[English] 日本語
 Yorodumi
Yorodumi- PDB-487d: SEVEN RIBOSOMAL PROTEINS FITTED TO A CRYO-ELECTRON MICROSCOPIC MA... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 487d | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
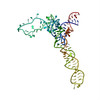
| Title | SEVEN RIBOSOMAL PROTEINS FITTED TO A CRYO-ELECTRON MICROSCOPIC MAP OF THE LARGE 50S SUBUNIT AT 7.5 ANGSTROMS RESOLUTION | |||||||||
 Components Components | (50S ribosomal protein ...) x 7 | |||||||||
 Keywords Keywords | RIBOSOME / LARGE RIBOSOMAL SUBUNIT / RIBOSOMAL PROTEIN / PROTEIN BIOSYNTHESIS / EM-RECONSTRUCTION / ATOMIC STRUCTURE / 3D ARRANGEMENT / FITTING | |||||||||
| Function / homology |  Function and homology information Function and homology informationribosomal large subunit assembly / response to radiation / regulation of translation / transferase activity / 5S rRNA binding / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / rRNA binding ...ribosomal large subunit assembly / response to radiation / regulation of translation / transferase activity / 5S rRNA binding / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / rRNA binding / negative regulation of translation / ribosome / structural constituent of ribosome / translation / ribonucleoprotein complex / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   Thermus thermophilus (bacteria) Thermus thermophilus (bacteria)  Geobacillus stearothermophilus (bacteria) Geobacillus stearothermophilus (bacteria)  Thermotoga maritima (bacteria) Thermotoga maritima (bacteria) | |||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 7.5 Å | |||||||||
 Authors Authors | Brimacombe, R. / Mueller, F. | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2000 Journal: J Mol Biol / Year: 2000Title: The 3D arrangement of the 23 S and 5 S rRNA in the Escherichia coli 50 S ribosomal subunit based on a cryo-electron microscopic reconstruction at 7.5 A resolution. Authors: F Mueller / I Sommer / P Baranov / R Matadeen / M Stoldt / J Wöhnert / M Görlach / M van Heel / R Brimacombe /  Abstract: The Escherichia coli 23 S and 5 S rRNA molecules have been fitted helix by helix to a cryo-electron microscopic (EM) reconstruction of the 50 S ribosomal subunit, using an unfiltered version of the ...The Escherichia coli 23 S and 5 S rRNA molecules have been fitted helix by helix to a cryo-electron microscopic (EM) reconstruction of the 50 S ribosomal subunit, using an unfiltered version of the recently published 50 S reconstruction at 7.5 A resolution. At this resolution, the EM density shows a well-defined network of fine structural elements, in which the major and minor grooves of the rRNA helices can be discerned at many locations. The 3D folding of the rRNA molecules within this EM density is constrained by their well-established secondary structures, and further constraints are provided by intra and inter-rRNA crosslinking data, as well as by tertiary interactions and pseudoknots. RNA-protein cross-link and foot-print sites on the 23 S and 5 S rRNA were used to position the rRNA elements concerned in relation to the known arrangement of the ribosomal proteins as determined by immuno-electron microscopy. The published X-ray or NMR structures of seven 50 S ribosomal proteins or RNA-protein complexes were incorporated into the EM density. The 3D locations of cross-link and foot-print sites to the 23 S rRNA from tRNA bound to the ribosomal A, P or E sites were correlated with the positions of the tRNA molecules directly observed in earlier reconstructions of the 70 S ribosome at 13 A or 20 A. Similarly, the positions of cross-link sites within the peptidyl transferase ring of the 23 S rRNA from the aminoacyl residue of tRNA were correlated with the locations of the CCA ends of the A and P site tRNA. Sites on the 23 S rRNA that are cross-linked to the N termini of peptides of different lengths were all found to lie within or close to the internal tunnel connecting the peptidyl transferase region with the presumed peptide exit site on the solvent side of the 50 S subunit. The post-transcriptionally modified bases in the 23 S rRNA form a cluster close to the peptidyl transferase area. The minimum conserved core elements of the secondary structure of the 23 S rRNA form a compact block within the 3D structure and, conversely, the points corresponding to the locations of expansion segments in 28 S rRNA all lie on the outside of the structure. #1:  Journal: Cell(Cambridge,Mass.) / Year: 1999 Journal: Cell(Cambridge,Mass.) / Year: 1999Title: A Detailed View of a Ribosomal Active Site: The Structure of the Gtpase Center at 2.6 Angstroms Resolution. Authors: Wimberly, B.T. / Guymon, R. / Mc Cutcheon, J.P. / White, S. / Ramakrishnan, V. #2:  Journal: Embo J. / Year: 1999 Journal: Embo J. / Year: 1999Title: The NMR Structure of the 5S Rrna E-Domain-Protein C25 Complex Shows Pre-Formed and Induced Recognition. Authors: Stoldt, M. / Woehnert, J. / Ohlenschlaeger, O. / Goerlach, M. / Brown, L.R. #3:  Journal: Embo J. / Year: 1999 Journal: Embo J. / Year: 1999Title: The Three-Dimensional Structure of the RNA-Binding Domain of Ribosomal Protein L2; a Protein at the Peptidyl Transferase Center of the Ribosome. Authors: Nakagawa, A. / Nakashima, T. / Taniguchi, M. / Hosaka, H. / Kimura, M. / Tanaka, I. #4:  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: Ribosomal Protein L9: A Structure Determination