3UOR
 
 | |
2YMW
 
 | |
1AZW
 
 | | PROLINE IMINOPEPTIDASE FROM XANTHOMONAS CAMPESTRIS PV. CITRI | | Descriptor: | PROLINE IMINOPEPTIDASE | | Authors: | Medrano, F.J, Alonso, J, Garcia, J.L, Romero, A, Bode, W, Gomis-Ruth, F.X. | | Deposit date: | 1997-11-22 | | Release date: | 1999-01-13 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of proline iminopeptidase from Xanthomonas campestris pv. citri: a prototype for the prolyl oligopeptidase family.
EMBO J., 17, 1998
|
|
8BGD
 
 | | Structure of Mpro from SARS-CoV-2 in complex with FGA147 | | Descriptor: | (phenylmethyl) ~{N}-[(2~{S})-4-methyl-1-[[(2~{S})-4-nitro-1-[(3~{R})-2-oxidanylidenepyrrolidin-3-yl]butan-2-yl]amino]-1-oxidanylidene-pentan-2-yl]carbamate, 3C-like proteinase nsp5 | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2022-10-27 | | Release date: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.621 Å) | | Cite: | Peptidyl Nitroalkenes Inhibit SARS-CoV-2 Main Protease and Block Virus Replication
To Be Published
|
|
8BFO
 
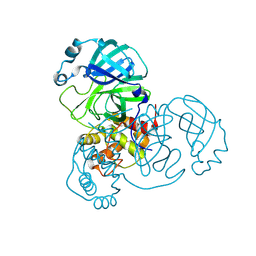 | |
8BGA
 
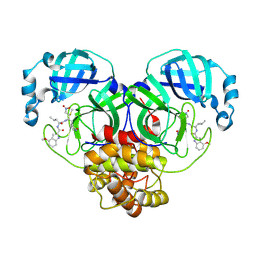 | | Structure of Mpro in complex with FGA146 | | Descriptor: | 3C-like proteinase nsp5, 4-methoxy-~{N}-[(2~{S})-4-methyl-1-[[(2~{S})-4-nitro-1-[(3~{S})-2-oxidanylidenepyrrolidin-3-yl]butan-2-yl]amino]-1-oxidanylidene-pentan-2-yl]-1~{H}-indole-2-carboxamide | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2022-10-27 | | Release date: | 2023-11-08 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.982 Å) | | Cite: | Peptidyl nitroalkene inhibitors of main protease rationalized by computational and crystallographic investigations as antivirals against SARS-CoV-2.
Commun Chem, 7, 2024
|
|
8BFQ
 
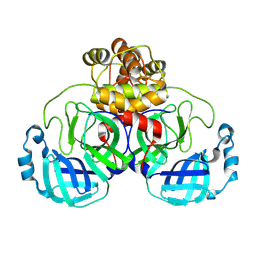 | |
1AU8
 | | HUMAN CATHEPSIN G | | Descriptor: | CATHEPSIN G, N-(3-carboxypropanoyl)-L-valyl-N-[(1R)-5-amino-1-phosphonopentyl]-L-prolinamide | | Authors: | Medrano, F.J, Bode, W, Banbula, A, Potempa, J. | | Deposit date: | 1997-09-12 | | Release date: | 1998-10-14 | | Last modified: | 2012-12-12 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | HUMAN CATHEPSIN G
to be published
|
|
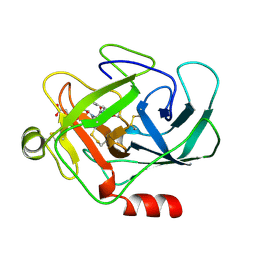
2NUH
 
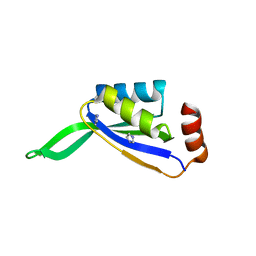 | |
4W7L
 
 | |
4W7N
 
 | |
4W7O
 
 | |
4W7J
 
 | |
4W7K
 
 | |
4W7M
 
 | |
8PAQ
 | |
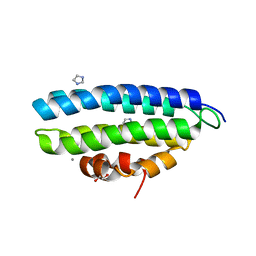
5ABN
 
 | | CRYSTAL STRUCTURE ANALYSIS OF FUNGAL VERSATILE PEROXIDASE FROM PLEUROTUS ERYNGII. MUTANT VPi. MUTATED RESIDUES D69S, T70D, S86E, D146T, Q202L, H232E, Q239R AND S301K. | | Descriptor: | CALCIUM ION, MAGNESIUM ION, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2015-08-07 | | Release date: | 2015-11-04 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.194 Å) | | Cite: | Improving the Ph-Stability of Versatile Peroxidase by Comparative Structural Analysis with a Naturally-Stable Manganese Peroxidase.
Plos One, 10, 2015
|
|
5ABO
 
