[English] 日本語
 Yorodumi
Yorodumi- EMDB-2420: Electron microscopy of the CSM/RNA complex from Sulfolobus solfat... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2420 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron microscopy of the CSM/RNA complex from Sulfolobus solfataricus | |||||||||
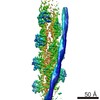
 Map data Map data | Structure of CSM/RNA complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CRISPR / CSM / Sulfolobus solfataricus | |||||||||
| Biological species |   Sulfolobus solfataricus (archaea) Sulfolobus solfataricus (archaea) | |||||||||
| Method | single particle reconstruction / Resolution: 30.0 Å | |||||||||
 Authors Authors | Rouillon C / Zhou M / Zhang J / Politis A / Beilsten V / Cannone G / Graham S / Robinson CV / Spagnolo L / White MF | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2013 Journal: Mol Cell / Year: 2013Title: Structure of the CRISPR interference complex CSM reveals key similarities with cascade. Authors: Christophe Rouillon / Min Zhou / Jing Zhang / Argyris Politis / Victoria Beilsten-Edmands / Giuseppe Cannone / Shirley Graham / Carol V Robinson / Laura Spagnolo / Malcolm F White /  Abstract: The Clustered Regularly Interspaced Palindromic Repeats (CRISPR) system is an adaptive immune system in prokaryotes. Interference complexes encoded by CRISPR-associated (cas) genes utilize small RNAs ...The Clustered Regularly Interspaced Palindromic Repeats (CRISPR) system is an adaptive immune system in prokaryotes. Interference complexes encoded by CRISPR-associated (cas) genes utilize small RNAs for homology-directed detection and subsequent degradation of invading genetic elements, and they have been classified into three main types (I-III). Type III complexes share the Cas10 subunit but are subclassifed as type IIIA (CSM) and type IIIB (CMR), depending on their specificity for DNA or RNA targets, respectively. The role of CSM in limiting the spread of conjugative plasmids in Staphylococcus epidermidis was first described in 2008. Here, we report a detailed investigation of the composition and structure of the CSM complex from the archaeon Sulfolobus solfataricus, using a combination of electron microscopy, mass spectrometry, and deep sequencing. This reveals a three-dimensional model for the CSM complex that includes a helical component strikingly reminiscent of the backbone structure of the type I (Cascade) family. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2420.map.gz emd_2420.map.gz | 346 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2420-v30.xml emd-2420-v30.xml emd-2420.xml emd-2420.xml | 6.7 KB 6.7 KB | Display Display |  EMDB header EMDB header |
| Images |  EMD-2420.png EMD-2420.png | 71.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2420 http://ftp.pdbj.org/pub/emdb/structures/EMD-2420 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2420 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2420 | HTTPS FTP |
-Validation report
| Summary document |  emd_2420_validation.pdf.gz emd_2420_validation.pdf.gz | 196.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2420_full_validation.pdf.gz emd_2420_full_validation.pdf.gz | 196.1 KB | Display | |
| Data in XML |  emd_2420_validation.xml.gz emd_2420_validation.xml.gz | 5.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2420 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2420 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2420 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2420 | HTTPS FTP |
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2420.map.gz / Format: CCP4 / Size: 1001 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2420.map.gz / Format: CCP4 / Size: 1001 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of CSM/RNA complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 7.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : CSM/RNA complex from Sulfolobus solfataricus
| Entire | Name: CSM/RNA complex from Sulfolobus solfataricus |
|---|---|
| Components |
|
-Supramolecule #1000: CSM/RNA complex from Sulfolobus solfataricus
| Supramolecule | Name: CSM/RNA complex from Sulfolobus solfataricus / type: sample / ID: 1000 / Number unique components: 1 |
|---|---|
| Molecular weight | Experimental: 428 KDa |
-Macromolecule #1: CSM complex
| Macromolecule | Name: CSM complex / type: protein_or_peptide / ID: 1 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:   Sulfolobus solfataricus (archaea) Sulfolobus solfataricus (archaea) |
| Recombinant expression | Organism:   Sulfolobus solfataricus (archaea) Sulfolobus solfataricus (archaea) |
-Experimental details
-Structure determination
 Processing Processing | single particle reconstruction |
|---|---|
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Dec 13, 2012 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 30.0 Å / Resolution method: OTHER / Number images used: 7829 |
|---|
 Movie
Movie Controller
Controller