[English] 日本語
 Yorodumi
Yorodumi- PDB-5a7x: negative stain EM of BG505 SOSIP.664 in complex with sCD4, 17b, a... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5a7x | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
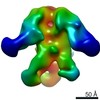
| Title | negative stain EM of BG505 SOSIP.664 in complex with sCD4, 17b, and 8ANC195 | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | VIRAL PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationhelper T cell enhancement of adaptive immune response / interleukin-16 binding / interleukin-16 receptor activity / maintenance of protein location in cell / T cell selection / MHC class II protein binding / cellular response to granulocyte macrophage colony-stimulating factor stimulus / interleukin-15-mediated signaling pathway / positive regulation of monocyte differentiation / Nef Mediated CD4 Down-regulation ...helper T cell enhancement of adaptive immune response / interleukin-16 binding / interleukin-16 receptor activity / maintenance of protein location in cell / T cell selection / MHC class II protein binding / cellular response to granulocyte macrophage colony-stimulating factor stimulus / interleukin-15-mediated signaling pathway / positive regulation of monocyte differentiation / Nef Mediated CD4 Down-regulation / Alpha-defensins / positive regulation of kinase activity / regulation of T cell activation / extracellular matrix structural constituent / T cell receptor complex / Other interleukin signaling / enzyme-linked receptor protein signaling pathway / Dectin-2 family / Translocation of ZAP-70 to Immunological synapse / Phosphorylation of CD3 and TCR zeta chains / regulation of calcium ion transport / macrophage differentiation / Generation of second messenger molecules / T cell differentiation / PD-1 signaling / positive regulation of protein kinase activity / Binding and entry of HIV virion / coreceptor activity / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / cell surface receptor protein tyrosine kinase signaling pathway / positive regulation of establishment of T cell polarity / T cell activation / positive regulation of calcium-mediated signaling / positive regulation of interleukin-2 production / protein tyrosine kinase binding / host cell endosome membrane / clathrin-coated endocytic vesicle membrane / Vpu mediated degradation of CD4 / calcium-mediated signaling / transmembrane signaling receptor activity / Cargo recognition for clathrin-mediated endocytosis / positive regulation of peptidyl-tyrosine phosphorylation / positive regulation of T cell activation / Clathrin-mediated endocytosis / virus receptor activity / Downstream TCR signaling / MHC class II protein complex binding / signaling receptor activity / positive regulation of canonical NF-kappaB signal transduction / clathrin-dependent endocytosis of virus by host cell / defense response to Gram-negative bacterium / adaptive immune response / positive regulation of ERK1 and ERK2 cascade / positive regulation of MAPK cascade / cell surface receptor signaling pathway / positive regulation of viral entry into host cell / early endosome / viral protein processing / cell adhesion / positive regulation of protein phosphorylation / membrane raft / immune response / endoplasmic reticulum lumen / external side of plasma membrane / fusion of virus membrane with host plasma membrane / virus-mediated perturbation of host defense response / fusion of virus membrane with host endosome membrane / lipid binding / viral envelope / endoplasmic reticulum membrane / positive regulation of DNA-templated transcription / virion attachment to host cell / protein kinase binding / apoptotic process / enzyme binding / host cell plasma membrane / structural molecule activity / virion membrane / signal transduction / protein homodimerization activity / zinc ion binding / identical protein binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   HUMAN IMMUNODEFICIENCY VIRUS 1 HUMAN IMMUNODEFICIENCY VIRUS 1 HOMO SAPIENS (human) HOMO SAPIENS (human) | |||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / negative staining / Resolution: 17 Å | |||||||||
 Authors Authors | Scharf, L. / Wang, H. / Gao, H. / Chen, S. / McDowall, A. / Bjorkman, P. | |||||||||
 Citation Citation |  Journal: Cell / Year: 2015 Journal: Cell / Year: 2015Title: Broadly Neutralizing Antibody 8ANC195 Recognizes Closed and Open States of HIV-1 Env. Authors: Louise Scharf / Haoqing Wang / Han Gao / Songye Chen / Alasdair W McDowall / Pamela J Bjorkman /  Abstract: The HIV-1 envelope (Env) spike contains limited epitopes for broadly neutralizing antibodies (bNAbs); thus, most neutralizing antibodies are strain specific. The 8ANC195 epitope, defined by crystal ...The HIV-1 envelope (Env) spike contains limited epitopes for broadly neutralizing antibodies (bNAbs); thus, most neutralizing antibodies are strain specific. The 8ANC195 epitope, defined by crystal and electron microscopy (EM) structures of bNAb 8ANC195 complexed with monomeric gp120 and trimeric Env, respectively, spans the gp120 and gp41 Env subunits. To investigate 8ANC195's gp41 epitope at higher resolution, we solved a 3.58 Å crystal structure of 8ANC195 complexed with fully glycosylated Env trimer, revealing 8ANC195 insertion into a glycan shield gap to contact gp120 and gp41 glycans and protein residues. To determine whether 8ANC195 recognizes the CD4-bound open Env conformation that leads to co-receptor binding and fusion, one of several known conformations of virion-associated Env, we solved EM structures of an Env/CD4/CD4-induced antibody/8ANC195 complex. 