1A91
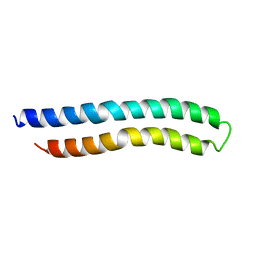 
 | | SUBUNIT C OF THE F1FO ATP SYNTHASE OF ESCHERICHIA COLI; NMR, 10 STRUCTURES | | Descriptor: | F1FO ATPASE SUBUNIT C | | Authors: | Girvin, M.E, Rastogi, V.K, Abildgaard, F, Markley, J.L, Fillingame, R.H. | | Deposit date: | 1998-04-15 | | Release date: | 1998-07-01 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the transmembrane H+-transporting subunit c of the F1F0 ATP synthase.
Biochemistry, 37, 1998
|
|
1BRZ
 
 | | SOLUTION STRUCTURE OF THE SWEET PROTEIN BRAZZEIN, NMR, 43 STRUCTURES | | Descriptor: | BRAZZEIN | | Authors: | Caldwell, J.E, Abildgaard, F, Dzakula, Z, Ming, D, Hellekant, G, Markley, J.L. | | Deposit date: | 1998-03-12 | | Release date: | 1998-07-01 | | Last modified: | 2019-12-25 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the thermostable sweet-tasting protein brazzein.
Nat.Struct.Biol., 5, 1998
|
|
1C0V
 
 | | SUBUNIT C OF THE F1FO ATP SYNTHASE OF ESCHERICHIA COLI; NMR, 10 STRUCTURES | | Descriptor: | PROTEIN (F1FO ATPASE SUBUNIT C) | | Authors: | Girvin, M.E, Rastogi, V.K, Abildgaard, F, Markley, J.L, Fillingame, R.H. | | Deposit date: | 1999-07-22 | | Release date: | 1999-08-18 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the transmembrane H+-transporting subunit c of the F1F0 ATP synthase.
Biochemistry, 37, 1998
|
|
1G90
 
 | |
1SYZ
 
 | |
2BRZ
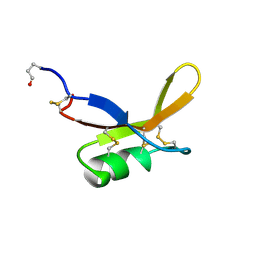 
 | | SOLUTION NMR STRUCTURE OF THE SWEET PROTEIN BRAZZEIN, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | BRAZZEIN | | Authors: | Caldwell, J.E, Abildgaard, F, Dzakula, Z, Ming, D, Hellekant, G, Markley, J.L. | | Deposit date: | 1998-04-30 | | Release date: | 1998-07-01 | | Last modified: | 2019-12-25 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the thermostable sweet-tasting protein brazzein.
Nat.Struct.Biol., 5, 1998
|
|
2ARF
 
 | | Solution structure of the Wilson ATPase N-domain in the presence of ATP | | Descriptor: | WILSON DISEASE ATPASE | | Authors: | Dmitriev, O, Tsivkovskii, R, Abildgaard, F, Morgan, C.T, Markley, J.L, Lutsenko, S. | | Deposit date: | 2005-08-19 | | Release date: | 2006-02-28 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the N-domain of Wilson disease protein: Distinct nucleotide-binding environment and effects of disease mutations
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
1JOR
 
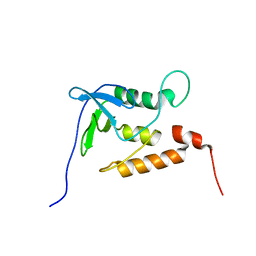 | | Ensemble structures for unligated Staphylococcal nuclease-H124L | | Descriptor: | staphylococcal nuclease | | Authors: | Wang, J, Truckses, D.M, Abildgaard, F, Dzakula, Z, Zolnai, Z, Markley, J.L. | | Deposit date: | 2001-07-30 | | Release date: | 2001-08-22 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | Solution structures of staphylococcal nuclease from multidimensional, multinuclear NMR: nuclease-H124L and its ternary complex with Ca2+ and thymidine-3',5'-bisphosphate.
J.Biomol.NMR, 10, 1997
|
|
1JOO
 
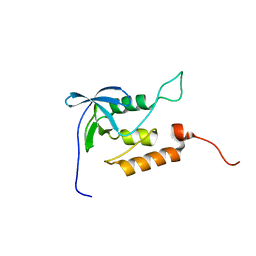 | | Averaged structure for unligated Staphylococcal nuclease-H124L | | Descriptor: | staphylococcal nuclease | | Authors: | Wang, J, Truckses, D.M, Abildgaard, F, Dzakula, Z, Zolnai, Z, Markley, J.L. | | Deposit date: | 2001-07-30 | | Release date: | 2001-08-22 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | Solution structures of staphylococcal nuclease from multidimensional, multinuclear NMR: nuclease-H124L and its ternary complex with Ca2+ and thymidine-3',5'-bisphosphate.
J.Biomol.NMR, 10, 1997
|
|
1JOK
 
