-phenyl)-urea | | Descriptor: | 1-{5-[2-(thieno[3,2-d]pyrimidin-4-ylamino)ethyl]-1,3-thiazol-2-yl}-3-[3-(trifluoromethyl)phenyl]urea, serine/threonine kinase 6 | | Authors: | Elling, R.A, Baskaran, S, Allen, D.A, Oslob, J.D, Romanowski, M.J. | | Deposit date: | 2008-05-04 | | Release date: | 2008-08-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Discovery of a potent and selective aurora kinase inhibitor.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
6W3H
 
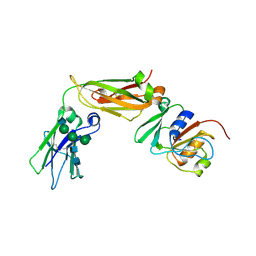 | | Brain delivery of therapeutic proteins using an Fc fragment blood-brain barrier transport vehicle in mice and monkeys | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ATV Fc, Transferrin receptor protein 1,Transferrin receptor protein 1 | | Authors: | Srivastava, A, Kariolis, M, Wells, R. | | Deposit date: | 2020-03-09 | | Release date: | 2020-06-10 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.38 Å) | | Cite: | Brain delivery of therapeutic proteins using an Fc fragment blood-brain barrier transport vehicle in mice and monkeys.
Sci Transl Med, 12, 2020
|
|
3SZB
 
 | |
3SZA
 
 | | Crystal structure of human ALDH3A1 - apo form | | Descriptor: | ACETATE ION, Aldehyde dehydrogenase, dimeric NADP-preferring, ... | | Authors: | Khanna, M, Hurley, T.D. | | Deposit date: | 2011-07-18 | | Release date: | 2011-11-02 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Discovery of a novel class of covalent inhibitor for aldehyde dehydrogenases.
J.Biol.Chem., 286, 2011
|
|
3SZ9
 
 | |
3D15
 
 | | Crystal structure of mouse Aurora A (Asn186->Gly, Lys240->Arg, Met302->Leu) in complex with 1-(3-chloro-phenyl)-3-{5-[2-(thieno[3,2-d]pyrimidin-4-ylamino)- ethyl]-thiazol-2-yl}-urea [SNS-314] | | Descriptor: | 1-(3-chlorophenyl)-3-{5-[2-(thieno[3,2-d]pyrimidin-4-ylamino)ethyl]-1,3-thiazol-2-yl}urea, serine/threonine kinase 6 | | Authors: | Elling, R.A, Bui, M, Allen, D.A, Oslob, J.D, Romanowski, M.J. | | Deposit date: | 2008-05-04 | | Release date: | 2009-05-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Discovery of a potent and selective aurora kinase inhibitor.
Bioorg.Med.Chem.Lett., 18, 2008
|
|
1RWW
 
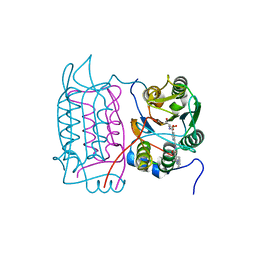 | |
1RWX
 
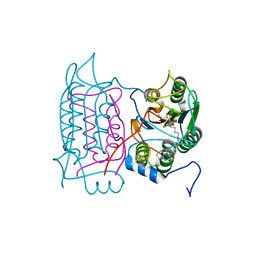 | |
5SUL
 
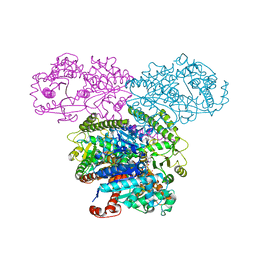 | | Inhibited state structure of yGsy2p | | Descriptor: | Glycogen [starch] synthase isoform 2, URIDINE-5'-DIPHOSPHATE, URIDINE-5'-MONOPHOSPHATE | | Authors: | Mahalingan, K.K, Hurley, T.D. | | Deposit date: | 2016-08-03 | | Release date: | 2017-06-14 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Redox Switch for the Inhibited State of Yeast Glycogen Synthase Mimics Regulation by Phosphorylation.
Biochemistry, 56, 2017
|
|
3D2I
 
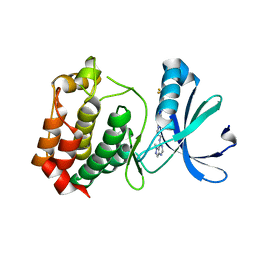 | | Crystal structure of mouse Aurora A (Asn186->Gly, Lys240->Arg, Met302->Leu) in complex with 1-{5-[2-(1-methyl-1H-pyrazolo[4,3-d]pyrimidin-7-ylamino)-ethyl]-thiazol-2-yl}-3-(3-trifluoromethyl-phenyl)-urea | | Descriptor: | 1-(5-{2-[(1-methyl-1H-pyrazolo[4,3-d]pyrimidin-7-yl)amino]ethyl}-1,3-thiazol-2-yl)-3-[3-(trifluoromethyl)phenyl]urea, serine/threonine kinase 6 | | Authors: | Oslob, J.D, Yu, C, Romanowski, M.J. | | Deposit date: | 2008-05-08 | | Release date: | 2009-05-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Discovery of Aurora-A-selective inhibitors
To be Published
|
|
3D2K
 
 | | Crystal structure of mouse Aurora A (Asn186->Gly, Lys240->Arg, Met302->Leu) in complex with [7-(2-{2-[3-(3-chloro-phenyl)-ureido]-thiazol-5-yl}-ethylamino)-pyrazolo[4,3-d]pyrimidin-1-yl]-acetic acid | | Descriptor: | (7-{[2-(2-{[(3-chlorophenyl)carbamoyl]amino}-1,3-thiazol-5-yl)ethyl]amino}-1H-pyrazolo[4,3-d]pyrimidin-1-yl)acetic acid, serine/threonine kinase 6 | | Authors: | Elling, R.A, Oslob, J.D, Yu, C, Romanowski, M.J. | | Deposit date: | 2008-05-08 | | Release date: | 2009-05-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Discovery of Aurora-A-selective inhibitors
To be Published
|
|
4KQM
 
 | |
4KQ1
 
 | |
4KQ2
 
 | | Glucose1,2cyclic phosphate bound activated state of Yeast Glycogen Synthase | | Descriptor: | (2R,3aR,5R,6S,7S,7aR)-5-(hydroxymethyl)tetrahydro-3aH-[1,3,2]dioxaphospholo[4,5-b]pyran-2,6,7-triol 2-oxide, 6-O-phosphono-alpha-D-glucopyranose, BARIUM ION, ... | | Authors: | Chikwana, V.M, Hurley, T.D. | | Deposit date: | 2013-05-14 | | Release date: | 2013-12-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural basis for 2'-phosphate incorporation into glycogen by glycogen synthase.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