8ANC195 binding partially closed the CD4-bound trimer, confirming structural plasticity of Env by revealing a previously unseen conformation. 8ANC195's ability to bind different Env conformations suggests advantages for potential therapeutic applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5a7x.cif.gz 5a7x.cif.gz | 788.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5a7x.ent.gz pdb5a7x.ent.gz | 644.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5a7x.json.gz 5a7x.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  5a7x_validation.pdf.gz 5a7x_validation.pdf.gz | 985.9 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  5a7x_full_validation.pdf.gz 5a7x_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  5a7x_validation.xml.gz 5a7x_validation.xml.gz | 131.9 KB | Display | |
| Data in CIF |  5a7x_validation.cif.gz 5a7x_validation.cif.gz | 197.9 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/a7/5a7x https://data.pdbj.org/pub/pdb/validation_reports/a7/5a7x ftp://data.pdbj.org/pub/pdb/validation_reports/a7/5a7x ftp://data.pdbj.org/pub/pdb/validation_reports/a7/5a7x | HTTPS FTP |
-Related structure data
| Related structure data |  3086MC  3096C  5a8hC  5cjxC C: citing same article ( M: map data used to model this data |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Protein , 2 types, 6 molecules AEIBFJ
| #1: Protein | Mass: 34838.691 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   HUMAN IMMUNODEFICIENCY VIRUS 1 / Strain: YU2 / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host: HUMAN IMMUNODEFICIENCY VIRUS 1 / Strain: YU2 / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host:  HOMO SAPIENS (human) / References: UniProt: P35961*PLUS HOMO SAPIENS (human) / References: UniProt: P35961*PLUS#2: Protein | Mass: 20129.896 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host: HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host:  HOMO SAPIENS (human) / References: UniProt: P01730 HOMO SAPIENS (human) / References: UniProt: P01730 |
|---|
-Antibody , 4 types, 12 molecules CGKDHLMOQNPR
| #3: Antibody | Mass: 23399.898 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host: HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host:  HOMO SAPIENS (human) HOMO SAPIENS (human)#4: Antibody | Mass: 24457.387 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host: HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host:  HOMO SAPIENS (human) HOMO SAPIENS (human)#5: Antibody | Mass: 23401.984 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host: HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host:  HOMO SAPIENS (human) HOMO SAPIENS (human)#6: Antibody | Mass: 26125.270 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host: HOMO SAPIENS (human) / Plasmid: PTT5 / Cell line (production host): HEK293-6E / Production host:  HOMO SAPIENS (human) HOMO SAPIENS (human) |
|---|
-Sugars , 1 types, 45 molecules 
| #7: Sugar | ChemComp-NAG / |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: BG505 SOSIP.664 IN COMPLEX WITH 17B, SCD4, AND 8ANC195 Type: COMPLEX |
|---|---|
| Buffer solution | Name: 20MM TRIS, 50 MM NACL / pH: 8 / Details: 20MM TRIS, 50 MM NACL |
| Specimen | Conc.: 0.01 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: YES / Vitrification applied: NO |
| EM staining | Type: NEGATIVE / Material: uranyl acetate |
| Specimen support | Details: HOLEY CARBON |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: FEI TECNAI 12 / Date: Apr 7, 2015 |
|---|---|
| Electron gun | Electron source: LAB6 / Accelerating voltage: 120 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 42000 X / Cs: 2.2 mm |
| Image recording | Film or detector model: GATAN ULTRASCAN 1000 (2k x 2k) |
| Image scans | Num. digital images: 642 |
- Processing
Processing
| EM software |
| ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Details: INDIVIDUAL PARTICLES | ||||||||||||||||
| Symmetry | Point symmetry: C3 (3 fold cyclic) | ||||||||||||||||
| 3D reconstruction | Method: REFERENCE-BASED REFINEMENT / Resolution: 17 Å / Num. of particles: 7174 / Actual pixel size: 2.5 Å Details: SUBMISSION BASED ON EXPERIMENTAL DATA FROM EMDB EMD-3086. (DEPOSITION ID: 13327). Symmetry type: POINT | ||||||||||||||||
| Atomic model building | Protocol: OTHER / Details: METHOD--MANUAL DOCKING AND LOCAL CORRELATION | ||||||||||||||||
| Refinement | Highest resolution: 17 Å | ||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 17 Å
|
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera



 PDBj
PDBj





