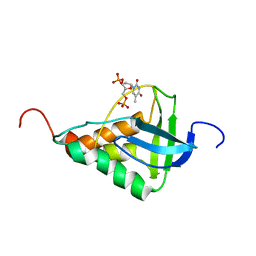 | | Averaged structure for Staphylococcal nuclease-H124L in ternary complex with Ca2+ and thymidine-3',5'-bisphosphate | | Descriptor: | THYMIDINE-3',5'-DIPHOSPHATE, staphylococcal nuclease | | Authors: | Wang, J, Truckses, D.M, Abildgaard, F, Dzakula, Z, Zolnai, Z, Markley, J.L. | | Deposit date: | 2001-07-30 | | Release date: | 2001-08-22 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | Solution structures of staphylococcal nuclease from multidimensional, multinuclear NMR: nuclease-H124L and its ternary complex with Ca2+ and thymidine-3',5'-bisphosphate.
J.Biomol.NMR, 10, 1997
|
|
1JOQ
 
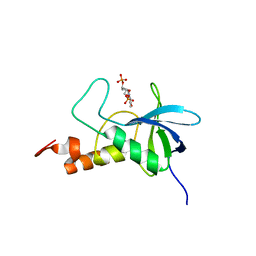 | | Ensemble structures for Staphylococcal nuclease-H124L in ternary complex with Ca2+ and thymidine-3',5'-bisphosphate | | Descriptor: | THYMIDINE-3',5'-DIPHOSPHATE, staphylococcal nuclease | | Authors: | Wang, J, Truckses, D.M, Abildgaard, F, Dzakula, Z, Zolnai, Z, Markley, J.L. | | Deposit date: | 2001-07-30 | | Release date: | 2001-08-22 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | Solution structures of staphylococcal nuclease from multidimensional, multinuclear NMR: nuclease-H124L and its ternary complex with Ca2+ and thymidine-3',5'-bisphosphate.
J.Biomol.NMR, 10, 1997
|
|
1L6T
 
 | |
1BLR
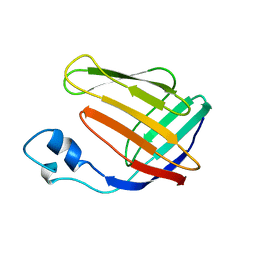 
 | | NMR SOLUTION STRUCTURE OF HUMAN CELLULAR RETINOIC ACID BINDING PROTEIN-TYPE II, 22 STRUCTURES | | Descriptor: | CELLULAR RETINOIC ACID BINDING PROTEIN-TYPE II | | Authors: | Wang, L, Li, Y, Abilddard, F, Yan, H, Markely, J. | | Deposit date: | 1998-07-20 | | Release date: | 1999-01-13 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of type II human cellular retinoic acid binding protein: implications for ligand binding.
Biochemistry, 37, 1998
|
|
1SYM
 
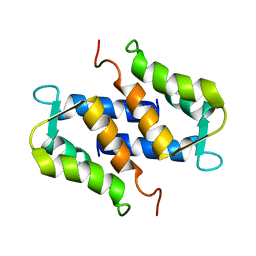 | |
1B4C
 
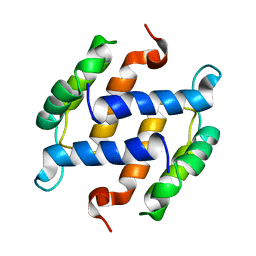 | | SOLUTION STRUCTURE OF RAT APO-S100B USING DIPOLAR COUPLINGS | | Descriptor: | PROTEIN (S-100 PROTEIN, BETA CHAIN) | | Authors: | Weber, D.J, Drohat, A.C, Tjandra, N, Baldisseri, D.M. | | Deposit date: | 1998-12-17 | | Release date: | 1998-12-30 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | The use of dipolar couplings for determining the solution structure of rat apo-S100B(betabeta).
Protein Sci., 8, 1999
|
|
1CO1
 
 | | FOLD OF THE CBFA | | Descriptor: | CORE BINDING FACTOR ALPHA | | Authors: | Berardi, M.J, Bushweller, J.H. | | Deposit date: | 1999-05-31 | | Release date: | 2000-06-07 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The Ig fold of the core binding factor alpha Runt domain is a member of a family of structurally and functionally related Ig-fold DNA-binding domains.
Structure Fold.Des., 7, 1999
|
|
1QLK
 
