+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8014 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Body region of the U4/U6.U5 tri-snRNP | |||||||||
 Map data Map data | Central region of the U4/U6.U5 tri-snRNP comprising Prp8, Dib1, Prp31, Prp6, Prp4, Prp3, Snu13, U4 snRNA and U6 snRNA | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationspliceosomal conformational changes to generate catalytic conformation / snoRNA splicing / snoRNA guided rRNA 2'-O-methylation / positive regulation of RNA binding / box C/D sno(s)RNA 3'-end processing / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / box C/D methylation guide snoRNP complex / U4/U6 snRNP / spliceosomal tri-snRNP complex /  U4 snRNA binding ...spliceosomal conformational changes to generate catalytic conformation / snoRNA splicing / snoRNA guided rRNA 2'-O-methylation / positive regulation of RNA binding / box C/D sno(s)RNA 3'-end processing / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / box C/D methylation guide snoRNP complex / U4/U6 snRNP / spliceosomal tri-snRNP complex / U4 snRNA binding ...spliceosomal conformational changes to generate catalytic conformation / snoRNA splicing / snoRNA guided rRNA 2'-O-methylation / positive regulation of RNA binding / box C/D sno(s)RNA 3'-end processing / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / box C/D methylation guide snoRNP complex / U4/U6 snRNP / spliceosomal tri-snRNP complex /  U4 snRNA binding / U4 snRNP / U3 snoRNA binding / precatalytic spliceosome / spliceosomal complex assembly / generation of catalytic spliceosome for second transesterification step / Major pathway of rRNA processing in the nucleolus and cytosol / mRNA 3'-splice site recognition / mRNA 5'-splice site recognition / spliceosomal tri-snRNP complex assembly / U4 snRNA binding / U4 snRNP / U3 snoRNA binding / precatalytic spliceosome / spliceosomal complex assembly / generation of catalytic spliceosome for second transesterification step / Major pathway of rRNA processing in the nucleolus and cytosol / mRNA 3'-splice site recognition / mRNA 5'-splice site recognition / spliceosomal tri-snRNP complex assembly /  U5 snRNA binding / U5 snRNP / spliceosomal snRNP assembly / U5 snRNA binding / U5 snRNP / spliceosomal snRNP assembly /  U2 snRNA binding / U2 snRNA binding /  U6 snRNA binding / pre-mRNA intronic binding / maturation of SSU-rRNA / U6 snRNA binding / pre-mRNA intronic binding / maturation of SSU-rRNA /  U1 snRNA binding / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of LSU-rRNA / U4/U6 x U5 tri-snRNP complex / catalytic step 2 spliceosome / small-subunit processome / U1 snRNA binding / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of LSU-rRNA / U4/U6 x U5 tri-snRNP complex / catalytic step 2 spliceosome / small-subunit processome /  spliceosomal complex / spliceosomal complex /  mRNA splicing, via spliceosome / mRNA splicing, via spliceosome /  metallopeptidase activity / cytosolic large ribosomal subunit / metallopeptidase activity / cytosolic large ribosomal subunit /  RNA helicase activity / RNA helicase activity /  nucleic acid binding / nucleic acid binding /  RNA helicase / RNA helicase /  mRNA binding / mRNA binding /  nucleolus / nucleolus /  ATP hydrolysis activity / ATP hydrolysis activity /  mitochondrion / mitochondrion /  RNA binding / RNA binding /  nucleoplasm / nucleoplasm /  ATP binding / identical protein binding / ATP binding / identical protein binding /  nucleus / nucleus /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Saccharomyces cerevisiae (brewer's yeast) / Saccharomyces cerevisiae (brewer's yeast) /   Baker's yeast (brewer's yeast) Baker's yeast (brewer's yeast) | |||||||||
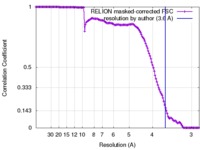
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.6 Å cryo EM / Resolution: 3.6 Å | |||||||||
