1IBA
 
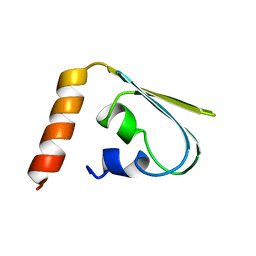 | | GLUCOSE PERMEASE (DOMAIN IIB), NMR, 11 STRUCTURES | | Descriptor: | GLUCOSE PERMEASE | | Authors: | Eberstadt, M, Grdadolnik, S.G, Gemmecker, G, Kessler, H, Buhr, A, Erni, B. | | Deposit date: | 1996-03-23 | | Release date: | 1996-10-14 | | Last modified: | 2017-11-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the IIB domain of the glucose transporter of Escherichia coli.
Biochemistry, 35, 1996
|
|
1PDO
 
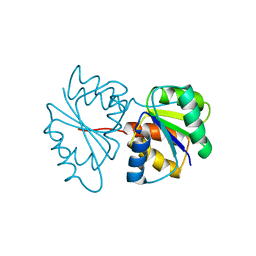 | |
2BTD
 
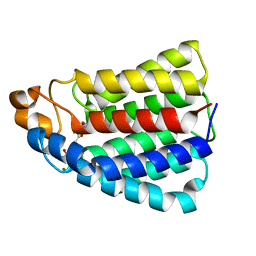 | | Crystal structure of DhaL from E. coli | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, PTS-DEPENDENT DIHYDROXYACETONE KINASE | | Authors: | Oberholzer, A.E, Schneider, P, Bachler, C, Baumann, U, Erni, B. | | Deposit date: | 2005-05-27 | | Release date: | 2006-06-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structure of the Nucleotide-Binding Subunit Dhal of the Escherichia Coli Dihydroxyacetone Kinase.
J.Mol.Biol., 359, 2006
|
|
2BG5
 | | Crystal Structure of the Phosphoenolpyruvate-binding Enzyme I-Domain from the Thermoanaerobacter tengcongensis PEP: Sugar Phosphotransferase System (PTS) | | Descriptor: | PHOSPHOENOLPYRUVATE-PROTEIN KINASE | | Authors: | Oberholzer, A.E, Bumann, M, Schneider, P, Baechler, C, Siebold, C, Baumann, U, Erni, B. | | Deposit date: | 2004-12-17 | | Release date: | 2005-02-02 | | Last modified: | 2018-03-07 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Crystal Structure of the Phosphoenolpyruvate-Binding Enzyme I-Domain from the Thermoanaerobacter Tengcongensis Pep: Sugar Phosphotransferase System (Pts)
J.Mol.Biol., 346, 2005
|
|
2WQD
 
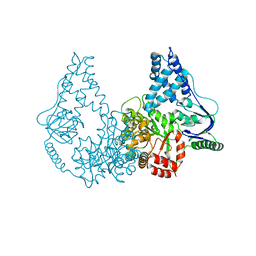 | | Crystal structure of enzyme I of the phosphoenolpyruvate:sugar phosphotransferase system in the dephosphorylated state | | Descriptor: | CALCIUM ION, PHOSPHOENOLPYRUVATE-PROTEIN PHOSPHOTRANSFERASE | | Authors: | Oberholzer, A.E, Schneider, P, Siebold, C, Baumann, U, Erni, B. | | Deposit date: | 2009-08-19 | | Release date: | 2009-10-13 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of Enzyme I of the Phosphoenolpyruvate:Sugar Phosphotransferase System in the Dephosphorylated State.
J.Biol.Chem., 284, 2009
|
|
3CT4
 
