[English] 日本語
 Yorodumi
Yorodumi- EMDB-6235: Negative stain single particle electron microscopy of Marburg Vir... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6235 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Negative stain single particle electron microscopy of Marburg Virus Glycoproteins bound to antibodies from a human survivor | |||||||||
 Map data Map data | Reconstruction of Marburg Virus GP bound to antibody MR82 from a human survivor | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Marburg Virus / Antibodies / Neutralization / Glycoprotein | |||||||||
| Biological species |  Marburg marburgvirus / Marburg marburgvirus /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 25.0 Å | |||||||||
 Authors Authors | Flyak A / Ilinykh PA / Murin CD / Garron T / Shen X / Fusco ML / Hashiguchi T / Bornholdt ZA / Slaughter C / Sapparapu G ...Flyak A / Ilinykh PA / Murin CD / Garron T / Shen X / Fusco ML / Hashiguchi T / Bornholdt ZA / Slaughter C / Sapparapu G / Ksiazek TG / Ward AB / Saphire EO / Bukreyev A / Crowe JE | |||||||||
 Citation Citation |  Journal: Cell / Year: 2015 Journal: Cell / Year: 2015Title: Mechanism of human antibody-mediated neutralization of Marburg virus. Authors: Andrew I Flyak / Philipp A Ilinykh / Charles D Murin / Tania Garron / Xiaoli Shen / Marnie L Fusco / Takao Hashiguchi / Zachary A Bornholdt / James C Slaughter / Gopal Sapparapu / Curtis ...Authors: Andrew I Flyak / Philipp A Ilinykh / Charles D Murin / Tania Garron / Xiaoli Shen / Marnie L Fusco / Takao Hashiguchi / Zachary A Bornholdt / James C Slaughter / Gopal Sapparapu / Curtis Klages / Thomas G Ksiazek / Andrew B Ward / Erica Ollmann Saphire / Alexander Bukreyev / James E Crowe /  Abstract: The mechanisms by which neutralizing antibodies inhibit Marburg virus (MARV) are not known. We isolated a panel of neutralizing antibodies from a human MARV survivor that bind to MARV glycoprotein ...The mechanisms by which neutralizing antibodies inhibit Marburg virus (MARV) are not known. We isolated a panel of neutralizing antibodies from a human MARV survivor that bind to MARV glycoprotein (GP) and compete for binding to a single major antigenic site. Remarkably, several of the antibodies also bind to Ebola virus (EBOV) GP. Single-particle EM structures of antibody-GP complexes reveal that all of the neutralizing antibodies bind to MARV GP at or near the predicted region of the receptor-binding site. The presence of the glycan cap or mucin-like domain blocks binding of neutralizing antibodies to EBOV GP, but not to MARV GP. The data suggest that MARV-neutralizing antibodies inhibit virus by binding to infectious virions at the exposed MARV receptor-binding site, revealing a mechanism of filovirus inhibition. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6235.map.gz emd_6235.map.gz | 2.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6235-v30.xml emd-6235-v30.xml emd-6235.xml emd-6235.xml | 11 KB 11 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6235.jpg emd_6235.jpg | 585.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6235 http://ftp.pdbj.org/pub/emdb/structures/EMD-6235 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6235 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6235 | HTTPS FTP |
-Validation report
| Summary document |  emd_6235_validation.pdf.gz emd_6235_validation.pdf.gz | 78.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6235_full_validation.pdf.gz emd_6235_full_validation.pdf.gz | 77.7 KB | Display | |
| Data in XML |  emd_6235_validation.xml.gz emd_6235_validation.xml.gz | 493 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6235 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6235 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6235 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6235 | HTTPS FTP |
-Related structure data
| Related structure data |  6232C  6233C  6234C 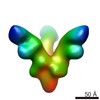 6236C  6237C  6238C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6235.map.gz / Format: CCP4 / Size: 3.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6235.map.gz / Format: CCP4 / Size: 3.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of Marburg Virus GP bound to antibody MR82 from a human survivor | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Fab Fragment of MR82 human IgG1 monoclonal antibody to Marburg Vi...
| Entire | Name: Fab Fragment of MR82 human IgG1 monoclonal antibody to Marburg Virus GP |
|---|---|
| Components |
|
-Supramolecule #1000: Fab Fragment of MR82 human IgG1 monoclonal antibody to Marburg Vi...
| Supramolecule | Name: Fab Fragment of MR82 human IgG1 monoclonal antibody to Marburg Virus GP type: sample / ID: 1000 / Oligomeric state: trimer / Number unique components: 2 |
|---|---|
| Molecular weight | Theoretical: 310 KDa |
-Macromolecule #1: GP, mucin-deleted
| Macromolecule | Name: GP, mucin-deleted / type: protein_or_peptide / ID: 1 / Name.synonym: glycoprotein Details: Recombinant Marburg Virus GP; mucin-like domains were not included in the construct. Number of copies: 1 / Oligomeric state: trimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Marburg marburgvirus / synonym: Marburg virus Marburg marburgvirus / synonym: Marburg virus |
| Molecular weight | Theoretical: 160 MDa |
| Recombinant expression | Organism:  |
-Macromolecule #2: Human Monoclonal Antibody IgG1
| Macromolecule | Name: Human Monoclonal Antibody IgG1 / type: protein_or_peptide / ID: 2 / Name.synonym: IgG1, monoclonal mAb / Details: Human monoclonal antibody IgG1, MR82 / Number of copies: 1 / Oligomeric state: monomer / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human / Tissue: Blood / Cell: Peripheral blood mononuclear cells Homo sapiens (human) / synonym: Human / Tissue: Blood / Cell: Peripheral blood mononuclear cells |
| Molecular weight | Theoretical: 150 KDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.03 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 20 mM Tris, 150 mM NaCl |
| Staining | Type: NEGATIVE Details: Grids with absorbed protein stained with 2% uranyl formate for 30 seconds |
| Grid | Details: 400 mesh copper grid with carbon support, plasma cleaned 20 seconds (Ar/O2 mix, Gatan) |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Date | Nov 12, 2013 |
| Image recording | Category: CCD / Film or detector model: OTHER / Number real images: 166 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 52000 |
| Sample stage | Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Particles were selected using DoG picker. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 25.0 Å / Resolution method: OTHER / Software - Name: EMAN2 / Number images used: 2554 |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera

