3TV0
 
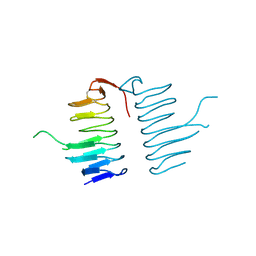 | |
4TGL
 
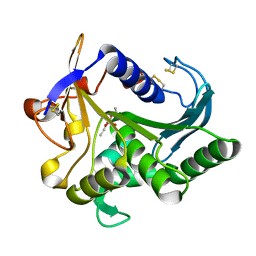 | | CATALYSIS AT THE INTERFACE: THE ANATOMY OF A CONFORMATIONAL CHANGE IN A TRIGLYCERIDE LIPASE | | Descriptor: | DIETHYL PHOSPHONATE, TRIACYL-GLYCEROL ACYLHYDROLASE | | Authors: | Derewenda, U, Brzozowski, A.M, Lawson, D, Derewenda, Z.S. | | Deposit date: | 1991-07-29 | | Release date: | 1993-07-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Catalysis at the interface: the anatomy of a conformational change in a triglyceride lipase.
Biochemistry, 31, 1992
|
|
6B6Z
 
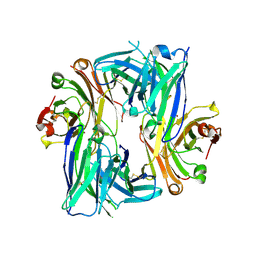 | | Crystal structure of the Apo Antibody fragment (Fab) raised against C-terminal domain of Ebola nucleoprotein (EBOV, TAFV, BDBV strains) | | Descriptor: | Apo Fab Heavy Chain, Apo Fab Light Chain, ZINC ION | | Authors: | Radwanska, M.J, Derewenda, U, Kossiakoff, A.A, Derewenda, Z.S. | | Deposit date: | 2017-10-03 | | Release date: | 2018-12-19 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.112 Å) | | Cite: | Crystal structure of the Apo Antibody fragment (Fab) raised against C-terminal domain of Ebola nucleoprotein (EBOV, TAFV, BDBV strains)
To Be Published
|
|
6TAA
 
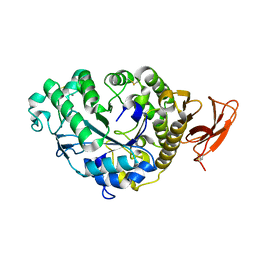 | | STRUCTURE AND MOLECULAR MODEL REFINEMENT OF ASPERGILLUS ORYZAE (TAKA) ALPHA-AMYLASE: AN APPLICATION OF THE SIMULATED-ANNEALING METHOD | | Descriptor: | ALPHA-AMYLASE, CALCIUM ION | | Authors: | Swift, H.J, Brady, L, Derewenda, Z.S, Dodson, E.J, Turkenburg, J.P, Wilkinson, A.J. | | Deposit date: | 1992-08-21 | | Release date: | 1993-10-31 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure and molecular model refinement of Aspergillus oryzae (TAKA) alpha-amylase: an application of the simulated-annealing method.
Acta Crystallogr.,Sect.B, 47, 1991
|
|
5E2X
 
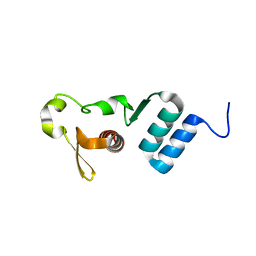 | | The crystal structure of the C-terminal domain of Ebola (Tai Forest) nucleoprotein | | Descriptor: | NONAETHYLENE GLYCOL, NP | | Authors: | Baker, L.E, Handing, K.B, Derewenda, U, Utepbergenov, D, Derewenda, Z.S. | | Deposit date: | 2015-10-01 | | Release date: | 2015-10-21 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Molecular architecture of the nucleoprotein C-terminal domain from the Ebola and Marburg viruses.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
5DSD
 
