+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2b6b | ||||||
|---|---|---|---|---|---|---|---|
| Title | Cryo EM structure of Dengue complexed with CRD of DC-SIGN | ||||||
 Components Components |
| ||||||
 Keywords Keywords | Virus/Receptor / Cryo EM dengue CRD DC-SIGN / Icosahedral virus / Virus-Receptor COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationB cell adhesion / cell-cell recognition / modulation by virus of host process / intracellular transport of virus / peptide antigen transport / Butyrophilin (BTN) family interactions / positive regulation of viral life cycle / virion binding / heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules / leukocyte cell-cell adhesion ...B cell adhesion / cell-cell recognition / modulation by virus of host process / intracellular transport of virus / peptide antigen transport / Butyrophilin (BTN) family interactions / positive regulation of viral life cycle / virion binding / heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules / leukocyte cell-cell adhesion / regulation of T cell proliferation / RSV-host interactions / stimulatory C-type lectin receptor signaling pathway / D-mannose binding / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of host TYK2 activity / antigen processing and presentation / host cell mitochondrion / symbiont-mediated suppression of host JAK-STAT cascade via inhibition of STAT2 activity / positive regulation of T cell proliferation / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / CD209 (DC-SIGN) signaling / ribonucleoside triphosphate phosphatase activity / viral genome replication / endocytosis / peptide antigen binding / viral capsid / host cell / double-stranded RNA binding / virus receptor activity / protein complex oligomerization / monoatomic ion channel activity / carbohydrate binding / clathrin-dependent endocytosis of virus by host cell / mRNA (nucleoside-2'-O-)-methyltransferase activity / adaptive immune response / mRNA 5'-cap (guanine-N7-)-methyltransferase activity / membrane => GO:0016020 / RNA helicase activity / host cell endoplasmic reticulum membrane / protein dimerization activity / intracellular signal transduction / symbiont-mediated suppression of host type I interferon-mediated signaling pathway / symbiont entry into host cell / induction by virus of host autophagy / immune response / external side of plasma membrane / serine-type endopeptidase activity / innate immune response / viral RNA genome replication / virus-mediated perturbation of host defense response / RNA-dependent RNA polymerase activity / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / structural molecule activity / virion membrane / cell surface / proteolysis / extracellular region / ATP binding / membrane / metal ion binding / plasma membrane / cytoplasm Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human)  Dengue virus Dengue virus | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 25 Å | ||||||
 Authors Authors | Pokidysheva, E. / Zhang, Y. / Battisti, A.J. / Bator-Kelly, C.M. / Chipman, P.R. / Gregorio, G. / Hendrickson, W.A. / Kuhn, R.J. / Rossmann, M.G. | ||||||
 Citation Citation |  Journal: Cell / Year: 2006 Journal: Cell / Year: 2006Title: Cryo-EM reconstruction of dengue virus in complex with the carbohydrate recognition domain of DC-SIGN. Authors: Elena Pokidysheva / Ying Zhang / Anthony J Battisti / Carol M Bator-Kelly / Paul R Chipman / Chuan Xiao / G Glenn Gregorio / Wayne A Hendrickson / Richard J Kuhn / Michael G Rossmann /  Abstract: Dengue virus (DENV) is a significant human pathogen that causes millions of infections and results in about 24,000 deaths each year. Dendritic cell-specific ICAM3 grabbing nonintegrin (DC-SIGN), ...Dengue virus (DENV) is a significant human pathogen that causes millions of infections and results in about 24,000 deaths each year. Dendritic cell-specific ICAM3 grabbing nonintegrin (DC-SIGN), abundant in immature dendritic cells, was previously reported as being an ancillary receptor interacting with the surface of DENV. The structure of DENV in complex with the carbohydrate recognition domain (CRD) of DC-SIGN was determined by cryo-electron microscopy at 25 A resolution. One CRD monomer was found to bind to two glycosylation sites at Asn67 of two neighboring glycoproteins in each icosahedral asymmetric unit, leaving the third Asn67 residue vacant. The vacancy at the third Asn67 site is a result of the nonequivalence of the glycoprotein environments, leaving space for the primary receptor binding to domain III of E. The use of carbohydrate moieties for receptor binding sites suggests a mechanism for avoiding immune surveillance. | ||||||
