2D29
 
 | |
2D3K
 
 | |
6L6K
 
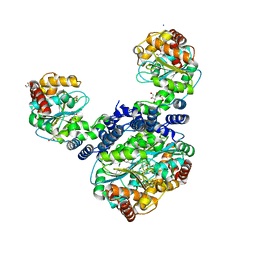
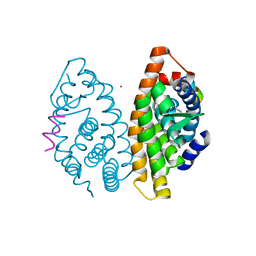 | | Crystal structure of dimeric RXRalpha-LBD complexed with partial agonist CBt-PMN and SRC1 | | Descriptor: | 1-(3,5,5,8,8-pentamethyl-6,7-dihydronaphthalen-2-yl)benzotriazole-5-carboxylic acid, CALCIUM ION, Nuclear receptor coactivator 1, ... | | Authors: | Shimizu, K, Numoto, N, Nakano, S, Makishima, M, Kakuta, H, Ito, N. | | Deposit date: | 2019-10-29 | | Release date: | 2020-11-04 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of dimeric RXRalpha-LBD complexed with partial agonist CBt-PMN and SRC1
To Be Published
|
|
1V8G
 
 | |
1X1O
 
 | |
1WN2
 
 | |
1WS9
 
 | |
1WU8
 
 | |
2DSL
 
 | |
2DEK
 
 | |
2DSJ
 
 | |
2DVL
 
 | |
2DY0
 
 | |
2E9Y
 
 | |
2ENI
 
 | |
2E18
 
 | |
2ENW
 
 | |
2EJJ
 
 | |
2ELD
 
 | |
2E4R
 
 | |
2EJK
 
 | |
2ED3
 
 | |
2EEQ
 
 | |
2EMU
 
 | |
2ELC
 
 | |