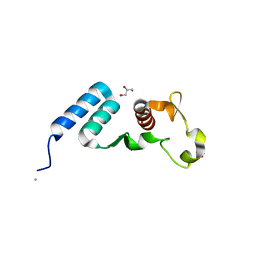 | | The crystal structure of the C-terminal domain of Ebola (Bundibugyo) nucleoprotein | | Descriptor: | CHLORIDE ION, GLYCEROL, Nucleoprotein | | Authors: | Baker, L, Handing, K.B, Utepbergenov, D, Derewenda, U, Derewenda, Z.S. | | Deposit date: | 2015-09-17 | | Release date: | 2015-09-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Molecular architecture of the nucleoprotein C-terminal domain from the Ebola and Marburg viruses.
Acta Crystallogr D Struct Biol, 72, 2016
|
|
2V71
 
 | | Coiled-coil region of NudEL | | Descriptor: | NUCLEAR DISTRIBUTION PROTEIN NUDE-LIKE 1 | | Authors: | Derewenda, U, Cooper, D.R, Kim, M.H, Derewenda, Z.S. | | Deposit date: | 2007-07-25 | | Release date: | 2007-11-27 | | Last modified: | 2017-06-28 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | The Structure of the Coiled-Coil Domain of Ndel1 and the Basis of its Interaction with Lis1, the Causal Protein of Miller-Dieker Lissencephaly.
Structure, 15, 2007
|
|
4QAZ
 
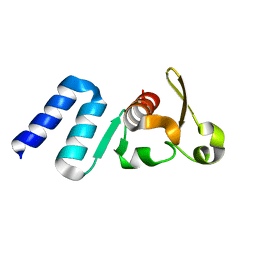 | |
4QB0
 
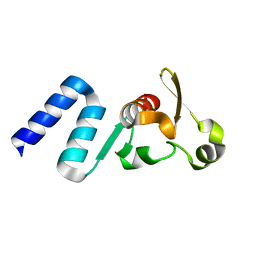 | |
4EL9
 
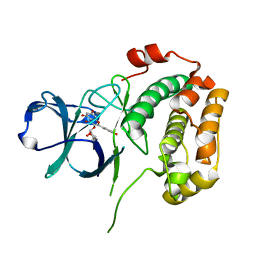 | | Structure of N-terminal kinase domain of RSK2 with afzelin | | Descriptor: | 5,7-dihydroxy-2-(4-hydroxyphenyl)-4-oxo-4H-chromen-3-yl 6-deoxy-alpha-L-mannopyranoside, Ribosomal protein S6 kinase alpha-3 | | Authors: | Utepbergenov, D, Derewenda, U, Derewenda, Z.S. | | Deposit date: | 2012-04-10 | | Release date: | 2012-09-05 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Insights into the Inhibition of the p90 Ribosomal S6 Kinase (RSK) by the Flavonol Glycoside SL0101 from the 1.5 A Crystal Structure of the N-Terminal Domain of RSK2 with Bound Inhibitor.
Biochemistry, 51, 2012
|
|
4GUE
 
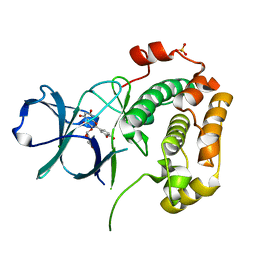 | | Structure of N-terminal kinase domain of RSK2 with flavonoid glycoside quercitrin | | Descriptor: | 2-(3,4-dihydroxyphenyl)-5,7-dihydroxy-4-oxo-4H-chromen-3-yl 6-deoxy-alpha-L-mannopyranoside, MAGNESIUM ION, Ribosomal protein S6 kinase alpha-3, ... | | Authors: | Derewenda, U, Utepbergenov, D, Szukalska, G, Derewenda, Z.S. | | Deposit date: | 2012-08-29 | | Release date: | 2013-01-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Identification of quercitrin as an inhibitor of the p90 S6 ribosomal kinase (RSK): structure of its complex with the N-terminal domain of RSK2 at 1.8 A resolution.
Acta Crystallogr.,Sect.D, 69, 2013
|
|
3TGL
 
