2LW8
 
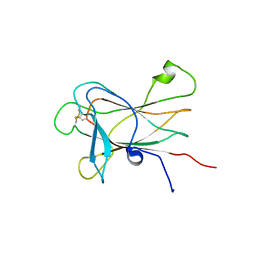 | |
2KOU
 
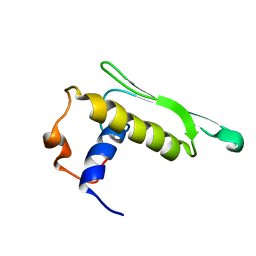 | | DICER LIKE protein | | Descriptor: | Dicer-like protein 4 | | Authors: | Qin, H, Song, J, Yuan, Y.A. | | Deposit date: | 2009-09-30 | | Release date: | 2010-02-16 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Structure of the Arabidopsis thaliana DCL4 DUF283 domain reveals a noncanonical double-stranded RNA-binding fold for protein-protein interaction
Rna, 2010
|
|
2MDK
 
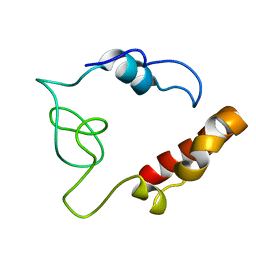 | |
3YGS
 
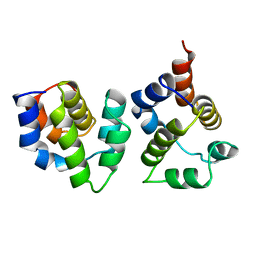 | | APAF-1 CARD IN COMPLEX WITH PRODOMAIN OF PROCASPASE-9 | | Descriptor: | APOPTOTIC PROTEASE ACTIVATING FACTOR 1, PROCASPASE 9 | | Authors: | Qin, H, Srinivasula, S, Wu, G, Fernandes-Alnemri, T, Alnemri, E, Shi, Y. | | Deposit date: | 1999-05-08 | | Release date: | 2000-04-19 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis of procaspase-9 recruitment by the apoptotic protease-activating factor 1.
Nature, 399, 1999
|
|
5ZSY
 
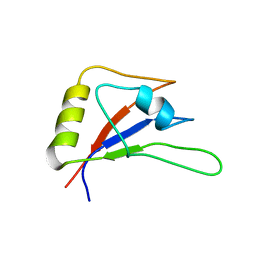 | | RBM10-RRM2 domain and its lung cancer related mutant | | Descriptor: | RNA-binding protein 10 | | Authors: | Qin, H. | | Deposit date: | 2018-04-30 | | Release date: | 2019-06-12 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural, dynamics, and RNA binding comparison between RBM10-RRM2 domain and its lung cancer related mutation.
To be published
|
|
5ZSW
 
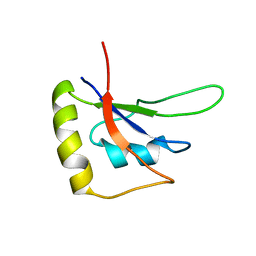 | | RBM10-RRM2 domain and its lung cancer related mutant | | Descriptor: | RNA-binding protein 10 | | Authors: | Qin, H. | | Deposit date: | 2018-04-30 | | Release date: | 2019-05-22 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural, dynamics, and RNA binding comparison between RBM10-RRM2 domain and its lung cancer related mutation.
To be published
|
|
8U1T
 
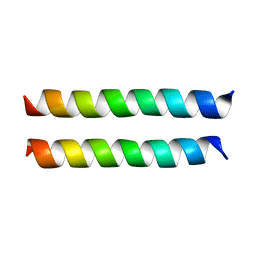 | | SARS-CoV-2 Envelope Protein Transmembrane Domain: Dimeric Structure Determined by Solid-State NMR | | Descriptor: | Envelope small membrane protein | | Authors: | Zhang, R, Qin, H, Prasad, R, Fu, R, Zhou, H.X, Cross, T. | | Deposit date: | 2023-09-02 | | Release date: | 2023-11-15 | | Method: | SOLID-STATE NMR | | Cite: | Dimeric Transmembrane Structure of the SARS-CoV-2 E Protein.
Commun Biol, 6, 2023
|
|
2L0J
 