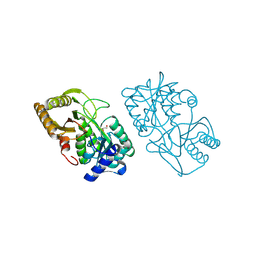 | | Structure of Dha-kinase subunit DhaK from L. Lactis | | Descriptor: | Dihydroxyacetone, PTS-dependent dihydroxyacetone kinase, dihydroxyacetone-binding subunit dhaK | | Authors: | Jeckelmann, J.M, Zurbriggen, A, Christen, S, Baumann, U, Erni, B. | | Deposit date: | 2008-04-11 | | Release date: | 2008-10-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.498 Å) | | Cite: | X-ray Structures of the Three Lactococcus lactis Dihydroxyacetone Kinase Subunits and of a Transient Intersubunit Complex.
J.Biol.Chem., 283, 2008
|
|
3CR3
 
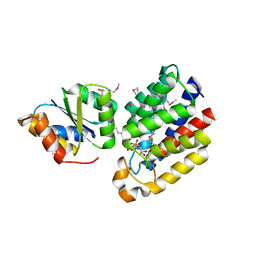 | | Structure of a transient complex between Dha-kinase subunits DhaM and DhaL from Lactococcus lactis | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, PTS-dependent dihydroxyacetone kinase, ... | | Authors: | Jeckelmann, J.M, Zurbriggen, A, Christen, S, Baumann, U, Erni, B. | | Deposit date: | 2008-04-04 | | Release date: | 2008-10-21 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | X-ray Structures of the Three Lactococcus lactis Dihydroxyacetone Kinase Subunits and of a Transient Intersubunit Complex.
J.Biol.Chem., 283, 2008
|
|
2XZ9
 
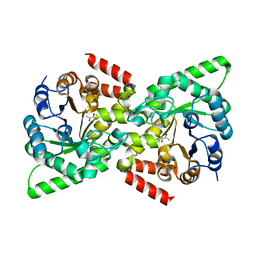 | | CRYSTAL STRUCTURE FROM THE PHOSPHOENOLPYRUVATE-BINDING DOMAIN OF ENZYME I IN COMPLEX WITH PYRUVATE FROM THE THERMOANAEROBACTER TENGCONGENSIS PEP-SUGAR PHOSPHOTRANSFERASE SYSTEM (PTS) | | Descriptor: | MAGNESIUM ION, PHOSPHOENOLPYRUVATE-PROTEIN KINASE (PTS SYSTEM EI COMPONENT IN BACTERIA), PYRUVIC ACID | | Authors: | Oberholzer, A.E, Waltersperger, S.M, Schneider, P, Baumann, U, Erni, B. | | Deposit date: | 2010-11-24 | | Release date: | 2011-06-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.677 Å) | | Cite: | Phosphoenolpyruvate: Sugar Phosphotransferase System from the Hyperthermophilic Thermoanaerobacter Tengcongensis.
Biochemistry, 50, 2011
|
|
2XZ7
 
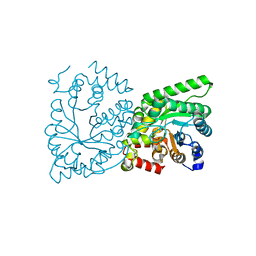 | | CRYSTAL STRUCTURE OF THE PHOSPHOENOLPYRUVATE-BINDING DOMAIN OF ENZYME I IN COMPLEX WITH PHOSPHOENOLPYRUVATE FROM THE THERMOANAEROBACTER TENGCONGENSIS PEP-SUGAR PHOSPHOTRANSFERASE SYSTEM (PTS) | | Descriptor: | MAGNESIUM ION, PHOSPHOENOLPYRUVATE, PHOSPHOENOLPYRUVATE-PROTEIN KINASE (PTS SYSTEM EI COMPONENT IN BACTERIA) | | Authors: | Waltersperger, S.M, Oberholzer, A.E, Schneider, P, Baumann, U, Erni, B. | | Deposit date: | 2010-11-23 | | Release date: | 2011-06-15 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Phosphoenolpyruvate: Sugar Phosphotransferase System from the Hyperthermophilic Thermoanaerobacter Tengcongensis.
Biochemistry, 50, 2011
|
|
1NRZ
 
