2K8V
 
 | | Solution structure of Oxidised ERp18 | | Descriptor: | Thioredoxin domain-containing protein 12 | | Authors: | Rowe, M.L, Alanen, H.I, Ruddock, L.W, Kelly, G, Schmidt, J.M, Williamson, R.A, Howard, M.J. | | Deposit date: | 2008-09-25 | | Release date: | 2009-06-02 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of ERp18, a small endoplasmic reticulum resident oxidoreductase .
Biochemistry, 48, 2009
|
|
6XNE
 
 | |
6XNL
 
 | |
6XNM
 
 | |
6XNF
 | |
7EP7
 
 | | The complex structure of Gpsm2 and Whirlin | | Descriptor: | G-protein-signaling modulator 2, Whirlin | | Authors: | Lin, L, Shi, Y, Wang, C, Zhu, J. | | Deposit date: | 2021-04-26 | | Release date: | 2022-05-04 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Promotion of row 1-specific tip complex condensates by Gpsm2-G alpha i provides insights into row identity of the tallest stereocilia.
Sci Adv, 8, 2022
|
|
5ITW
 
 | | Crystal structure of Bacillus subtilis BacC Dihydroanticapsin 7-dehydrogenase | | Descriptor: | Dihydroanticapsin 7-dehydrogenase, SULFATE ION | | Authors: | Perinbam, K, Balaram, H, Row, T.N.G, Gopal, B. | | Deposit date: | 2016-03-17 | | Release date: | 2017-02-22 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.19 Å) | | Cite: | Probing the influence of non-covalent contact networks identified by charge density analysis on the oxidoreductase BacC.
Protein Eng. Des. Sel., 30, 2017
|
|
5ITV
 
 | | Crystal structure of Bacillus subtilis BacC Dihydroanticapsin 7-dehydrogenase in complex with NADH | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, Dihydroanticapsin 7-dehydrogenase | | Authors: | Perinbam, K, Balaram, H, Row, T.N.G, Gopal, B. | | Deposit date: | 2016-03-17 | | Release date: | 2017-02-22 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Probing the influence of non-covalent contact networks identified by charge density analysis on the oxidoreductase BacC.
Protein Eng. Des. Sel., 30, 2017
|
|
5LO9
 
 | | Thiosulfate dehydrogenase (TsdBA) from Marichromatium purpuratum - "as isolated" form | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Cytochrome C, ... | | Authors: | Brito, J.A, Kurth, J.M, Reuter, J, Flegler, A, Koch, T, Franke, T, Klein, E, Rowe, S, Butt, J.N, Denkmann, K, Pereira, I.A.C, Dahl, C, Archer, M. | | Deposit date: | 2016-08-08 | | Release date: | 2016-10-12 | | Last modified: | 2017-09-06 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Electron Accepting Units of the Diheme Cytochrome c TsdA, a Bifunctional Thiosulfate Dehydrogenase/Tetrathionate Reductase.
J.Biol.Chem., 291, 2016
|
|
5DSB
 
