4DQW
 
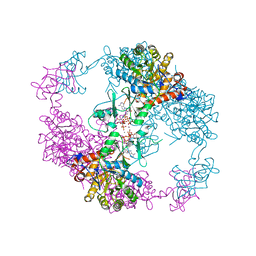 | | Crystal Structure Analysis of PA3770 | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, GLYCEROL, Inosine-5'-monophosphate dehydrogenase, ... | | Authors: | Labesse, G, Munier-Lehmann, H. | | Deposit date: | 2012-02-16 | | Release date: | 2013-03-06 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | MgATP Regulates Allostery and Fiber Formation in IMPDHs.
Structure, 21, 2013
|
|
2W1Z
 
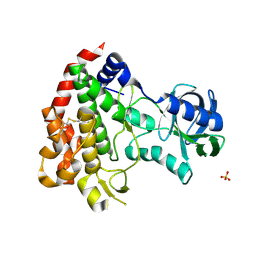 | | ROP2 from Toxoplasma gondii: A virulence factor with a protein- kinase fold and no enzymatic activity. | | Descriptor: | ROP2, SULFATE ION | | Authors: | Labesse, G, Gelin, M, Bessin, Y, Lebrun, M, Arold, S.T, Dubremetz, J.-F. | | Deposit date: | 2008-10-21 | | Release date: | 2008-12-09 | | Last modified: | 2011-09-21 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Rop2 from Toxoplasma Gondii: A Virulence Factor with a Protein-Kinase Fold and No Enzymatic Activity.
Structure, 17, 2009
|
|
3NWY
 
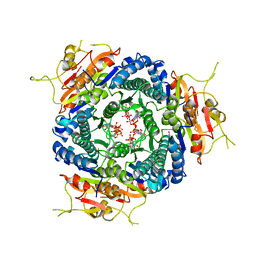 | |
5AHN
 | | IMP-bound form of the D199N mutant of IMPDH from Pseudomonas aeruginosa | | Descriptor: | INOSINE-5'-MONOPHOSPHATE DEHYDROGENASE, INOSINIC ACID, MAGNESIUM ION | | Authors: | Labesse, G, Alexandre, T, Gelin, M, Haouz, A, Munier-Lehmann, H. | | Deposit date: | 2015-02-06 | | Release date: | 2015-07-15 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.652 Å) | | Cite: | Crystallographic Studies of Two Variants of Pseudomonas Aeruginosa Impdh with Impaired Allosteric Regulation
Acta Crystallogr.,Sect.D, 71, 2015
|
|
5AHL
 
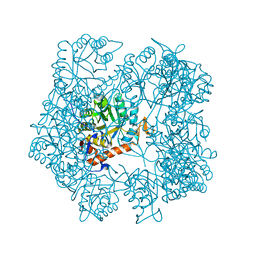 | | Apo-form of the DeltaCBS mutant of IMPDH from Pseudomonas aeruginosa | | Descriptor: | INOSINE-5'-MONOPHOSPHATE DEHYDROGENASE, SODIUM ION | | Authors: | Labesse, G, Alexandre, T, Gelin, M, Haouz, A, Munier-Lehmann, H. | | Deposit date: | 2015-02-06 | | Release date: | 2015-07-15 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.951 Å) | | Cite: | Crystallographic Studies of Two Variants of Pseudomonas Aeruginosa Impdh with Impaired Allosteric Regulation
Acta Crystallogr.,Sect.D, 71, 2015
|
|
5AHM
 
 | | IMP-bound form of the DeltaCBS mutant of IMPDH from Pseudomonas aeruginosa | | Descriptor: | INOSINE-5'-MONOPHOSPHATE DEHYDROGENASE, INOSINIC ACID, PHOSPHATE ION | | Authors: | Labesse, G, Alexandre, T, Gelin, M, Haouz, A, Munier-Lehmann, H. | | Deposit date: | 2015-02-06 | | Release date: | 2015-07-15 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Crystallographic Studies of Two Variants of Pseudomonas Aeruginosa Impdh with Impaired Allosteric Regulation
Acta Crystallogr.,Sect.D, 71, 2015
|
|
7QBJ
 
