1XP4
 
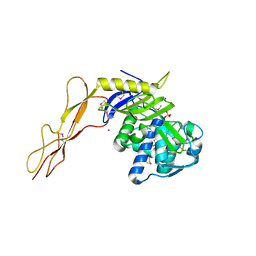 | | Crystal structure of a peptidoglycan synthesis regulatory factor (PBP3) from Streptococcus pneumoniae | | Descriptor: | D-alanyl-D-alanine carboxypeptidase, IODIDE ION, SULFATE ION | | Authors: | Morlot, C, Pernot, L, Le Gouellec, A, Di Guilmi, A.M, Vernet, T, Dideberg, O, Dessen, A. | | Deposit date: | 2004-10-08 | | Release date: | 2004-11-09 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of a peptidoglycan synthesis regulatory factor (PBP3) from Streptococcus pneumoniae
J.Biol.Chem., 280, 2005
|
|
6FQB
 
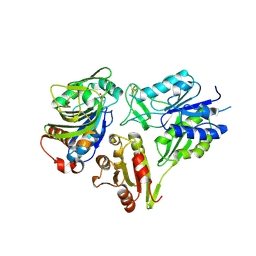 | | MurT/GatD peptidoglycan amidotransferase complex from Streptococcus pneumoniae R6 | | Descriptor: | Cobyric acid synthase, GLUTAMINE, Mur ligase family protein | | Authors: | Morlot, C, Contreras-Martel, C, Leisico, F, Straume, D, Peters, K, Hegnar, O.A, Simon, N, Villard, A.M, Breukink, E, Gravier-Pelletier, C, Le Corre, L, Vollmer, W, Pietrancosta, N, Havarstein, L.S, Zapun, A. | | Deposit date: | 2018-02-13 | | Release date: | 2018-08-22 | | Last modified: | 2018-11-07 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of the essential peptidoglycan amidotransferase MurT/GatD complex from Streptococcus pneumoniae.
Nat Commun, 9, 2018
|
|
2V9T
 
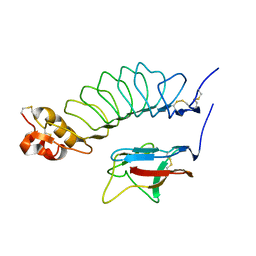 | |
2V9Q
 
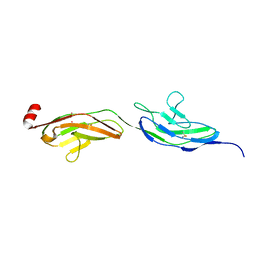 | |
2V9S
 
 | | Second LRR domain of human Slit2 | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, SLIT HOMOLOG 2 PROTEIN N-PRODUCT | | Authors: | Morlot, C, Cusack, S, McCarthy, A.A. | | Deposit date: | 2007-08-25 | | Release date: | 2007-09-25 | | Last modified: | 2014-11-05 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Insights Into the Slit-Robo Complex.
Proc.Natl.Acad.Sci.USA, 104, 2007
|
|
2V9R
 
 | |
2V70
 
 | | Third LRR domain of human Slit2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, SLIT HOMOLOG 2 PROTEIN N-PRODUCT | | Authors: | Morlot, C, Cusack, S, McCarthy, A.A. | | Deposit date: | 2007-07-24 | | Release date: | 2007-08-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.01 Å) | | Cite: | Production of Slit2 Lrr Domains in Mammalian Cells for Structural Studies and the Structure of Human Slit2 Domain 3.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
4BH5
 
 | | LytM domain of EnvC, an activator of cell wall amidases in Escherichia coli | | Descriptor: | CHLORIDE ION, GLYCEROL, IODIDE ION, ... | | Authors: | Morlot, C, Peters, N.T, Yang, D.C, Uehara, T, Vernet, T, Bernhardt, T.G. | | Deposit date: | 2013-03-29 | | Release date: | 2013-07-03 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Structure-Function Analysis of the Lytm Domain of Envc, an Activator of Cell Wall Remodeling at the Escherichia Coli Division Site.
Mol.Microbiol., 89, 2013
|
|
7PEG
 
 | | Structure of the sporulation/germination protein YhcN from Bacillus subtilis | | Descriptor: | Probable spore germination lipoprotein YhcN | | Authors: | Liu, B, Chan, H, Bauda, E, Contreras-Martel, C, Bellard, L, Villard, A.M, Mas, C, Neumann, E, Fenel, D, Favier, A, Serrano, M, Henriques, A.O.H, Rodrigues, C.D.A, Morlot, C. | | Deposit date: | 2021-08-10 | | Release date: | 2022-08-24 | | Last modified: | 2022-09-07 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Structural insights into ring-building motif domains involved in bacterial sporulation.
J.Struct.Biol., 214, 2022
|
|
5MKC
 
 | | Crystal structure of the RrgA Jo.In complex | | Descriptor: | CALCIUM ION, Cell wall surface anchor family protein (Jo),Cell wall surface anchor family protein (In), NICKEL (II) ION, ... | | Authors: | Bonnet, J, Cartannaz, J, Tourcier, G, Contreras-Martel, C, Kleman, J.P, Fenel, D, Schoehn, G, Morlot, C, Vernet, T, Di Guilmi, A.M. | | Deposit date: | 2016-12-03 | | Release date: | 2017-03-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Autocatalytic association of proteins by covalent bond formation: a Bio Molecular Welding toolbox derived from a bacterial adhesin.
Sci Rep, 7, 2017
|
|
4LJB
 