by the Combined Use of X-Ray Crystallography and NMR Spectroscopy. Authors: Hoffman, D.W. / Cameron, C.S. / Davies, C. / White, S.W. / Ramakrishnan, V. #5:  Journal: Embo J. / Year: 1996 Journal: Embo J. / Year: 1996Title: Crystal Structure of the RNA Binding Ribosomal Protein L1 from Thermus Thermophilus. Authors: Nikonov, S. / Nevskaya, N. / Eliseikina, I. / Fomenkova, N. / Nikulin, A. / Ossina, N. / Garber, M. / Jonsson, B.H. / Briand, C. / Al-Karadaghi, S. / Svensson, A. / Aevarsson, A. / Liljas, A. #6:  Journal: Structure / Year: 1996 Journal: Structure / Year: 1996Title: The Crystal Structure of Ribosomal Protein L14 Reveals an Important Organizational Component of the Translational Apparatus. Authors: Davies, C. / White, S.W. / Ramakrishnan, V. #7:  Journal: Embo J. / Year: 1993 Journal: Embo J. / Year: 1993Title: Ribosomal Protein L6: Structural Evidence of Gene Duplication from a Primitive RNA Binding Proetin. Authors: Golden, B.L. / Ramakrishnan, V. / White, S.W. #8:  Journal: Structure / Year: 1999 Journal: Structure / Year: 1999Title: The Escherichia coli large ribosomal subunit at 7.5 A resolution Authors: Matadeen, R. / Patwardhan, A. / Gowen, B. / Orlova, E.V. / Pape, T. / Cuff, M. / Mueller, F. / Brimacombe, R. / van Heel, M. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  487d.cif.gz 487d.cif.gz | 224 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb487d.ent.gz pdb487d.ent.gz | 176.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  487d.json.gz 487d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  487d_validation.pdf.gz 487d_validation.pdf.gz | 396.8 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  487d_full_validation.pdf.gz 487d_full_validation.pdf.gz | 439.5 KB | Display | |
| Data in XML |  487d_validation.xml.gz 487d_validation.xml.gz | 25.7 KB | Display | |
| Data in CIF |  487d_validation.cif.gz 487d_validation.cif.gz | 38.8 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/87/487d https://data.pdbj.org/pub/pdb/validation_reports/87/487d ftp://data.pdbj.org/pub/pdb/validation_reports/87/487d ftp://data.pdbj.org/pub/pdb/validation_reports/87/487d | HTTPS FTP |
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-50S ribosomal protein ... , 7 types, 7 molecules HIJKLMN
| #1: Protein | Mass: 24331.074 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Thermus thermophilus (bacteria) / References: UniProt: P27150, UniProt: Q5SLP7*PLUS Thermus thermophilus (bacteria) / References: UniProt: P27150, UniProt: Q5SLP7*PLUS |
|---|---|
| #2: Protein | Mass: 14759.666 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Geobacillus stearothermophilus (bacteria) / References: UniProt: P04257 Geobacillus stearothermophilus (bacteria) / References: UniProt: P04257 |
| #3: Protein | Mass: 17811.447 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Geobacillus stearothermophilus (bacteria) / References: UniProt: P02391 Geobacillus stearothermophilus (bacteria) / References: UniProt: P02391 |
| #4: Protein | Mass: 16341.037 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Geobacillus stearothermophilus (bacteria) / References: UniProt: P02417 Geobacillus stearothermophilus (bacteria) / References: UniProt: P02417 |
| #5: Protein | Mass: 14294.913 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Thermotoga maritima (bacteria) / References: UniProt: P29395 Thermotoga maritima (bacteria) / References: UniProt: P29395 |
| #6: Protein | Mass: 13369.613 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   Geobacillus stearothermophilus (bacteria) / References: UniProt: P04450 Geobacillus stearothermophilus (bacteria) / References: UniProt: P04450 |
| #7: Protein | Mass: 10713.465 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: LARGE 50S RIBOSOMAL SUBUNIT / Type: RIBOSOME |
|---|---|
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Crystal | Description: THE CRYST1 AND SCALE RECORDS ARE MEANINGLESS. |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: FEI/PHILIPS CM200FEG / Details: from Structure 1999 citation |
|---|---|
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 38000 X / Nominal defocus max: 2000 nm / Nominal defocus min: 1000 nm |
| Specimen holder | Specimen holder model: GATAN LIQUID NITROGEN |
| Image recording | Film or detector model: GENERIC FILM |
| Image scans | Scanner model: IMAGE SCIENCE PATCHWORK DENSITOMETER |
- Processing
Processing
| Symmetry | Point symmetry: C1 (asymmetric) | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3D reconstruction | Resolution: 7.5 Å / Resolution method: FSC 3 SIGMA CUT-OFF / Num. of particles: 16000 / Symmetry type: POINT | ||||||||||||
| Atomic model building | Space: REAL Details: DETAILS--OTHER REFINEMENT REMARKS- CRYO-EM RECONSTRUCTION THIS FILE HAS BEEN GENERATED BY THE USE OF ALL RELEVANT BIOCHEMICAL CONSTRAINTS AND THE CONSTRAINTS GIVEN BY THE ELECTRON DENSITY ...Details: DETAILS--OTHER REFINEMENT REMARKS- CRYO-EM RECONSTRUCTION THIS FILE HAS BEEN GENERATED BY THE USE OF ALL RELEVANT BIOCHEMICAL CONSTRAINTS AND THE CONSTRAINTS GIVEN BY THE ELECTRON DENSITY CONTOUR OF THE RIBOSOME, WHICH WAS DERIVED FROM THE CRYO-ELECTRON MICROSCOPIC RECONSTRUCTION. | ||||||||||||
| Atomic model building |
| ||||||||||||
| Refinement | Highest resolution: 7.5 Å | ||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 7.5 Å
|
 Movie
Movie Controller
Controller





 PDBj
PDBj