 | | bacterial IMPDH chimera | | Descriptor: | Inosine-5'-monophosphate dehydrogenase | | Authors: | Labesse, G, Gelin, M, Munier-Lehmann, H, Gedeon, A, Haouz, A. | | Deposit date: | 2021-11-19 | | Release date: | 2023-05-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Insight into the role of the Bateman domain at the molecular and physiological levels through engineered IMP dehydrogenases.
Protein Sci., 32, 2023
|
|
7QDX
 
 | | bacterial IMPDH chimera | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Inosine-5'-monophosphate dehydrogenase, MAGNESIUM ION | | Authors: | Labesse, G, Gelin, M, Munier-Lehmann, H, Gedeon, A, Haouz, A. | | Deposit date: | 2021-11-30 | | Release date: | 2023-06-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Insight into the role of the Bateman domain at the molecular and physiological levels through engineered IMP dehydrogenases.
Protein Sci., 32, 2023
|
|
7R4J
 
 | | Crystal structure of human mitochondrial NAD kinase | | Descriptor: | CALCIUM ION, NAD kinase 2, mitochondrial | | Authors: | Labesse, G, Mary, C, Gelin, M, Lionne, C. | | Deposit date: | 2022-02-08 | | Release date: | 2022-07-06 | | Last modified: | 2022-09-14 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Crystal structure of human NADK2 reveals a dimeric organization and active site occlusion by lysine acetylation.
Mol.Cell, 82, 2022
|
|
7R4M
 
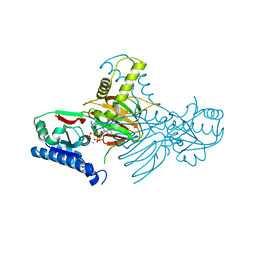 | | Crystal structure of mitochondrial NAD kinase | | Descriptor: | CALCIUM ION, NAD kinase 2, mitochondrial, ... | | Authors: | Labesse, G, Mary, C, Gelin, M, Lionne, C. | | Deposit date: | 2022-02-08 | | Release date: | 2022-07-06 | | Last modified: | 2022-09-14 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Crystal structure of human NADK2 reveals a dimeric organization and active site occlusion by lysine acetylation.
Mol.Cell, 82, 2022
|
|
7R4K
 
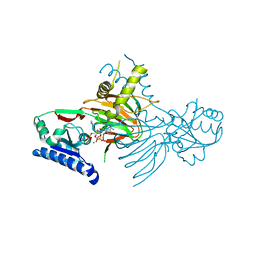 | | Crystal structure of human mitochondrial NAD kinase | | Descriptor: | MAGNESIUM ION, NAD kinase 2, mitochondrial, ... | | Authors: | Labesse, G, Mary, C, Gelin, M, Lionne, C. | | Deposit date: | 2022-02-08 | | Release date: | 2022-07-06 | | Last modified: | 2022-09-14 | | Method: | X-RAY DIFFRACTION (3.33 Å) | | Cite: | Crystal structure of human NADK2 reveals a dimeric organization and active site occlusion by lysine acetylation.
Mol.Cell, 82, 2022
|
|
7R4L
 | | Crystal structure of human mitochondrial NAD kinase | | Descriptor: | FE (III) ION, NAD kinase 2, mitochondrial, ... | | Authors: | Labesse, G, Mary, C, Gelin, M, Lionne, C. | | Deposit date: | 2022-02-08 | | Release date: | 2022-07-06 | | Last modified: | 2022-09-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of human NADK2 reveals a dimeric organization and active site occlusion by lysine acetylation.
Mol.Cell, 82, 2022
|
|
7BIX
 