 Authors Authors | Nguyen THD / Galej WP / Oubridge C / Bai XC / Newman A / Scheres S / Nagai K | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2016 Journal: Nature / Year: 2016Title: Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution. Authors: Thi Hoang Duong Nguyen / Wojciech P Galej / Xiao-Chen Bai / Chris Oubridge / Andrew J Newman / Sjors H W Scheres / Kiyoshi Nagai /  Abstract: U4/U6.U5 tri-snRNP represents a substantial part of the spliceosome before activation. A cryo-electron microscopy structure of Saccharomyces cerevisiae U4/U6.U5 tri-snRNP at 3.7 Å resolution led ...U4/U6.U5 tri-snRNP represents a substantial part of the spliceosome before activation. A cryo-electron microscopy structure of Saccharomyces cerevisiae U4/U6.U5 tri-snRNP at 3.7 Å resolution led to an essentially complete atomic model comprising 30 proteins plus U4/U6 and U5 small nuclear RNAs (snRNAs). The structure reveals striking interweaving interactions of the protein and RNA components, including extended polypeptides penetrating into subunit interfaces. The invariant ACAGAGA sequence of U6 snRNA, which base-pairs with the 5'-splice site during catalytic activation, forms a hairpin stabilized by Dib1 and Prp8 while the adjacent nucleotides interact with the exon binding loop 1 of U5 snRNA. Snu114 harbours GTP, but its putative catalytic histidine is held away from the γ-phosphate by hydrogen bonding to a tyrosine in the amino-terminal domain of Prp8. Mutation of this histidine to alanine has no detectable effect on yeast growth. The structure provides important new insights into the spliceosome activation process leading to the formation of the catalytic centre. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8014.map.gz emd_8014.map.gz | 196.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8014-v30.xml emd-8014-v30.xml emd-8014.xml emd-8014.xml | 28.9 KB 28.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8014_fsc.xml emd_8014_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_8014.png emd_8014.png | 233.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8014 http://ftp.pdbj.org/pub/emdb/structures/EMD-8014 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8014 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8014 | HTTPS FTP |
-Related structure data
| Related structure data |  5gapMC  8006C  8007C  8008C  8009C  8010C  8011C  8012C  8013C  5gamC  5ganC  5gaoC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8014.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8014.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Central region of the U4/U6.U5 tri-snRNP comprising Prp8, Dib1, Prp31, Prp6, Prp4, Prp3, Snu13, U4 snRNA and U6 snRNA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.43 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
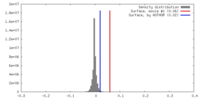
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Body region of the U4/U6.U5 tri-snRNP complex
+Supramolecule #1: Body region of the U4/U6.U5 tri-snRNP complex
+Macromolecule #1: U4 snRNA, 5' region, nucleotides 1-67
+Macromolecule #2: U6 snRNA
+Macromolecule #3: U5 snRNA
+Macromolecule #4: unknown protein
+Macromolecule #5: Pre-mRNA-splicing factor 8
+Macromolecule #6: U4/U6 small nuclear ribonucleoprotein PRP4
+Macromolecule #7: Pre-mRNA-splicing factor 6
+Macromolecule #8: Spliceosomal protein DIB1
+Macromolecule #9: Pre-mRNA-processing factor 31
+Macromolecule #10: U4/U6 small nuclear ribonucleoprotein PRP3
+Macromolecule #11: 13 kDa ribonucleoprotein-associated protein
+Macromolecule #12: Pre-mRNA-splicing helicase BRR2
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 7.9 / Component - Concentration: 1.0 mM / Component - Name: DTT |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 6.0 nm / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: OTHER |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III Details: Grids were blotted at 4 deg C for 2 seconds before plunging.. |
| Details | 3.5 microlitre of sample was applied to grid. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 35714 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.0 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 81000 Bright-field microscopy / Cs: 2.0 mm / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 81000 |
| Specialist optics | Energy filter - Name: GIF Quantum |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-20 / Number real images: 2477 / Average exposure time: 16.0 sec. / Average electron dose: 38.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-5gap: |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)