 | | CRYSTAL STRUCTURE ANALYSIS OF FUNGAL VERSATILE PEROXIDASE FROM PLEUROTUS ERYNGII. MUTANT VPi-br. MUTATED RESIDUES T2K, D69S, T70D, S86E, A131K, D146T, Q202L, Q219K, H232E, Q239R, L288R, S301K, A308R, A309K AND A314R. | | Descriptor: | CALCIUM ION, PROTOPORPHYRIN IX CONTAINING FE, VERSATILE PEROXIDASE VPL2 | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2015-08-07 | | Release date: | 2015-11-04 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.095 Å) | | Cite: | Improving the Ph-Stability of Versatile Peroxidase by Comparative Structural Analysis with a Naturally-Stable Manganese Peroxidase.
Plos One, 10, 2015
|
|
5ABQ
 
 | | CRYSTAL STRUCTURE ANALYSIS OF FUNGAL VERSATILE PEROXIDASE FROM PLEUROTUS ERYNGII. MUTANT VPi-SS. MUTATED RESIDUES T2K, A49C, A61C, D69S, T70D, S86E, A131K, D146T, Q202L, Q219K, H232E, Q239R, L288R, S301K, A308R,A309K AND A314R. | | Descriptor: | CALCIUM ION, PROTOPORPHYRIN IX CONTAINING FE, VERSATILE PEROXIDASE | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2015-08-07 | | Release date: | 2015-11-04 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.293 Å) | | Cite: | Improving the Ph-Stability of Versatile Peroxidase by Comparative Structural Analysis with a Naturally-Stable Manganese Peroxidase.
Plos One, 10, 2015
|
|
5FNE
 
 | |
5FNB
 
 | | CRYSTAL STRUCTURE OF FUNGAL VERSATILE PEROXIDASE FROM PLEUROTUS ERYNGII SEPTUPLE MUTANT E37K, H39R, V160A, T184M, Q202L, D213A & G330R | | Descriptor: | CALCIUM ION, PROTOPORPHYRIN IX CONTAINING FE, SULFATE ION, ... | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2015-11-13 | | Release date: | 2016-07-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.792 Å) | | Cite: | Unveiling the Basis of Alkaline Stability of an Evolved Versatile Peroxidase.
Biochem.J., 473, 2016
|
|
1P19
 
 | |
1P17
 
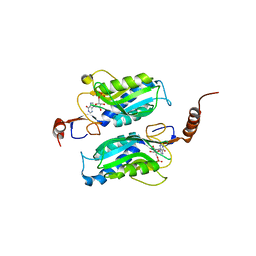 | | Hypoxanthine Phosphoribosyltransferase from Trypanosoma cruzi, K68R mutant, complexed with the product IMP | | Descriptor: | INOSINIC ACID, hypoxanthine phosphoribosyltransferase | | Authors: | Medrano, F.J, Eakin, A.E, Craig III, S.P. | | Deposit date: | 2003-04-11 | | Release date: | 2004-05-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Interactions at the dimer interface influence the relative efficiencies for purine nucleotide synthesis and pyrophosphorolysis in a phosphoribosyltransferase.
J.Mol.Biol., 335, 2004
|
|
8BZ3
 
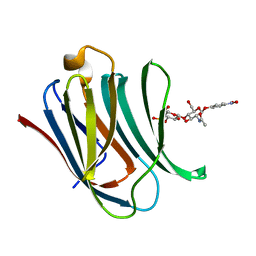 | | Structure od the carbohydrate reconition domain of Gal3 in comples with SAF-2-010 | | Descriptor: | Galectin-3, [(2~{S},3~{R},4~{S},5~{S},6~{R})-2-[(2~{R},3~{S},4~{R},5~{R},6~{S})-5-acetamido-2-(hydroxymethyl)-6-(4-nitrophenoxy)-4-oxidanyl-oxan-3-yl]oxy-6-(hydroxymethyl)-3,5-bis(oxidanyl)oxan-4-yl] hydrogen sulfate | | Authors: | Medrano, F.J, Romero, A. | | Deposit date: | 2022-12-14 | | Release date: | 2023-03-01 | | Last modified: | 2023-03-08 | | Method: | X-RAY DIFFRACTION (1.31 Å) | | Cite: | Selectively Modified Lactose and N -Acetyllactosamine Analogs at Three Key Positions to Afford Effective Galectin-3 Ligands.
Int J Mol Sci, 24, 2023
|
|
5ANH
 
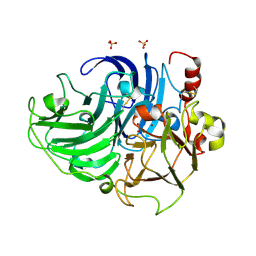 | |