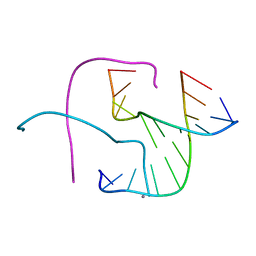 | | Crystal structure of Holliday junctions stabilized by 5-hydroxymethylcytosine in GCC junction core | | Descriptor: | 5'-D(*CP*CP*GP*GP*CP*GP*5HCP*CP*GP*G)-3', CALCIUM ION | | Authors: | Vander Zanden, C.M, Rowe, R.K, Broad, A.J, Ho, P.S. | | Deposit date: | 2015-09-17 | | Release date: | 2016-09-28 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.4959 Å) | | Cite: | Effect of Hydroxymethylcytosine on the Structure and Stability of Holliday Junctions.
Biochemistry, 55, 2016
|
|
2KMS
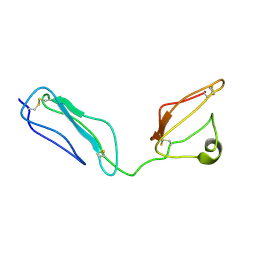 
 | | Combined high- and low-resolution techniques reveal compact structure in central portion of factor H despite long inter-modular linkers | | Descriptor: | Complement factor H | | Authors: | Schmidt, C.Q, Herbert, A.P, Guariento, M, Mertens, H.D.T, Soares, D.C, Uhrin, D, Rowe, A.J, Svergun, D.I, Barlow, P.N. | | Deposit date: | 2009-08-04 | | Release date: | 2009-11-03 | | Last modified: | 2020-02-26 | | Method: | SOLUTION NMR | | Cite: | The Central Portion of Factor H (Modules 10-15) Is Compact and Contains a Structurally Deviant CCP Module
J.Mol.Biol., 395, 2010
|
|
2L4C
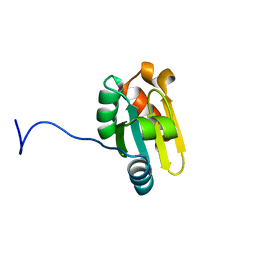 
 | | Solution Structure of the b domain of Human ERp27 | | Descriptor: | Endoplasmic reticulum resident protein 27 | | Authors: | Amin, N.T, Wallis, K, Rowe, M.L, Kelly, G, Frenkiel, T.A, Williamson, R.A, Howard, M.J, Freedman, R.B. | | Deposit date: | 2010-10-03 | | Release date: | 2010-11-10 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the b domain of human ERp27
To be Published
|
|
1W9R
 
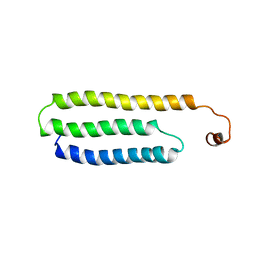 | | Solution Structure of Choline Binding Protein A, Domain R2, the Major Adhesin of Streptococcus pneumoniae | | Descriptor: | CHOLINE BINDING PROTEIN A | | Authors: | Luo, R, Mann, B, Lewis, W.S, Rowe, A, Heath, R, Stewart, M.L, Hamburger, A.E, Bjorkman, P.J, Sivakolundu, S, Lacy, E.R, Tuomanen, E, Kriwacki, R.W. | | Deposit date: | 2004-10-15 | | Release date: | 2005-02-22 | | Last modified: | 2011-07-13 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of Choline Binding Protein A, the Major Adhesin of Streptococcus Pneumoniae
Embo J., 24, 2005
|
|
4B20
 
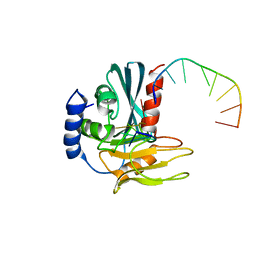 | | Structural basis of DNA loop recognition by Endonuclease V | | Descriptor: | 5'-D(*AP*TP*CP*TP*TP*GP*TP*CP*GP*CP)-3', 5'-D(*GP*CP*GP*AP*CP*AP*GP)-3', ENDONUCLEASE V, ... | | Authors: | Rosnes, I, Rowe, A.D, Forstrom, R.J, Alseth, I, Bjoras, M, Dalhus, B. | | Deposit date: | 2012-07-12 | | Release date: | 2013-04-17 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structural Basis of DNA Loop Recognition by Endonuclease V.
Structure, 21, 2013
|
|
3MKP
 
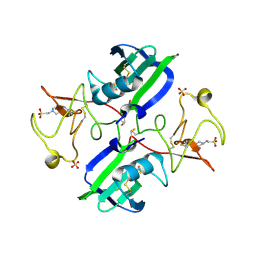 | | Crystal structure of 1K1 mutant of Hepatocyte Growth Factor/Scatter Factor fragment NK1 in complex with heparin | | Descriptor: | 2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose-(1-4)-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose, 2-deoxy-6-O-sulfo-2-(sulfoamino)-alpha-D-glucopyranose, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, ... | | Authors: | Gherardi, E, Chirgadze, D.Y, Blundell, T.L. | | Deposit date: | 2010-04-15 | | Release date: | 2010-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Engineering a fragment of Hepatocyte Growth Factor/Scatter Factor for tissue and organ regeneration
To be Published
|
|
1HFO
 
