+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5725 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
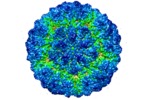
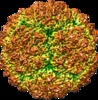
| Title | Three-dimensional Structure of Victorivirus HvV190S | |||||||||
 Map data Map data | Reconstruction of HvV190S virion | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Helminthosporium victoriae virus 190S Helminthosporium victoriae virus 190S | |||||||||
| Method | single particle reconstruction / cryo EM / negative staining / Resolution: 7.1 Å | |||||||||
 Authors Authors | Dun SE / Li H / Cardone G / Nibert ML / Gabrial SA / Baker TS | |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2013 Journal: PLoS Pathog / Year: 2013Title: Three-dimensional structure of victorivirus HvV190S suggests coat proteins in most totiviruses share a conserved core. Authors: Sarah E Dunn / Hua Li / Giovanni Cardone / Max L Nibert / Said A Ghabrial / Timothy S Baker /  Abstract: Double-stranded (ds)RNA fungal viruses are currently assigned to six different families. Those from the family Totiviridae are characterized by nonsegmented genomes and single-layer capsids, 300-450 ...Double-stranded (ds)RNA fungal viruses are currently assigned to six different families. Those from the family Totiviridae are characterized by nonsegmented genomes and single-layer capsids, 300-450 Å in diameter. Helminthosporium victoriae virus 190S (HvV190S), prototype of recently recognized genus Victorivirus, infects the filamentous fungus Helminthosporium victoriae (telomorph: Cochliobolus victoriae), which is the causal agent of Victoria blight of oats. The HvV190S genome is 5179 bp long and encompasses two large, slightly overlapping open reading frames that encode the coat protein (CP, 772 aa) and the RNA-dependent RNA polymerase (RdRp, 835 aa). To our present knowledge, victoriviruses uniquely express their RdRps via a coupled termination-reinitiation mechanism that differs from the well-characterized Saccharomyces cerevisiae virus L-A (ScV-L-A, prototype of genus Totivirus), in which the RdRp is expressed as a CP/RdRp fusion protein due to ribosomal frameshifting. Here, we used transmission electron cryomicroscopy and three-dimensional image reconstruction to determine the structures of HvV190S virions and two types of virus-like particles (capsids lacking dsRNA and capsids lacking both dsRNA and RdRp) at estimated resolutions of 7.1, 7.5, and 7.6 Å, respectively. The HvV190S capsid is thin and smooth, and contains 120 copies of CP arranged in a "T = 2" icosahedral lattice characteristic of ScV-L-A and other dsRNA viruses. For aid in our interpretations, we developed and used an iterative segmentation procedure to define the boundaries of the two, chemically identical CP subunits in each asymmetric unit. Both subunits have a similar fold, but one that differs from ScV-L-A in many details except for a core α-helical region that is further predicted to be conserved among many other totiviruses. In particular, we predict the structures of other victoriviruses to be highly similar to HvV190S and the structures of most if not all totiviruses including, Leishmania RNA virus 1, to be similar as well. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5725.map.gz emd_5725.map.gz | 29.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5725-v30.xml emd-5725-v30.xml emd-5725.xml emd-5725.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5725_1.jpg emd_5725_1.jpg | 270.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5725 http://ftp.pdbj.org/pub/emdb/structures/EMD-5725 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5725 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5725 | HTTPS FTP |
-Validation report
| Summary document |  emd_5725_validation.pdf.gz emd_5725_validation.pdf.gz | 78.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5725_full_validation.pdf.gz emd_5725_full_validation.pdf.gz | 77.3 KB | Display | |
| Data in XML |  emd_5725_validation.xml.gz emd_5725_validation.xml.gz | 492 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5725 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5725 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5725 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5725 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5725.map.gz / Format: CCP4 / Size: 234.2 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) Download / File: emd_5725.map.gz / Format: CCP4 / Size: 234.2 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of HvV190S virion | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Helminthosporium victoriae virus 190S
| Entire | Name:  Helminthosporium victoriae virus 190S Helminthosporium victoriae virus 190S |
|---|---|
| Components |
|
-Supramolecule #1000: Helminthosporium victoriae virus 190S
| Supramolecule | Name: Helminthosporium victoriae virus 190S / type: sample / ID: 1000 / Oligomeric state: Icosahedral / Number unique components: 1 |
|---|
-Supramolecule #1: Helminthosporium victoriae virus 190S
| Supramolecule | Name: Helminthosporium victoriae virus 190S / type: virus / ID: 1 / Name.synonym: HvV190S / NCBI-ID: 45237 / Sci species name: Helminthosporium victoriae virus 190S / Sci species strain: A-9 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No / Syn species name: HvV190S |
|---|---|
| Host (natural) | Organism:  Cochliobolus victoriae (fungus) / synonym: FUNGI Cochliobolus victoriae (fungus) / synonym: FUNGI |
| Host system | Organism: unidentified (others) |
| Virus shell | Shell ID: 1 / Diameter: 462 Å / T number (triangulation number): 2 |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 5 mg/mL |
|---|---|
| Buffer | pH: 7.8 / Details: 50 mM Tris-HCl, 5 mM EDTA, 150 mM NaCl, pH 7.8 |
| Staining | Type: NEGATIVE Details: Samples were adsorbed to continuous carbon grids that had been glow-discharged for ~25 seconds in an Emitech K350 evaporation unit. Samples were subsequently stained with 1% aqueous uranyl ...Details: Samples were adsorbed to continuous carbon grids that had been glow-discharged for ~25 seconds in an Emitech K350 evaporation unit. Samples were subsequently stained with 1% aqueous uranyl acetate and rinsed with ddH2O. |
| Grid | Details: 200 mesh gold grid with thin carbon support |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 90 K / Instrument: FEI VITROBOT MARK III Details: Vitrification carried out after applying sample to carbon-coated grids and waiting 5 minutes before inserting the grid into the Vitrobot. Method: Blot for 3.5 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Temperature | Min: 90 K / Max: 92 K / Average: 91 K |
| Alignment procedure | Legacy - Astigmatism: Objective lens astigmatism was corrected at 120,000 times magnification using FFT |
| Date | Aug 8, 2009 |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 1.09 µm / Number real images: 176 / Average electron dose: 24 e/Å2 / Bits/pixel: 8 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 59318 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.3 mm / Nominal defocus max: 3.8 µm / Nominal defocus min: 0.72 µm / Nominal magnification: 59000 |
| Sample stage | Specimen holder model: OTHER |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: ROBEM |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 7.1 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: AUTO3DEM / Number images used: 20904 |
 Movie
Movie Controller
Controller