8GUM
 
 | |
8GUL
 
 | |
8CC5
 
 | | Vibrio cholerae GbpA (LPMO domain) | | Descriptor: | ACETATE ION, COPPER (II) ION, GlcNAc-binding protein A, ... | | Authors: | Montserrat-Canals, M, Sorensen, H.V, Cordara, G, Krengel, U. | | Deposit date: | 2023-01-26 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Perdeuterated GbpA Enables Neutron Scattering Experiments of a Lytic Polysaccharide Monooxygenase.
Acs Omega, 8, 2023
|
|
8CC3
 
 | | Vibrio cholerae GbpA (LPMO domain) | | Descriptor: | ACETATE ION, COPPER (II) ION, GlcNAc-binding protein A, ... | | Authors: | Montserrat-Canals, M, Sorensen, H.V, Cordara, G, Krengel, U. | | Deposit date: | 2023-01-26 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.128 Å) | | Cite: | Perdeuterated GbpA Enables Neutron Scattering Experiments of a Lytic Polysaccharide Monooxygenase.
Acs Omega, 8, 2023
|
|
8C5N
 | | Sub-atomic resolution structure of the chitin-binding protein D (CbpD) from Pseudomonas aeruginosa | | Descriptor: | CHLORIDE ION, Chitin-binding protein CbpD | | Authors: | Cordara, G, Krengel, U, Golten, O, Vaaje-Kolstad, G, Vinther Soerensen, H. | | Deposit date: | 2023-01-09 | | Release date: | 2023-06-28 | | Last modified: | 2023-07-26 | | Method: | X-RAY DIFFRACTION (0.75 Å) | | Cite: | Immunization with lytic polysaccharide monooxygenase CbpD induces protective immunity against Pseudomonas aeruginosa pneumonia.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
7ZJB
 
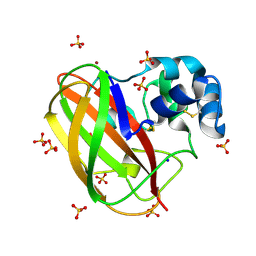 | | Structural and functional characterization of the bacterial lytic polysaccharide Monooxygenase ScLPMO10D | | Descriptor: | COPPER (II) ION, Putative secreted cellulose-binding protein, SODIUM ION, ... | | Authors: | Votvik, A.K, Rohr, A.K, Stepnov, A.A, Bissaro, B, Sorlie, M, Eijsink, V.G.H, Forsberg, Z. | | Deposit date: | 2022-04-10 | | Release date: | 2023-04-19 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.37 Å) | | Cite: | Structural and functional characterization of the catalytic domain of a cell-wall anchored bacterial lytic polysaccharide monooxygenase from Streptomyces coelicolor.
Sci Rep, 13, 2023
|
|
7ZE9
 
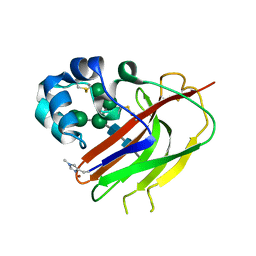 | | Structure of an AA16 LPMO-like protein | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, COPPER (II) ION, ... | | Authors: | Huang, Z, Banerjee, S, Muderspach, S.J, Sun, P, van Berkel, W.J.H, Kabel, M.A, Lo Leggio, L. | | Deposit date: | 2022-03-30 | | Release date: | 2023-03-15 | | Last modified: | 2023-04-26 | | Method: | X-RAY DIFFRACTION (2.646 Å) | | Cite: | AA16 Oxidoreductases Boost Cellulose-Active AA9 Lytic Polysaccharide Monooxygenases from Myceliophthora thermophila.
Acs Catalysis, 13, 2023
|
|
7SQX
 
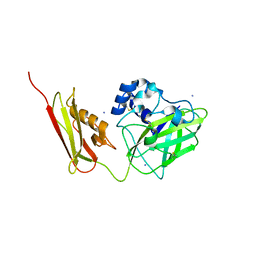 | | Crystal Structure of Pseudomonas aeruginosa lytic polysaccharide monooxygenase CbpD | | Descriptor: | AMMONIUM ION, Chitin-binding protein CbpD | | Authors: | Dade, C, Douzi, B, Ball, G, Voulhoux, R, Forest, K.T. | | Deposit date: | 2021-11-07 | | Release date: | 2022-07-20 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The crystal structure of CbpD clarifies substrate-specificity motifs in chitin-active lytic polysaccharide monooxygenases.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
7PB7
 | |
7PB6
 