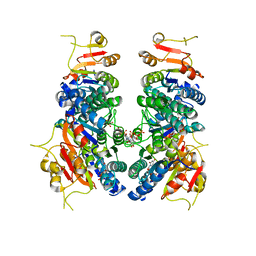 | | Crystal structure of UMPK from M. tuberculosis in complex with UDP and UTP (C2 form) | | Descriptor: | URIDINE 5'-TRIPHOSPHATE, URIDINE-5'-DIPHOSPHATE, Uridylate kinase | | Authors: | Labesse, G, Walter, P, Haouz, A, Mechaly, A.E, Munier-Lehmann, H. | | Deposit date: | 2021-01-13 | | Release date: | 2022-03-02 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.12 Å) | | Cite: | Structural basis for the allosteric inhibition of UMP kinase from Gram-positive bacteria, a promising antibacterial target.
Febs J., 289, 2022
|
|
7QEM
 
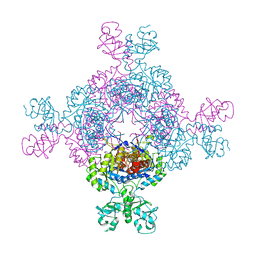 | | bacterial IMPDH chimera | | Descriptor: | INOSINIC ACID, Inosine-5'-monophosphate dehydrogenase | | Authors: | Labesse, G, Gelin, M, Gedeon, A, Haouz, A, Munier-Lehmann, H. | | Deposit date: | 2021-12-03 | | Release date: | 2023-06-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.09 Å) | | Cite: | Insight into the role of the Bateman domain at the molecular and physiological levels through engineered IMP dehydrogenases.
Protein Sci., 32, 2023
|
|
4FYH
 | | Crystal structure of rcl with phospho-triciribine | | Descriptor: | 5-methyl-1-(5-O-phosphono-beta-D-ribofuranosyl)-1,5-dihydro-1,4,5,6,8-pentaazaacenaphthylen-3-amine, Deoxyribonucleoside 5'-monophosphate N-glycosidase, SULFATE ION | | Authors: | Labesse, G, Padilla, A, Kaminski, P.A. | | Deposit date: | 2012-07-04 | | Release date: | 2013-01-02 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Structure of the oncoprotein Rcl bound to three nucleotide analogues.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4FYK
 
 | | Crystal structure of rcl with 5'-phiosphorothioate-adenosine | | Descriptor: | ADENOSINE -5'-THIO-MONOPHOSPHATE, Deoxyribonucleoside 5'-monophosphate N-glycosidase, SULFATE ION | | Authors: | Labesse, G, Padilla, A, Kaminski, P.A. | | Deposit date: | 2012-07-04 | | Release date: | 2013-01-02 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structure of the oncoprotein Rcl bound to three nucleotide analogues.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
4FYI
 
 | | Crystal structure of rcl with 6-cyclopentyl-AMP | | Descriptor: | Deoxyribonucleoside 5'-monophosphate N-glycosidase, N-cyclopentyladenosine 5'-(dihydrogen phosphate) | | Authors: | Labesse, G, Padilla, A, Kaminski, P.A. | | Deposit date: | 2012-07-04 | | Release date: | 2013-01-02 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Structure of the oncoprotein Rcl bound to three nucleotide analogues.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
2I2A
 
 | | Crystal structure of LmNADK1 from Listeria monocytogenes | | Descriptor: | GLYCEROL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Probable inorganic polyphosphate/ATP-NAD kinase 1 | | Authors: | Labesse, G. | | Deposit date: | 2006-08-16 | | Release date: | 2007-08-07 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | NAD kinases use substrate-assisted catalysis for specific recognition of NAD.
J.Biol.Chem., 282, 2007
|
|
6GK9
 