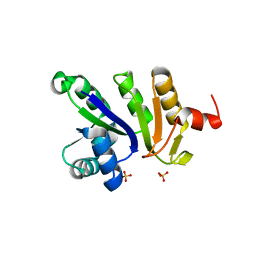 | |
1OI2
 
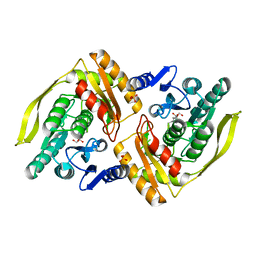 | | X-ray structure of the dihydroxyacetone kinase from Escherichia coli | | Descriptor: | GLYCEROL, HYPOTHETICAL PROTEIN YCGT, SULFATE ION | | Authors: | Siebold, C, Garcia-Alles, L.-F, Erni, B, Baumann, U. | | Deposit date: | 2003-06-04 | | Release date: | 2003-06-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | A Mechanism of Covalent Substrate Binding in the X-Ray Structure of Subunit K of the Escherichia Coli Dihydroxyacetone Kinase
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1OI3
 
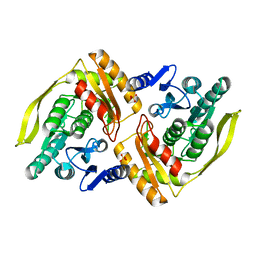 | | X-ray structure of the dihydroxyacetone kinase from Escherichia coli | | Descriptor: | HYPOTHETICAL PROTEIN YCGT, SULFATE ION | | Authors: | Siebold, C, Garcia-Alles, L.-F, Erni, B, Baumann, U. | | Deposit date: | 2003-06-04 | | Release date: | 2003-07-14 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A Mechanism of Covalent Substrate Binding in the X-Ray Structure of Subunit K of the Escherichia Coli Dihydroxyacetone Kinase
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
2IU5
 
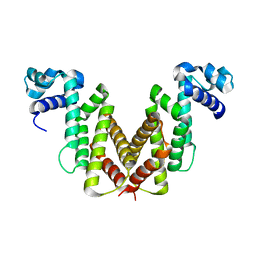 | | Dihydroxyacetone kinase operon activator DhaS | | Descriptor: | HTH-TYPE DHAKLM OPERON TRANSCRIPTIONAL ACTIVATOR DHAS | | Authors: | Srinivas, A, Christen, S, Baumann, U, Erni, B. | | Deposit date: | 2006-05-27 | | Release date: | 2006-06-13 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Regulation of the Dha Operon of Lactococcus Lactis: A Deviation from the Rule Followed by the Tetr Family of Transcription Regulators
J.Biol.Chem., 281, 2006
|
|
2IU4
 
 | | Dihydroxyacetone kinase operon co-activator Dha-DhaQ | | Descriptor: | DIHYDROXYACETONE KINASE, SULFATE ION | | Authors: | Srinivas, A, Christen, S, Baumann, U, Erni, B. | | Deposit date: | 2006-05-27 | | Release date: | 2006-06-13 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Regulation of the Dha Operon of Lactococcus Lactis: A Deviation from the Rule Followed by the Tetr Family of Transcription Regulators
J.Biol.Chem., 281, 2006
|
|
2IU6
 
 | | REGULATION OF THE DHA OPERON OF LACTOCOCCUS LACTIS | | Descriptor: | DIHYDROXYACETONE KINASE, GLYCEROL | | Authors: | Srinivas, A, Christen, S, Baumann, U, Erni, B. | | Deposit date: | 2006-05-27 | | Release date: | 2006-06-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Regulation of the Dha Operon of Lactococcus Lactis: A Deviation from the Rule Followed by the Tetr Family of Transcription Regulators
J.Biol.Chem., 281, 2006
|
|
1UOD
 
