[English] 日本語
 Yorodumi
Yorodumi- EMDB-2218: DOLORS: Versatile Strategy for Internal Labeling and Domain Local... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2218 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | DOLORS: Versatile Strategy for Internal Labeling and Domain Localization in Electron Microscopy | |||||||||
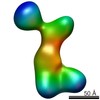
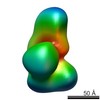
 Map data Map data | DUF283-After-labeled Dicer | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Dicer Enzyme / Ribonuclease III / MicroRNA Processing / Single Particle Electron Microscopy | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  Streptomyces avidinii (bacteria) Streptomyces avidinii (bacteria) | |||||||||
| Method | single particle reconstruction / negative staining | |||||||||
 Authors Authors | Lau PW / Potter CS / Carragher B / MacRae IJ | |||||||||
 Citation Citation |  Journal: Structure / Year: 2012 Journal: Structure / Year: 2012Title: DOLORS: versatile strategy for internal labeling and domain localization in electron microscopy. Authors: Pick-Wei Lau / Clinton S Potter / Bridget Carragher / Ian J MacRae /  Abstract: Single-particle electron microscopy (EM) is a powerful tool for studying the structures of large biological molecules. However, the achievable resolution does not always allow for direct recognition ...Single-particle electron microscopy (EM) is a powerful tool for studying the structures of large biological molecules. However, the achievable resolution does not always allow for direct recognition of individual protein domains. Labels that can be visualized by EM have been developed for protein termini, but tagging internal domains remains a challenge. We describe a robust strategy for determining the position of internal sites within EM maps, termed domain localization by RCT sampling (DOLORS). DOLORS uses monovalent streptavidin added posttranslationally to tagged sites in the target protein. Internal labels generally display less conformational flexibility than terminal labels, providing more precise positional information. Automated methods are used to rapidly generate assemblies of unique 3D models allowing the attachment sites of labeled domains to be accurately identified and thus provide an overall architectural map of the molecule. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2218.map.gz emd_2218.map.gz | 1018.3 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2218-v30.xml emd-2218-v30.xml emd-2218.xml emd-2218.xml | 9 KB 9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_2218.png emd_2218.png | 47.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2218 http://ftp.pdbj.org/pub/emdb/structures/EMD-2218 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2218 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2218 | HTTPS FTP |
-Validation report
| Summary document |  emd_2218_validation.pdf.gz emd_2218_validation.pdf.gz | 190.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2218_full_validation.pdf.gz emd_2218_full_validation.pdf.gz | 189.8 KB | Display | |
| Data in XML |  emd_2218_validation.xml.gz emd_2218_validation.xml.gz | 5.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2218 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2218 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2218 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2218 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2218.map.gz / Format: CCP4 / Size: 3.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2218.map.gz / Format: CCP4 / Size: 3.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | DUF283-After-labeled Dicer | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.52 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Dicer labeled with streptavidin at the loop following the DUF283 ...
| Entire | Name: Dicer labeled with streptavidin at the loop following the DUF283 domain |
|---|---|
| Components |
|
-Supramolecule #1000: Dicer labeled with streptavidin at the loop following the DUF283 ...
| Supramolecule | Name: Dicer labeled with streptavidin at the loop following the DUF283 domain type: sample / ID: 1000 / Details: The sample was mostly monodisperse Oligomeric state: One human Dicer and one streptavidin molecule Number unique components: 2 |
|---|---|
| Molecular weight | Theoretical: 280 KDa |
-Macromolecule #1: Human Dicer
| Macromolecule | Name: Human Dicer / type: protein_or_peptide / ID: 1 / Name.synonym: hDicer / Number of copies: 1 / Oligomeric state: Monomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: Human Homo sapiens (human) / synonym: Human |
| Molecular weight | Theoretical: 220 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #2: Streptavidin
| Macromolecule | Name: Streptavidin / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Oligomeric state: Tetramer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Streptomyces avidinii (bacteria) Streptomyces avidinii (bacteria) |
| Molecular weight | Theoretical: 60 KDa |
| Recombinant expression | Organism:  |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 150mM KCl, 25mM HEPES |
|---|---|
| Staining | Type: NEGATIVE Details: Grids with adsorbed protein floated on 2% w/v uranyl acetate |
| Grid | Details: holey C-flat grids covered with an additional layer of thin carbon |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Jan 10, 2011 |
| Image recording | Number real images: 570 / Average electron dose: 20 e/Å2 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 62000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC / Tilt angle max: 50 |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | Processing was done using APPION pipeline |
|---|---|
| CTF correction | Details: Each micrograph |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Algorithm: OTHER / Software - Name: Spider / Number images used: 48164 |
| Final two d classification | Number classes: 1 |
 Movie
Movie Controller
Controller