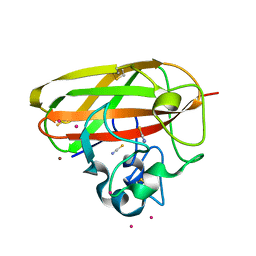 | |
7OKR
 
 | | Catalytic domain from the Aliivibrio salmonicida lytic polysaccharide monooxygenase AsLPMO10B | | Descriptor: | COPPER (II) ION, Chitinase B, DI(HYDROXYETHYL)ETHER | | Authors: | Cordara, G, Leitl, K.D, Loose, J.S.M, Vaaje-Kolstad, G, Krengel, U. | | Deposit date: | 2021-05-18 | | Release date: | 2022-06-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Chitinolytic enzymes contribute to the pathogenicity of Aliivibrio salmonicida LFI1238 in the invasive phase of cold-water vibriosis.
Bmc Microbiol., 22, 2022
|
|
6TWE
 
 | |
6TC4
 
 | | AA13 Lytic polysaccharide monooxygenase from Aspergillus oryzae measured with SSX | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, AoAA13, CHLORIDE ION, ... | | Authors: | Tandrup, T, Muderspach, S.J, Frandsen, K.E.H, Santoni, G, Poulsen, J.C.N, Lo Leggio, L. | | Deposit date: | 2019-11-05 | | Release date: | 2020-03-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Further structural studies of the lytic polysaccharide monooxygenase AoAA13 belonging to the starch-active AA13 family
Amylase, 3(1), 2019
|
|
6TBR
 
 | | Glycosylated AA13 Lytic polysaccharide monooxygenase from Aspergillus oryzae in P1 space group | | Descriptor: | AoAA13, ZINC ION | | Authors: | Frandsen, K.E.H, Muderspach, S.J, Tandrup, T, Poulsen, J.C.N, Lo Leggio, L. | | Deposit date: | 2019-11-04 | | Release date: | 2020-03-18 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Further structural studies of the lytic polysaccharide monooxygenase AoAA13 belonging to the starch-active AA13 family
Amylase, 3(1), 2019
|
|
6TBQ
 
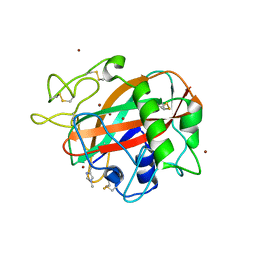 | | AA13 Lytic polysaccharide monooxygenase from Aspergillus oryzae partially in Cu(II) state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, AoAA13, COPPER (II) ION, ... | | Authors: | Muderspach, S.J, Lo Leggio, L, Tandrup, T, Frandsen, K.E.H, Poulsen, J.C.N. | | Deposit date: | 2019-11-04 | | Release date: | 2020-03-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Further structural studies of the lytic polysaccharide monooxygenase AoAA13 belonging to the starch-active AA13 family
Amylase, 3(1), 2019
|
|
6T5Z
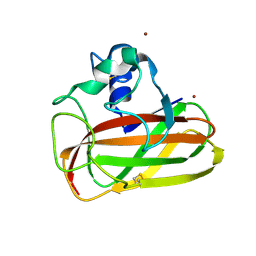 
 | | Crystal structure of an AA10 LPMO from Photorhabdus luminescens | | Descriptor: | COPPER (II) ION, Chitin-binding type-4 domain-containing protein | | Authors: | Munzone, A, El Kerdi, B, Reglier, M, Royant, A, Simaan, A.J, Decroos, C. | | Deposit date: | 2019-10-17 | | Release date: | 2020-01-15 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.60000312 Å) | | Cite: | Characterization of a bacterial copper-dependent lytic polysaccharide monooxygenase with an unusual second coordination sphere.
Febs J., 287, 2020
|
|
6RW7
 
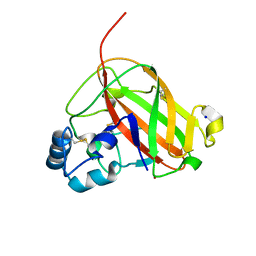 | | An AA10 LPMO from the shipworm symbiont Teredinibacter turnerae | | Descriptor: | COPPER (I) ION, COPPER (II) ION, Carbohydrate binding module family 33 and 10 domain protein, ... | | Authors: | Fowler, C.A, Sabbadin, F, Ciano, L, Hemsworth, G.R, Elias, L, Bruce, N.C, McQueen-Mason, S, Walton, P, Davies, G.J. | | Deposit date: | 2019-06-04 | | Release date: | 2019-10-09 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Discovery, activity and characterisation of an AA10 lytic polysaccharide oxygenase from the shipworm symbiontTeredinibacter turnerae.
Biotechnol Biofuels, 12, 2019
|
|
6IF7
 