| History |
| ||||||
| Remark 999 | SEQUENCE The proteins in this entry contain CA only. |
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2b6b.cif.gz 2b6b.cif.gz | 55.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2b6b.ent.gz pdb2b6b.ent.gz | 32.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2b6b.json.gz 2b6b.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2b6b_validation.pdf.gz 2b6b_validation.pdf.gz | 750 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2b6b_full_validation.pdf.gz 2b6b_full_validation.pdf.gz | 749.5 KB | Display | |
| Data in XML |  2b6b_validation.xml.gz 2b6b_validation.xml.gz | 20.9 KB | Display | |
| Data in CIF |  2b6b_validation.cif.gz 2b6b_validation.cif.gz | 30.8 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/b6/2b6b https://data.pdbj.org/pub/pdb/validation_reports/b6/2b6b ftp://data.pdbj.org/pub/pdb/validation_reports/b6/2b6b ftp://data.pdbj.org/pub/pdb/validation_reports/b6/2b6b | HTTPS FTP |
-Related structure data
| Related structure data |  1166MC 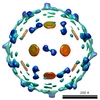 1167MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 | x 60
|
| 2 |
|
| 3 | x 5
|
| 4 | x 6
|
| 5 | 
|
| Symmetry | Point symmetry: (Hermann–Mauguin notation: 532 / Schoenflies symbol: I (icosahedral)) |
- Components
Components
| #1: Protein | Mass: 43819.391 Da / Num. of mol.: 3 / Source method: isolated from a natural source / Source: (natural)   Dengue virus / Genus: Flavivirus / References: GenBank: 323503, UniProt: Q9WDA7*PLUS Dengue virus / Genus: Flavivirus / References: GenBank: 323503, UniProt: Q9WDA7*PLUS#2: Protein | | Mass: 19920.029 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: CD209 / Production host: Homo sapiens (human) / Gene: CD209 / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: DENGUE VIRUS COMPLEXED WITH CRD OF DC-SIGN / Type: VIRUS |
|---|---|
| Details of virus | Host category: INSECT / Type: VIRION |
| Natural host | Strain: C6/36 |
| Buffer solution | pH: 7.5 Details: 50mM Tris 50mM NaCl 0.5 mM EDTA, 5 mM CaCl2, pH 7.5 |
| Specimen | Conc.: 20 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Vitrification | Instrument: HOMEMADE PLUNGER / Cryogen name: ETHANE Details: SAMPLES WERE PREPARED AS THIN LAYERS OF VITREOUS ICE AND MAINTAINED AT LIQUID NITROGEN TEMPERATURE IN THE ELECTRON MICROSCOPE |
- Electron microscopy imaging
Electron microscopy imaging
| Microscopy | Model: FEI/PHILIPS CM300FEG/T / Date: Nov 15, 2004 |
|---|---|
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 50000 X / Nominal defocus max: 3000 nm / Nominal defocus min: 1100 nm / Cs: 2 mm |
| Specimen holder | Temperature: 87 K / Tilt angle max: 0 ° / Tilt angle min: 0 ° |
| Image recording | Electron dose: 27 e/Å2 / Film or detector model: KODAK SO-163 FILM |
- Processing
Processing
| EM software |
| |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Details: EACH VIRAL IMAGE WAS CTF CORRECTED BEFORE RECONSTRUCTION, BASED ON THE FOLLOWING EQUATION: F(CORR)=F(OBS)/[|CTF|+WIENER*(1-|CTF|)] | |||||||||||||||||||||
| Symmetry | Point symmetry: I (icosahedral) | |||||||||||||||||||||
| 3D reconstruction | Method: MODEL-BASED / Resolution: 25 Å / Num. of particles: 830 / Nominal pixel size: 2.95 Å / Symmetry type: POINT | |||||||||||||||||||||
| Atomic model building | Protocol: OTHER / Details: METHOD--PLEASE SEE CITATION | |||||||||||||||||||||
| Atomic model building |
| |||||||||||||||||||||
| Refinement step | Cycle: LAST
|
 Movie
Movie Controller
Controller











 PDBj
PDBj