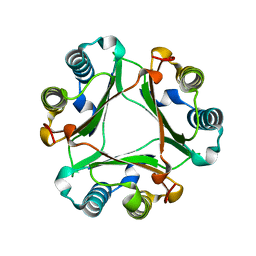 | |
5DSA
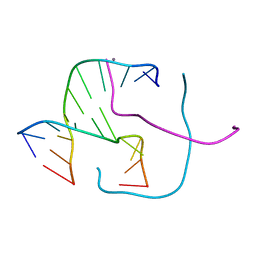 
 | |
4IRA
 
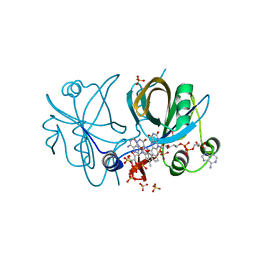 | | CobR in complex with FAD | | Descriptor: | 4-hydroxyphenylacetate 3-monooxygenase, FLAVIN-ADENINE DINUCLEOTIDE, SULFATE ION | | Authors: | Lawrence, A.D, Scott, A.F, Warren, M.J, Pickersgill, R.W. | | Deposit date: | 2013-01-14 | | Release date: | 2014-01-29 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Biophysical characterisation and structure-function analysis of Brucella melitensis CobR: Protein-flavin interactions determine function and stability
To be Published
|
|
5V7F
 
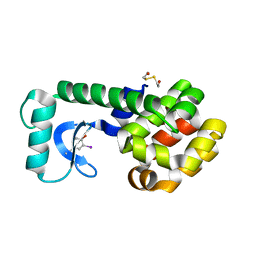 | | T4 lysozyme Y18Ymi | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, Lysozyme | | Authors: | Carlsson, A.-C.C. | | Deposit date: | 2017-03-20 | | Release date: | 2018-06-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Increasing Enzyme Stability and Activity through Hydrogen Bond-Enhanced Halogen Bonds.
Biochemistry, 57, 2018
|
|
5V7E
 
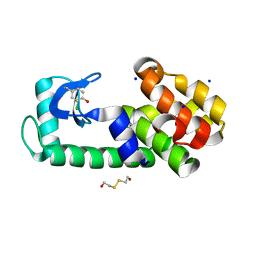 | | T4 lysozyme Y18Ymcl | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, Lysozyme, SODIUM ION | | Authors: | Carlsson, A.-C.C. | | Deposit date: | 2017-03-20 | | Release date: | 2018-06-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.36 Å) | | Cite: | Increasing Enzyme Stability and Activity through Hydrogen Bond-Enhanced Halogen Bonds.
Biochemistry, 57, 2018
|
|
5V7D
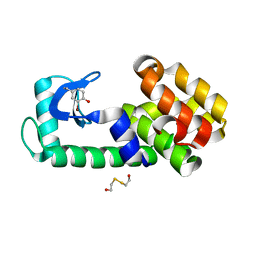 
 | | T4 lysozyme Y18Ymbr | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, Lysozyme | | Authors: | Carlsson, A.-C.C. | | Deposit date: | 2017-03-20 | | Release date: | 2018-06-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Increasing Enzyme Stability and Activity through Hydrogen Bond-Enhanced Halogen Bonds.
Biochemistry, 57, 2018
|
|
7M8J
 