 | | Crystal structure of the dihydroxyacetone kinase from E. coli in complex with dihydroxyacetone-phosphate | | Descriptor: | DIHYDROXYACETONE KINASE, GLYCERALDEHYDE-3-PHOSPHATE, SULFATE ION | | Authors: | Siebold, C, Garcia-Alles, L.F, Luthi-Nyffeler, T, Flukiger-Bruhwiler, K, Burgi, H.-B, Baumann, U, Erni, B. | | Deposit date: | 2003-09-16 | | Release date: | 2004-09-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Phosphoenolpyruvate- and ATP-Dependent Dihydroxyacetone Kinases: Covalent Substrate-Binding and Kinetic Mechanism
Biochemistry, 43, 2004
|
|
1UN8
 
 | | Crystal structure of the dihydroxyacetone kinase of C. freundii (native form) | | Descriptor: | (2R)-3-(PHOSPHONOOXY)-2-(TETRADECANOYLOXY)PROPYL PALMITATE, DIHYDROXYACETONE KINASE | | Authors: | Siebold, C, Arnold, I, Garcia-Alles, L.F, Baumann, U, Erni, B. | | Deposit date: | 2003-09-08 | | Release date: | 2003-10-10 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of the Citrobacter Freundii Dihydroxyacetone Kinase Reveals an Eight-Stranded Alpha-Helical Barrel ATP-Binding Domain
J.Biol.Chem., 278, 2003
|
|
1UN9
 
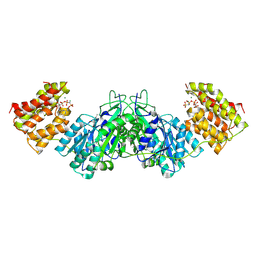 | | Crystal structure of the dihydroxyacetone kinase from C. freundii in complex with AMP-PNP and Mg2+ | | Descriptor: | DIHYDROXYACETONE, DIHYDROXYACETONE KINASE, MAGNESIUM ION, ... | | Authors: | Siebold, C, Arnold, I, Garcia-Alles, L.F, Baumann, U, Erni, B. | | Deposit date: | 2003-09-08 | | Release date: | 2003-10-30 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal Structure of the Citrobacter Freundii Dihydroxyacetone Kinase Reveals an Eight-Stranded Alpha-Helical Barrel ATP-Binding Domain
J.Biol.Chem., 278, 2003
|
|
1UOE
 
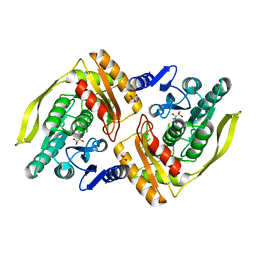 | | Crystal structure of the dihydroxyacetone kinase from E. coli in complex with glyceraldehyde | | Descriptor: | DIHYDROXYACETONE KINASE, GLYCEROL, SULFATE ION | | Authors: | Siebold, C, Garcia-Alles, L.F, Luthi-Nyffeler, T, Flukiger-Bruhwiler, K, Burgi, H.-B, Baumann, U, Erni, B. | | Deposit date: | 2003-09-16 | | Release date: | 2004-09-24 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Phosphoenolpyruvate- and ATP-Dependent Dihydroxyacetone Kinases: Covalent Substrate-Binding and Kinetic Mechanism
Biochemistry, 43, 2004
|
|
1BLE
 
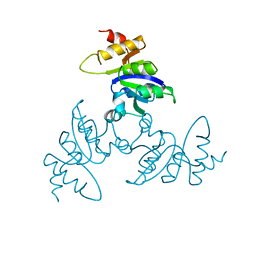 | |
3CT6
 
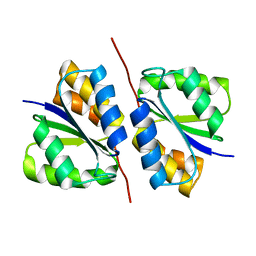 | | Crystal structure of DhaM of L. lactis | | Descriptor: | PTS-dependent dihydroxyacetone kinase, phosphotransferase subunit dhaM | | Authors: | Zurbriggen, A. | | Deposit date: | 2008-04-11 | | Release date: | 2008-10-21 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | X-ray Structures of the Three Lactococcus lactis Dihydroxyacetone Kinase Subunits and of a Transient Intersubunit Complex.
J.Biol.Chem., 283, 2008
|
|
