1HL2
 
 | |
8GEK
 
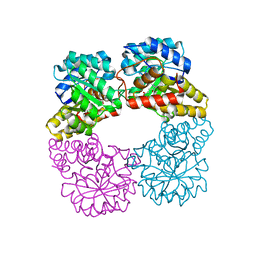 | | Dihydrodipicolinate synthase with pyruvate from Candidatus Liberibacter solanacearum | | Descriptor: | 4-hydroxy-tetrahydrodipicolinate synthase | | Authors: | Gilkes, J.M, Frampton, R.A, Board, A, Sheen, C.R, Smith, G.R, Dobson, R.C.J. | | Deposit date: | 2023-03-06 | | Release date: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Dihydrodipicolinate synthase with pyruvate from the plant pathogen, Candidatus Liberibacter solanacearum
To Be Published
|
|
2HMC
 
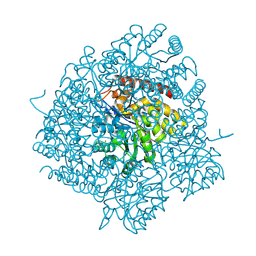 | | The Crystal Structure of Dihydrodipicolinate Synthase DapA from Agrobacterium tumefaciens | | Descriptor: | Dihydrodipicolinate synthase, MAGNESIUM ION | | Authors: | Kim, Y, Zhang, R, Xu, X, Zheng, H, Savchenko, A, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2006-07-11 | | Release date: | 2006-09-05 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The Crystal Structure of Dihydrodipicolinate Synthase DapA from Agrobacterium tumefaciens
To be Published, 2006
|
|
8DO5
 
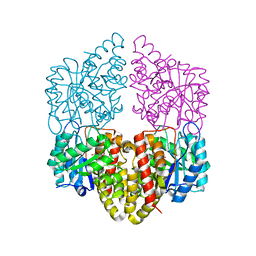 | | Crystal structure of NahE in complex with intermediate (R)-4-hydroxy-4-(2-hydroxyphenyl)-2-iminobutanoate | | Descriptor: | (4R)-4-hydroxy-4-(2-hydroxyphenyl)butanoic acid, DIMETHYL SULFOXIDE, Trans-ohydrobenzylidenepyruvate hydratase aldolase | | Authors: | LeVieux, J.A, Hardtke, H.A, Zhang, Y.J. | | Deposit date: | 2022-07-12 | | Release date: | 2022-12-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A mutagenic analysis of NahE, a hydratase-aldolase in the naphthalene degradative pathway.
Arch.Biochem.Biophys., 733, 2023
|
|
6MQH
 
 | |
6NVA
 
 | |
4R53
 
 | |
6P90
 
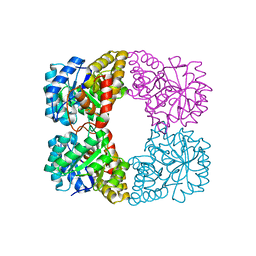
 | | Crystal structure of PaDHDPS2-H56Q mutant | | Descriptor: | 4-hydroxy-tetrahydrodipicolinate synthase, CHLORIDE ION, GLYCEROL | | Authors: | Impey, R.E, Panjikar, S, Hall, C.J, Bock, L.J, Sutton, J.M, Perugini, M.A, Soares da Costa, T.P. | | Deposit date: | 2019-06-08 | | Release date: | 2019-08-07 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification of two dihydrodipicolinate synthase isoforms from Pseudomonas aeruginosa that differ in allosteric regulation.
Febs J., 287, 2020
|
|
4U4M
 | | Crystal structure of 0.5M urea unfolded YagE, a KDG aldolase protein in complex with Pyruvate | | Descriptor: | 1,2-ETHANEDIOL, PYRUVIC ACID, UREA, ... | | Authors: | Manoj Kumar, P, Bhaskar, V, Manicka, S, Krishnaswamy, S. | | Deposit date: | 2014-07-24 | | Release date: | 2015-07-29 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.09 Å) | | Cite: | Crystal structure of 0.5M urea unfolded YagE, a KDG aldolase protein in complex with Pyruvate
To be published
|
|
7M06
 | | Dihydrodipicolinate synthase (DHDPS) from C.jejuni, Y110F mutant with R,R-bislysine bound at the allosteric site at 2.7 Angstrom | | Descriptor: | (2R,5R)-2,5-diamino-2,5-bis(4-aminobutyl)hexanedioic acid, 1,2-ETHANEDIOL, 4-hydroxy-tetrahydrodipicolinate synthase, ... | | Authors: | Saran, S, Sanders, D.A.R. | | Deposit date: | 2021-03-10 | | Release date: | 2022-03-16 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | B-FACTOR ANALYSIS SUGGEST THAT L-LYSINE AND R, R-BISLYSINE ALLOSTERICALLY INHIBIT Cj.DHDPS ENZYME BY DECREASING PROTEIN DYNAMICS
To Be Published
|
|
7MJF
 