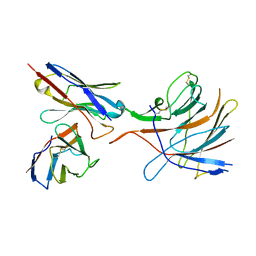 | | SARS-CoV-2 S-NTD + Fab CM25 | | Descriptor: | CM25 Fab - Heavy Chain, CM25 Fab - Light Chain, Spike protein S1 | | Authors: | Johnson, N.V, Mclellan, J.S. | | Deposit date: | 2021-03-29 | | Release date: | 2021-05-19 | | Last modified: | 2021-06-16 | | Method: | ELECTRON MICROSCOPY (3.48 Å) | | Cite: | Prevalent, protective, and convergent IgG recognition of SARS-CoV-2 non-RBD spike epitopes.
Science, 372, 2021
|
|
4PJ0
 
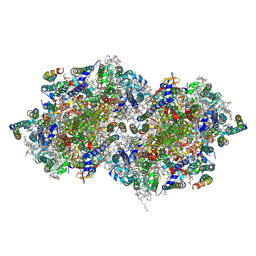 | | Structure of T.elongatus Photosystem II, rows of dimers crystal packing | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, ... | | Authors: | Hellmich, J, Bommer, M, Burkhardt, A, Ibrahim, M, Kern, J, Meents, A, Mueh, F, Dobbek, H, Zouni, A. | | Deposit date: | 2014-05-10 | | Release date: | 2014-10-22 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.437 Å) | | Cite: | Native-like Photosystem II Superstructure at 2.44 angstrom Resolution through Detergent Extraction from the Protein Crystal.
Structure, 22, 2014
|
|
6XHN
 
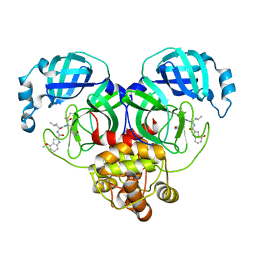 | | Covalent complex of SARS-CoV main protease with 4-methoxy-N-[(2S)-4-methyl-1-oxo-1-({(2S)-3-oxo-1-[(3S)-2-oxopyrrolidin-3-yl]butan-2-yl}amino)pentan-2-yl]-1H-indole-2-carboxamide | | Descriptor: | (3S)-3-{[N-(4-methoxy-1H-indole-2-carbonyl)-L-leucyl]amino}-2-oxo-4-[(3S)-2-oxopyrrolidin-3-yl]butyl 2-cyanobenzoate, 1,2-ETHANEDIOL, 3C-like proteinase | | Authors: | Gajiwala, K.S, Ferre, R.A, Ryan, K, Stewart, A.E. | | Deposit date: | 2020-06-19 | | Release date: | 2020-07-08 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.377 Å) | | Cite: | Discovery of Ketone-Based Covalent Inhibitors of Coronavirus 3CL Proteases for the Potential Therapeutic Treatment of COVID-19.
J.Med.Chem., 63, 2020
|
|
6XHM
 
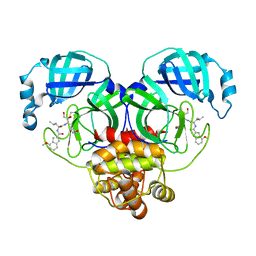 | | Covalent complex of SARS-CoV-2 main protease with N-[(2S)-1-({(2S,3S)-3,4-dihydroxy-1-[(3S)-2-oxopyrrolidin-3-yl]butan-2-yl}amino)-4-methyl-1-oxopentan-2-yl]-4-methoxy-1H-indole-2-carboxamide | | Descriptor: | 1,2-ETHANEDIOL, 3C-like proteinase, N-[(2S)-1-({(2S,3S)-3,4-dihydroxy-1-[(3S)-2-oxopyrrolidin-3-yl]butan-2-yl}amino)-4-methyl-1-oxopentan-2-yl]-4-methoxy-1H-indole-2-carboxamide | | Authors: | Gajiwala, K.S, Ferre, R.A, Ryan, K, Stewart, A.E. | | Deposit date: | 2020-06-19 | | Release date: | 2020-07-08 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.406 Å) | | Cite: | Discovery of Ketone-Based Covalent Inhibitors of Coronavirus 3CL Proteases for the Potential Therapeutic Treatment of COVID-19.
J.Med.Chem., 63, 2020
|
|