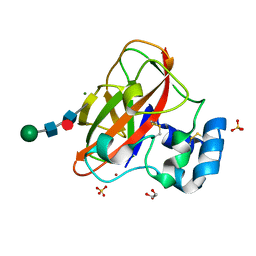 | | Crystal Structure of AA10 Lytic Polysaccharide Monooxygenase from Tectaria macrodonta | | Descriptor: | COPPER (II) ION, Chitin binding protein, GLYCEROL, ... | | Authors: | Archana, A, Yadav, S.K, Singh, P.K, Vasudev, P.G. | | Deposit date: | 2018-09-18 | | Release date: | 2019-04-24 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Insecticidal fern protein Tma12 is possibly a lytic polysaccharide monooxygenase.
Planta, 249, 2019
|
|
5WSZ
 
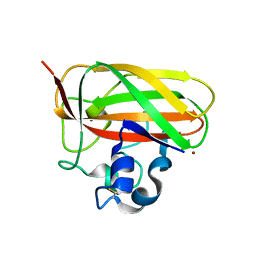 | | Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis | | Descriptor: | COPPER (II) ION, LpmO10A | | Authors: | Zhao, Y, Zhang, H, Yin, H. | | Deposit date: | 2016-12-08 | | Release date: | 2017-02-22 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.565 Å) | | Cite: | Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis
To Be Published
|
|
5VG1
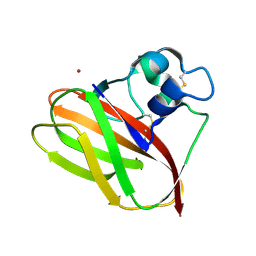 
 | |
5VG0
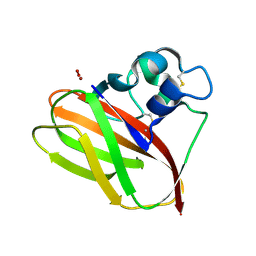 
 | |
5UIZ
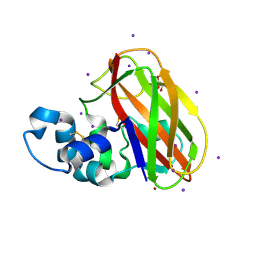 
 | | Structure of T.fusca AA10A | | Descriptor: | AA10A, COPPER (II) ION, GLYCEROL, ... | | Authors: | Alahuhta, P.M, Lunin, V.V. | | Deposit date: | 2017-01-16 | | Release date: | 2017-02-01 | | Last modified: | 2017-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of a Thermobifida fusca lytic polysaccharide monooxygenase and mutagenesis of key residues.
Biotechnol Biofuels, 10, 2017
|
|
5T7N
 
 | | X-ray crystal structure of AA13 LPMO | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, AoAA13, ZINC ION, ... | | Authors: | Frandsen, K.E.H, Poulsen, J.-C.N, Tovborg, M, Johansen, K.S, Lo Leggio, L. | | Deposit date: | 2016-09-05 | | Release date: | 2017-01-11 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Learning from oligosaccharide soaks of crystals of an AA13 lytic polysaccharide monooxygenase: crystal packing, ligand binding and active-site disorder.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
5T7K
 
 | | X-ray crystal structure of AA13 LPMO | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, AoAA13, ZINC ION | | Authors: | Frandsen, K.E.H, Poulsen, J.-C.N, Tovborg, M, Johansen, K.S, Lo Leggio, L. | | Deposit date: | 2016-09-05 | | Release date: | 2017-01-18 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Learning from oligosaccharide soaks of crystals of an AA13 lytic polysaccharide monooxygenase: crystal packing, ligand binding and active-site disorder.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
5T7J
 
 | | X-ray crystal structure of AA13 LPMO | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, AoAA13, ZINC ION | | Authors: | Frandsen, K.E.H, Poulsen, J.-C.N, Tovborg, M, Johansen, K.S, Lo Leggio, L. | | Deposit date: | 2016-09-05 | | Release date: | 2017-01-18 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Learning from oligosaccharide soaks of crystals of an AA13 lytic polysaccharide monooxygenase: crystal packing, ligand binding and active-site disorder.
Acta Crystallogr D Struct Biol, 73, 2017
|
|