 | | Inhibited structure of IMPDH from Pseudomonas aeruginosa | | Descriptor: | (5~{S})-7-azanyl-5-(4-chlorophenyl)-2,4-bis(oxidanylidene)-1,5-dihydropyrano[2,3-d]pyrimidine-6-carbonitrile, Inosine-5'-monophosphate dehydrogenase, SULFATE ION | | Authors: | Labesse, G, Alexandre, T, Haouz, A, Munier-Lehmann, H. | | Deposit date: | 2018-05-18 | | Release date: | 2019-02-27 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | First-in-class allosteric inhibitors of bacterial IMPDHs.
Eur J Med Chem, 167, 2019
|
|
6GJV
 | |
4U77
 
 | | BTB domain from Drosophila CP190 | | Descriptor: | Centrosome-associated zinc finger protein CP190, DIMETHYL SULFOXIDE, GLYCEROL | | Authors: | Allemand, F, Cohen-Gonsaud, M, Labesse, G, Nollmann, M. | | Deposit date: | 2014-07-30 | | Release date: | 2014-08-13 | | Last modified: | 2014-10-15 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Chromatin Insulator Factors Involved in Long-Range DNA Interactions and Their Role in the Folding of the Drosophila Genome.
Plos Genet., 10, 2014
|
|
4P5E
 
 | | CRYSTAL STRUCTURE OF HUMAN DNPH1 (RCL) WITH 6-NAPHTHYL-PURINE-RIBOSIDE-MONOPHOSPHATE | | Descriptor: | 2'-deoxynucleoside 5'-phosphate N-hydrolase 1, 6-(naphthalen-2-yl)-9-(5-O-phosphono-beta-D-ribofuranosyl)-9H-purine, CALCIUM ION | | Authors: | Padilla, A, Labesse, G, Kaminski, P.A. | | Deposit date: | 2014-03-16 | | Release date: | 2014-08-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | 6-(Hetero)Arylpurine nucleotides as inhibitors of the oncogenic target DNPH1: Synthesis, structural studies and cytotoxic activities.
Eur.J.Med.Chem., 85C, 2014
|
|
4P5D
 
 | | CRYSTAL STRUCTURE OF RAT DNPH1 (RCL) WITH 6-NAPHTHYL-PURINE-RIBOSIDE-MONOPHOSPHATE | | Descriptor: | 2'-deoxynucleoside 5'-phosphate N-hydrolase 1, 6-(naphthalen-2-yl)-9-(5-O-phosphono-beta-D-ribofuranosyl)-9H-purine, SULFATE ION | | Authors: | Padilla, A, Labesse, G, Kaminski, P.A. | | Deposit date: | 2014-03-16 | | Release date: | 2014-08-20 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | 6-(Hetero)Arylpurine nucleotides as inhibitors of the oncogenic target DNPH1: Synthesis, structural studies and cytotoxic activities.
Eur.J.Med.Chem., 85C, 2014
|
|
4XP3
 
 | | Crystal structure of ERK2 in complex with an inhibitor | | Descriptor: | 2-amino-1,9-dihydro-6H-purine-6-thione, DIMETHYL SULFOXIDE, Mitogen-activated protein kinase 1, ... | | Authors: | Gelin, M, Allemand, F, Labesse, G, Guichou, J.F. | | Deposit date: | 2015-01-16 | | Release date: | 2015-08-12 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.782 Å) | | Cite: | Combining `dry' co-crystallization and in situ diffraction to facilitate ligand screening by X-ray crystallography.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4XNE
 
 | | Crystal structure of ERK2 in complex with an inhibitor | | Descriptor: | 1-phenyl-1H-1,2,4-triazole-3,5-diamine, Mitogen-activated protein kinase 1, SULFATE ION | | Authors: | Gelin, M, Allemand, F, Labesse, G, Guichou, J.F. | | Deposit date: | 2015-01-15 | | Release date: | 2015-08-12 | | Last modified: | 2015-08-19 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Combining `dry' co-crystallization and in situ diffraction to facilitate ligand screening by X-ray crystallography.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