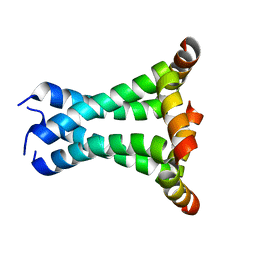 | | Solid State NMR structure of the M2 proton channel from Influenza A Virus in hydrated lipid bilayer | | Descriptor: | Matrix protein 2 | | Authors: | Sharma, M, Yi, M, Dong, H, Qin, H, Peterson, E, Busath, D.D, Zhou, H.X, Cross, T.A. | | Deposit date: | 2010-07-08 | | Release date: | 2010-11-03 | | Last modified: | 2011-07-13 | | Method: | SOLID-STATE NMR | | Cite: | Insight into the mechanism of the influenza a proton channel from a structure in a lipid bilayer.
Science, 330, 2010
|
|
2MBE
 
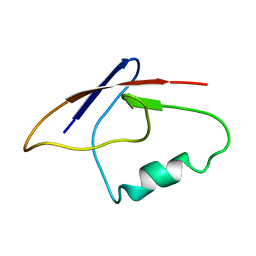 | |
3GXU
 
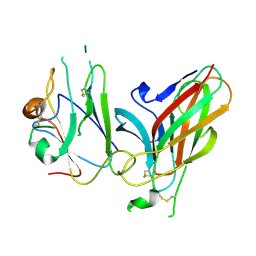 | | Crystal structure of Eph receptor and ephrin complex | | Descriptor: | Ephrin type-A receptor 4, Ephrin-B2 | | Authors: | Qin, H.N, Song, J.X. | | Deposit date: | 2009-04-03 | | Release date: | 2009-10-27 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural characterization of the EphA4-ephrin-B2 complex reveals new features enabling Eph-ephrin binding promiscuity
J.Biol.Chem., 2009
|
|
2N6E
 
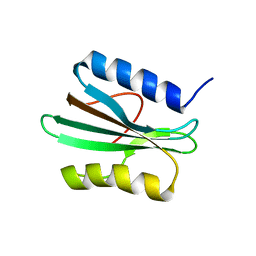 | |
3CKH
 
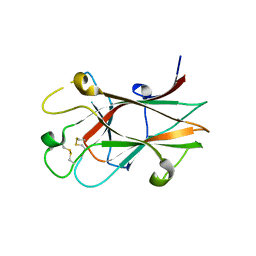 | | Crystal structure of Eph A4 receptor | | Descriptor: | Ephrin type-A receptor 4 | | Authors: | Shi, J.H, Song, J.X. | | Deposit date: | 2008-03-15 | | Release date: | 2008-09-23 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure and NMR Binding Reveal That Two Small Molecule Antagonists Target the High Affinity Ephrin-binding Channel of the EphA4 Receptor.
J.Biol.Chem., 283, 2008
|
|
2YGS
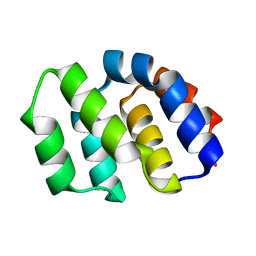 
 | | CARD DOMAIN FROM APAF-1 | | Descriptor: | APOPTOTIC PROTEASE ACTIVATING FACTOR 1 | | Authors: | Shi, Y. | | Deposit date: | 1999-05-08 | | Release date: | 2000-04-19 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural basis of procaspase-9 recruitment by the apoptotic protease-activating factor 1.
Nature, 399, 1999
|
|
5X57
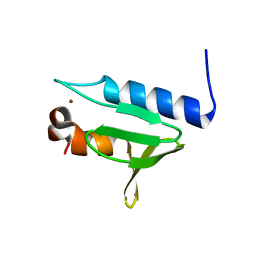 
 | | Structure of GAR domain of ACF7 | | Descriptor: | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5, NICKEL (II) ION | | Authors: | Yang, F, Wang, T, Zhang, Y, Wu, X.Y. | | Deposit date: | 2017-02-15 | | Release date: | 2017-07-05 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | ACF7 regulates inflammatory colitis and intestinal wound response by orchestrating tight junction dynamics.
Nat Commun, 8, 2017
|
|
8H41
 
 | | Crystal structure of a decarboxylase from Trichosporon moniliiforme in complex with o-nitrophenol | | Descriptor: | MAGNESIUM ION, O-NITROPHENOL, Salicylate decarboxylase | | Authors: | Gao, J, Zhao, Y.P, Li, Q, Liu, W.D, Sheng, X. | | Deposit date: | 2022-10-09 | | Release date: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | A Combined Computational-Experimental Study on the Substrate Binding and Reaction Mechanism of Salicylic Acid Decarboxylase
Catalysts, 12, 2022
|
|