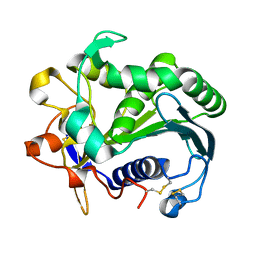 | | STRUCTURE AND MOLECULAR MODEL REFINEMENT OF RHIZOMUCOR MIEHEI TRIACYLGLYCERIDE LIPASE: A CASE STUDY OF THE USE OF SIMULATED ANNEALING IN PARTIAL MODEL REFINEMENT | | Descriptor: | TRIACYL-GLYCEROL ACYLHYDROLASE | | Authors: | Brady, L, Brzozowski, A.M, Derewenda, Z.S, Dodson, E.J, Dodson, G.G, Tolley, S.P, Turkenburg, J.P, Christiansen, L, Huge-Jensen, B, Norskov, L, Thim, L. | | Deposit date: | 1991-07-29 | | Release date: | 1993-07-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | STRUCTURE AND MOLECULAR-MODEL REFINEMENT OF RHIZOMUCOR-MIEHEI TRIACYLGLYCERIDE LIPASE - A CASE-STUDY OF THE USE OF SIMULATED ANNEALING IN PARTIAL MODEL REFINEMENT.
Acta Crystallogr.,Sect.B, 48, 1992
|
|
5TGL
 
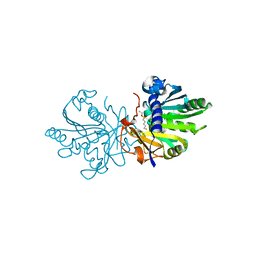 | | A MODEL FOR INTERFACIAL ACTIVATION IN LIPASES FROM THE STRUCTURE OF A FUNGAL LIPASE-INHIBITOR COMPLEX | | Descriptor: | LIPASE, N-HEXYLPHOSPHONATE ETHYL ESTER | | Authors: | Brzozowski, A.M, Derewenda, U, Derewenda, Z.S, Dodson, G.G, Lawson, D, Turkenburg, J.P, Bjorkling, F, Huge-Jensen, B, Patkar, S.R, Thim, L. | | Deposit date: | 1991-10-30 | | Release date: | 1994-01-31 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | A model for interfacial activation in lipases from the structure of a fungal lipase-inhibitor complex.
Nature, 351, 1991
|
|
3FMS
 
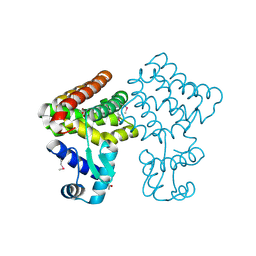 | | Crystal structure of TM0439, a GntR transcriptional regulator | | Descriptor: | ACETATE ION, NICKEL (II) ION, Transcriptional regulator, ... | | Authors: | Zheng, M, Cooper, D.R, Yu, M, Hung, L.-W, Derewenda, U, Derewenda, Z.S, Integrated Center for Structure and Function Innovation (ISFI) | | Deposit date: | 2008-12-22 | | Release date: | 2009-02-10 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of Thermotoga maritima TM0439: implications for the mechanism of bacterial GntR transcription regulators with Zn2+-binding FCD domains.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
3FHK
 
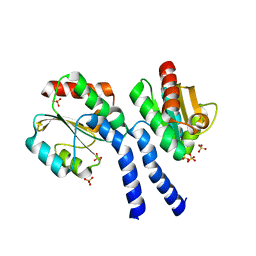 | | Crystal structure of APC1446, B.subtilis YphP disulfide isomerase | | Descriptor: | SULFATE ION, UPF0403 protein yphP | | Authors: | Derewenda, U, Boczek, T, Cooper, D.R, Yu, M, Hung, L, Derewenda, Z.S, Integrated Center for Structure and Function Innovation (ISFI) | | Deposit date: | 2008-12-09 | | Release date: | 2009-09-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure and function of Bacillus subtilis YphP, a prokaryotic disulfide isomerase with a CXC catalytic motif .
Biochemistry, 48, 2009
|
|
3MSX
 
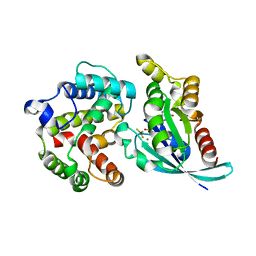 | | Crystal structure of RhoA.GDP.MgF3 in complex with GAP domain of ArhGAP20 | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Rho GTPase-activating protein 20, ... | | Authors: | Utepbergenov, D, Cooper, D.R, Derewenda, U, Derewenda, Z.S. | | Deposit date: | 2010-04-29 | | Release date: | 2011-03-16 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Mechanism of molecular specificity of RhoGAP domains towards small GTPases of RhoA family.
To be Published
|
|
3N77
 