 | | Crystal structure of Candidatus Liberibacter solanacearum dihydrodipicolinate synthase with pyruvate and succinic semi-aldehyde bound in active site | | Descriptor: | (4R)-4-oxidanyl-2-oxidanylidene-heptanedioic acid, (4S)-4-hydroxy-2-oxoheptanedioic acid, 4-hydroxy-tetrahydrodipicolinate synthase | | Authors: | Gilkes, J, Frampton, R.A, Board, A.J, Sheen, C.R, Smith, G.R, Dobson, R.C.J. | | Deposit date: | 2021-04-20 | | Release date: | 2021-07-14 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of Candidatus Liberibacter solanacearum dihydrodipicolinate synthase with pyruvate and succinic semi-aldehyde bound in active site
To Be Published
|
|
7MDS
 
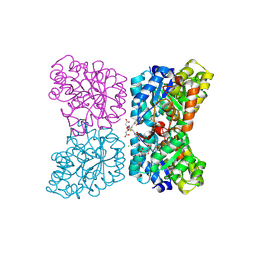 | | Crystal structure of AtDHDPS1 in complex with MBDTA-2 | | Descriptor: | 4-hydroxy-tetrahydrodipicolinate synthase 1, chloroplastic, CHLORIDE ION, ... | | Authors: | Hall, C.J, Soares da Costa, T.P, Panjikar, S. | | Deposit date: | 2021-04-06 | | Release date: | 2021-08-04 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.295 Å) | | Cite: | Towards novel herbicide modes of action by inhibiting lysine biosynthesis in plants.
Elife, 10, 2021
|
|
5T25
 
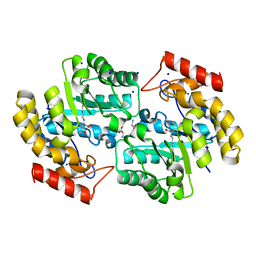 | | Kinetic, Spectral and Structural Characterization of the Slow Binding Inhibitor Acetopyruvate with Dihydrodipicolinate Synthase from Escherichia coli. | | Descriptor: | 4-hydroxy-tetrahydrodipicolinate synthase, LYSINE, SODIUM ION | | Authors: | Chooback, L, Thomas, L.M, Karsten, W.E, Fleming, C.D, Seabourn, P. | | Deposit date: | 2016-08-23 | | Release date: | 2017-03-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.991 Å) | | Cite: | Kinetic, Spectral and Structural Characterization of the Slow Binding Inhibitor Acetopyruvate with Dihydrodipicolinate Synthase from Escherichia coli.
To Be Published
|
|
7C0D
 | |
5T26
 
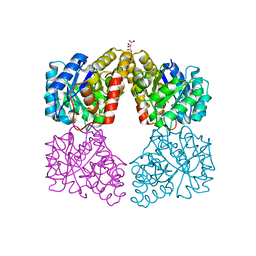 | | Kinetic, Spectral and Structural Characterization of the Slow Binding Inhibitor Acetopyruvate with Dihydrodipicolinate Synthase from Escherichia coli. | | Descriptor: | 4-hydroxy-tetrahydrodipicolinate synthase, GLYCEROL, L(+)-TARTARIC ACID, ... | | Authors: | Chooback, L, Thomas, L.M, Karsten, W.E, Fleming, C.D, Seabourn, P. | | Deposit date: | 2016-08-23 | | Release date: | 2016-10-05 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Kinetic, Spectral and Structural Characterization of the Slow Binding Inhibitor Acetopyruvate with Dihydrodipicolinate Synthase from Escherichia coli.
To Be Published
|
|
7C0E
 