 | | Structure of a photobleached state of IrisFP under high intensity laser-light | | Descriptor: | Green to red photoconvertible GPF-like protein EosFP, SULFATE ION, SULFITE ION | | Authors: | Duan, C, Adam, V, Byrdin, M, Ridard, J, Kieffer-Jacquinod, S, Morlot, C, Arcizet, D, Demachy, I, Bourgeois, D. | | Deposit date: | 2013-07-04 | | Release date: | 2013-10-09 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9019 Å) | | Cite: | Structural evidence for a two-regime photobleaching mechanism in a reversibly switchable fluorescent protein.
J.Am.Chem.Soc., 135, 2013
|
|
4LJD
 
 | | Structure of a photobleached state of IrisFP under low intensity laser-light | | Descriptor: | Green to red photoconvertible GPF-like protein EosFP, SULFATE ION, SULFITE ION | | Authors: | Duan, C, Adam, V, Byrdin, M, Ridard, J, Kieffer-Jacquinod, S, Morlot, C, Arcizet, D, Demachy, I, Bourgeois, D. | | Deposit date: | 2013-07-04 | | Release date: | 2013-10-09 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural evidence for a two-regime photobleaching mechanism in a reversibly switchable fluorescent protein.
J.Am.Chem.Soc., 135, 2013
|
|
4LJC
 
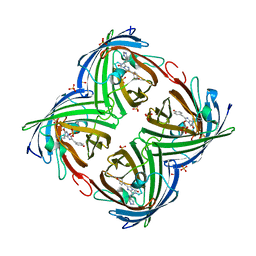 | | Structure of an X-ray-induced photobleached state of IrisFP | | Descriptor: | Green to red photoconvertible GPF-like protein EosFP, SULFATE ION, SULFITE ION | | Authors: | Duan, C, Adam, V, Byrdin, M, Ridard, J, Kieffer-Jacquinod, S, Morlot, C, Arcizet, D, Demachy, I, Bourgeois, D. | | Deposit date: | 2013-07-04 | | Release date: | 2013-10-09 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Structural evidence for a two-regime photobleaching mechanism in a reversibly switchable fluorescent protein.
J.Am.Chem.Soc., 135, 2013
|
|
6HWM
 
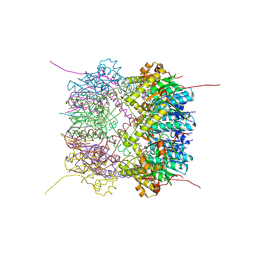 | | Structure of Thermus thermophilus ClpP in complex with bortezomib | | Descriptor: | ATP-dependent Clp protease proteolytic subunit, DI(HYDROXYETHYL)ETHER, N-[(1R)-1-(DIHYDROXYBORYL)-3-METHYLBUTYL]-N-(PYRAZIN-2-YLCARBONYL)-L-PHENYLALANINAMIDE | | Authors: | Felix, J, Schanda, P, Fraga, H, Morlot, C. | | Deposit date: | 2018-10-12 | | Release date: | 2019-09-18 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Mechanism of the allosteric activation of the ClpP protease machinery by substrates and active-site inhibitors.
Sci Adv, 5, 2019
|
|
6GZ8
 
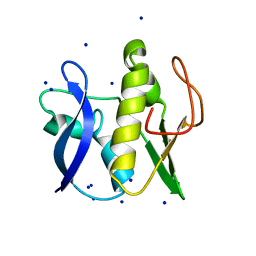 | | First GerMN domain of the sporulation protein GerM from Bacillus subtilis | | Descriptor: | 1,2-ETHANEDIOL, SODIUM ION, Spore germination protein GerM | | Authors: | Trouve, J, Mohamed, A, Leisico, F, Contreras-Martel, C, Liu, B, Mas, C, Rudner, D.Z, Rodrigues, C.D.A, Morlot, C. | | Deposit date: | 2018-07-03 | | Release date: | 2018-10-10 | | Last modified: | 2018-12-26 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | Structural characterization of the sporulation protein GerM from Bacillus subtilis.
J. Struct. Biol., 204, 2018
|
|
6GZB
 
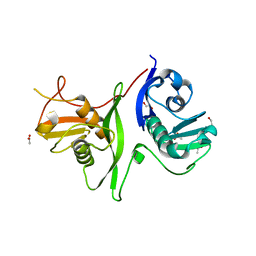 | | Tandem GerMN domains of the sporulation protein GerM from Bacillus subtilis | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Spore germination protein GerM | | Authors: | Trouve, J, Mohamed, A, Leisico, F, Contreras-Martel, C, Liu, B, Mas, C, Rudner, D.Z, Rodrigues, C.D.A, Morlot, C. | | Deposit date: | 2018-07-03 | | Release date: | 2018-10-10 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural characterization of the sporulation protein GerM from Bacillus subtilis.
J. Struct. Biol., 204, 2018
|
|
6HWN
 
 | | Structure of Thermus thermophilus ClpP in complex with a tripeptide. | | Descriptor: | ATP-dependent Clp protease proteolytic subunit, DI(HYDROXYETHYL)ETHER, Unknown tripeptide | | Authors: | Felix, J, Schanda, P, Fraga, H, Morlot, C. | | Deposit date: | 2018-10-12 | | Release date: | 2019-09-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Mechanism of the allosteric activation of the ClpP protease machinery by substrates and active-site inhibitors.
Sci Adv, 5, 2019
|
|
4X36
 
 | | Crystal structure of the autolysin LytA from Streptococcus pneumoniae TIGR4 | | Descriptor: | Autolysin, CHOLINE ION, GLYCEROL, ... | | Authors: | Cheng, W, Li, Q, Zhou, C.Z, Chen, Y.X. | | Deposit date: | 2014-11-28 | | Release date: | 2015-05-27 | | Last modified: | 2015-06-24 | | Method: | X-RAY DIFFRACTION (2.101 Å) | | Cite: | Full-length structure of the major autolysin LytA.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
6H9D
 
 | |