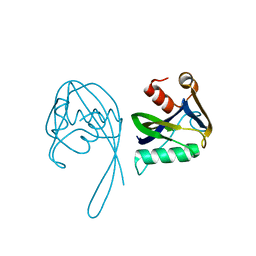 | | Crystal structure of Idp01880, putative NTP pyrophosphohydrolase of Salmonella typhimurium LT2 | | Descriptor: | Nucleoside triphosphatase nudI | | Authors: | Frydrysiak, J, Cooper, D.R, Derewenda, U, Anderson, W.F, Derewenda, Z.S, Integrated Center for Structure and Function Innovation (ISFI), Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2010-05-26 | | Release date: | 2010-08-18 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Crystal structure of Idp01880, putative NTP pyrophosphohydrolase of Salmonella typhimurium LT2
To be Published
|
|
3HD0
 
 | | Crystal structure of Tm1865, an Endonuclease V from Thermotoga Maritima | | Descriptor: | Endonuclease V | | Authors: | Utepbergenov, D, Cooper, D.R, Derewenda, U, Derewenda, Z.S, Integrated Center for Structure and Function Innovation (ISFI) | | Deposit date: | 2009-05-06 | | Release date: | 2009-07-14 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of Tm1865, an Endonuclease V from Thermotoga Maritima
To be Published
|
|
3H3E
 
 | | Crystal structure of Tm1679, A METAL-DEPENDENT HYDROLASE OF THE BETA-LACTAMASE SUPERFAMILY | | Descriptor: | ZINC ION, uncharacterized protein Tm1679 | | Authors: | Cooper, D.R, Olekhnovitch, N, Derewenda, U, Derewenda, Z.S, Integrated Center for Structure and Function Innovation (ISFI) | | Deposit date: | 2009-04-16 | | Release date: | 2009-07-14 | | Last modified: | 2021-10-13 | | Method: | X-RAY DIFFRACTION (2.749 Å) | | Cite: | Crystal structure of Tm1679, A METAL-DEPENDENT HYDROLASE OF THE BETA-LACTAMASE SUPERFAMILY
To be Published
|
|
5VIM
 
 | |
5VKD
 
 | | Crystal structure of C-terminal domain of Ebola (Bundibugyo) nucleoprotein in complex with Fab fragment | | Descriptor: | Fab Heavy Chain, Fab light chain, Nucleoprotein | | Authors: | Radwanska, M.J, Derewenda, U, Kossiakoff, A, Derewenda, Z.S. | | Deposit date: | 2017-04-21 | | Release date: | 2018-04-25 | | Last modified: | 2022-11-16 | | Method: | X-RAY DIFFRACTION (1.749 Å) | | Cite: | The structure of the C-terminal domain of the nucleoprotein from the Bundibugyo strain of the Ebola virus in complex with a pan-specific synthetic Fab.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
5VZ1
 
 | | Crystal structure of the Apo Antibody fragment (Fab) raised against C-terminal domain of Ebola nucleoprotein (EBOV, TAFV, BDBV strains) | | Descriptor: | Apo Antibody Fab Heavy Chain, Apo Antibody Fab Light Chain, ZINC ION | | Authors: | Radwanska, M.J, Derewenda, U, Kossiakoff, A, Derewenda, Z.S. | | Deposit date: | 2017-05-26 | | Release date: | 2018-06-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.112 Å) | | Cite: | Crystal structure of the Apo Antibody fragment (Fab) raised against C-terminal domain of Ebola nucleoprotein (EBOV, TAFV, BDBV strains)
To Be Published
|
|
5VI7
 
 | |
5W2B
 
 | | Crystal structure of C-terminal domain of Ebola (Reston) nucleoprotein in complex with Fab fragment | | Descriptor: | Fab Heavy Chain, Fab Light Chain, Nucleoprotein | | Authors: | Radwanska, M.J, Derewenda, U, Kossiakoff, A.A, Derewenda, Z.S. | | Deposit date: | 2017-06-06 | | Release date: | 2018-06-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | The structure of the C-terminal domain of the nucleoprotein from the Bundibugyo strain of the Ebola virus in complex with a pan-specific synthetic Fab.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
2JHY
 
 | |