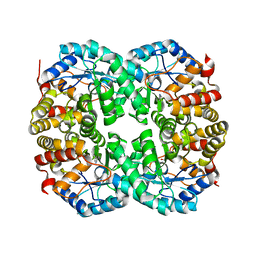 | |
4XKY
 
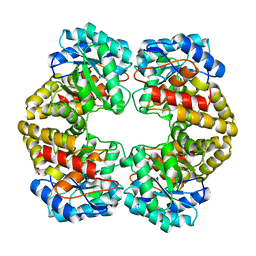 | | Structure of dihydrodipicolinate synthase from the commensal bacterium Bacteroides thetaiotaomicron at 2.1 A resolution | | Descriptor: | ACETATE ION, Dihydrodipicolinate synthase, SODIUM ION | | Authors: | Mank, N.J, Arnette, A.K, Klapper, V.G, Chruszcz, M. | | Deposit date: | 2015-01-12 | | Release date: | 2015-04-01 | | Last modified: | 2015-04-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of dihydrodipicolinate synthase from the commensal bacterium Bacteroides thetaiotaomicron at 2.1 angstrom resolution.
Acta Crystallogr.,Sect.F, 71, 2015
|
|
7C0C
 | |
5ZJM
 
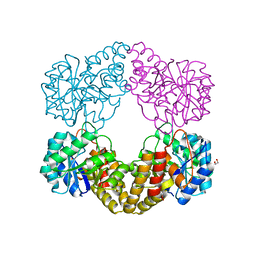 | | Crystal structure of N-acetylneuraminate lyase from Fusobacterium nucleatum | | Descriptor: | 1,2-ETHANEDIOL, N-acetylneuraminate lyase | | Authors: | Kumar, J.P, Rao, H, Nayak, V, Subramanian, R. | | Deposit date: | 2018-03-21 | | Release date: | 2019-01-23 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.323 Å) | | Cite: | Crystal structures and kinetics of N-acetylneuraminate lyase from Fusobacterium nucleatum
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
5ZKA
 
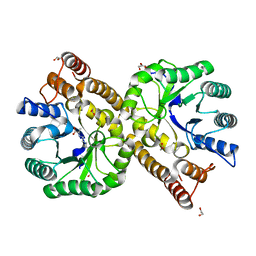 | | Crystal structure of N-acetylneuraminate lyase from Fusobacterium nucleatum complexed with Pyruvate | | Descriptor: | 1,2-ETHANEDIOL, N-acetylneuraminate lyase, TRIETHYLENE GLYCOL | | Authors: | Kumar, J.P, Rao, H, Nayak, V, Ramaswamy, S. | | Deposit date: | 2018-03-23 | | Release date: | 2018-12-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Crystal structures and kinetics of N-acetylneuraminate lyase from Fusobacterium nucleatum
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
5UD6
 | |
6ARH
 
 | |
5AFD
 
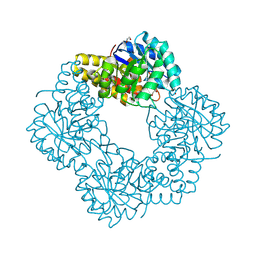 | | Native structure of N-acetylneuramininate lyase (sialic acid aldolase) from Aliivibrio salmonicida | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, N-ACETYLNEURAMINATE LYASE | | Authors: | Gurung, M.K, Altermark, B, Rader, I.L.U, Helland, R, Smalas, A.O. | | Deposit date: | 2015-01-21 | | Release date: | 2016-03-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Features and structure of a cold active N-acetylneuraminate lyase.
Plos One, 14, 2019
|
|
5A8G
 
 | | Crystal structure of the wild-type Staphylococcus aureus N- acetylneurminic acid lyase in complex with fluoropyruvate | | Descriptor: | N-ACETYLNEURAMINATE LYASE | | Authors: | Stockwell, J, Daniels, A.D, Windle, C.L, Harman, T, Woodhall, T, Trinh, C.H, Lebel, T, Pearson, A.R, Mulholland, K, Berry, A, Nelson, A. | | Deposit date: | 2015-07-15 | | Release date: | 2015-11-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Evaluation of Fluoropyruvate as Nucleophile in Reactions Catalysed by N-Acetyl Neuraminic Acid Lyase Variants: Scope, Limitations and Stereoselectivity.
Org.Biomol.Chem., 14, 2016
|
|
5KZE
 | | N-acetylneuraminate lyase from methicillin-resistant Staphylococcus aureus | | Descriptor: | GLYCEROL, N-acetylneuraminate lyase, SULFATE ION | | Authors: | North, R.A, Watson, A.J.A, Pearce, F.G, Muscroft-Taylor, A.C, Friemann, R, Fairbanks, A.J, Dobson, R.C.J. | | Deposit date: | 2016-07-25 | | Release date: | 2017-01-11 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Structure and inhibition of N-acetylneuraminate lyase from methicillin-resistant Staphylococcus aureus.
FEBS Lett., 590, 2016
|
|