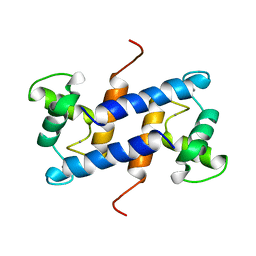 | | SOLUTION STRUCTURE OF CA(2+)-LOADED RAT S100B (BETABETA) NMR, 20 STRUCTURES | | Descriptor: | CALCIUM ION, S-100 PROTEIN | | Authors: | Drohat, A.C, Baldisseri, D.M, Rustandi, R.R, Weber, D.J. | | Deposit date: | 1997-09-26 | | Release date: | 1998-11-11 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Solution structure of calcium-bound rat S100B(betabeta) as determined by nuclear magnetic resonance spectroscopy,.
Biochemistry, 37, 1998
|
|
1C99
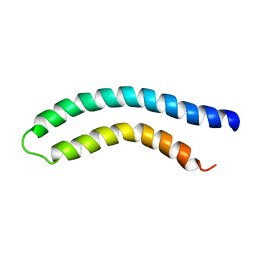 
 | |
2KEZ
 
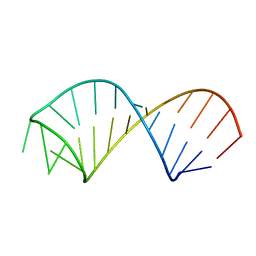 | | NMR structure of U6 ISL at pH 8.0 | | Descriptor: | RNA (5'-R(*GP*GP*UP*UP*CP*CP*CP*CP*UP*GP*CP*AP*UP*AP*AP*GP*GP*AP*UP*GP*AP*AP*CP*C)-3') | | Authors: | Venditti, V, Butcher, S.E. | | Deposit date: | 2009-02-08 | | Release date: | 2009-07-21 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Minimum-energy path for a u6 RNA conformational change involving protonation, base-pair rearrangement and base flipping.
J.Mol.Biol., 391, 2009
|
|
2KF0
 
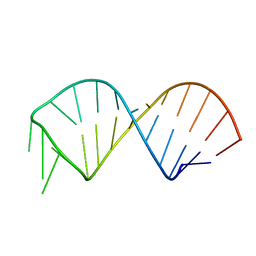 | | NMR structure of U6 ISL at pH 7.0 | | Descriptor: | RNA (5'-R(*GP*GP*UP*UP*CP*CP*CP*CP*UP*GP*CP*AP*UP*AP*AP*GP*GP*AP*UP*GP*AP*AP*CP*C)-3') | | Authors: | Venditti, V, Butcher, S.E. | | Deposit date: | 2009-02-08 | | Release date: | 2009-07-21 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Minimum-energy path for a u6 RNA conformational change involving protonation, base-pair rearrangement and base flipping.
J.Mol.Biol., 391, 2009
|
|
1MWN
 
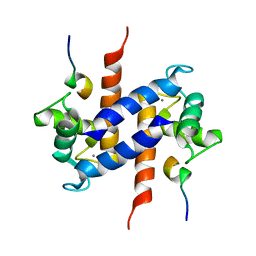 | | Solution NMR structure of S100B bound to the high-affinity target peptide TRTK-12 | | Descriptor: | CALCIUM ION, F-actin capping protein alpha-1 subunit, S-100 protein, ... | | Authors: | Inman, K.G, Yang, R, Rustandi, R.R, Miller, K.E, Baldisseri, D.M, Weber, D.J. | | Deposit date: | 2002-09-30 | | Release date: | 2002-12-18 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of S100B bound to the high-affinity target peptide TRTK-12
J.Mol.Biol., 324, 2002
|
|
1QO1
 
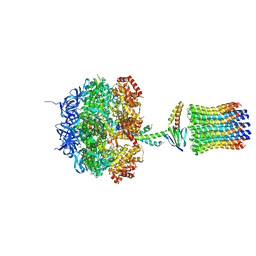 | | Molecular Architecture of the Rotary Motor in ATP Synthase from Yeast Mitochondria | | Descriptor: | ATP SYNTHASE ALPHA CHAIN, ATP SYNTHASE BETA CHAIN, ATP SYNTHASE DELTA CHAIN, ... | | Authors: | Stock, D, Leslie, A.G.W, Walker, J.E. | | Deposit date: | 1999-11-01 | | Release date: | 1999-11-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Molecular Architecture of the Rotary Motor in ATP Synthase
Science, 286, 1999
|
|
