[English] 日本語
 Yorodumi
Yorodumi- EMDB-2039: negative stain Electron Microscopy of the N-Ethylmaleimide Sensit... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-2039 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | negative stain Electron Microscopy of the N-Ethylmaleimide Sensitive Factor (NSF) | |||||||||
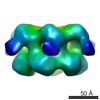
 Map data Map data | NSF-AMP-PNP | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NSF / N-Ethylmaleimide Sensitive Factor | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / Resolution: 20.0 Å | |||||||||
 Authors Authors | Moeller A / Zhao C / Fried MG / Wilson-Kubalek EM / Carragher B / Whiteheart SW | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2012 Journal: J Struct Biol / Year: 2012Title: Nucleotide-dependent conformational changes in the N-Ethylmaleimide Sensitive Factor (NSF) and their potential role in SNARE complex disassembly. Authors: Arne Moeller / Chunxia Zhao / Michael G Fried / Elizabeth M Wilson-Kubalek / Bridget Carragher / Sidney W Whiteheart /  Abstract: Homohexameric, N-Ethylmaleimide Sensitive Factor (NSF) disassembles Soluble NSF Attachment Protein Receptor (SNARE) complexes after membrane fusion, an essential step in vesicular trafficking. NSF ...Homohexameric, N-Ethylmaleimide Sensitive Factor (NSF) disassembles Soluble NSF Attachment Protein Receptor (SNARE) complexes after membrane fusion, an essential step in vesicular trafficking. NSF contains three domains (NSF-N, NSF-D1, and NSF-D2), each contributing to activity. We combined electron microscopic (EM) analysis, analytical ultracentrifugation (AU) and functional mutagenesis to visualize NSF's ATPase cycle. 3D density maps show that NSF-D2 remains stable, whereas NSF-N undergoes large conformational changes. NSF-Ns splay out perpendicular to the ADP-bound hexamer and twist upwards upon ATP binding, producing a more compact structure. These conformations were confirmed by hydrodynamic, AU measurements: NSF-ATP sediments faster with a lower frictional ratio (f/f(0)). Hydrodynamic analyses of NSF mutants, with specific functional defects, define the structures underlying these conformational changes. Mapping mutations onto our 3D models allows interpretation of the domain movement and suggests a mechanism for NSF binding to and disassembly of SNARE complexes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_2039.map.gz emd_2039.map.gz | 1.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-2039-v30.xml emd-2039-v30.xml emd-2039.xml emd-2039.xml | 7.1 KB 7.1 KB | Display Display |  EMDB header EMDB header |
| Images |  2039.jpg 2039.jpg | 171.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-2039 http://ftp.pdbj.org/pub/emdb/structures/EMD-2039 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2039 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-2039 | HTTPS FTP |
-Validation report
| Summary document |  emd_2039_validation.pdf.gz emd_2039_validation.pdf.gz | 220 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_2039_full_validation.pdf.gz emd_2039_full_validation.pdf.gz | 219.2 KB | Display | |
| Data in XML |  emd_2039_validation.xml.gz emd_2039_validation.xml.gz | 4.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2039 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2039 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2039 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-2039 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_2039.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_2039.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | NSF-AMP-PNP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.74 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : NSF-AMP-PNP
| Entire | Name: NSF-AMP-PNP |
|---|---|
| Components |
|
-Supramolecule #1000: NSF-AMP-PNP
| Supramolecule | Name: NSF-AMP-PNP / type: sample / ID: 1000 / Oligomeric state: hexamer / Number unique components: 1 |
|---|
-Macromolecule #1: N-Ethylmaleimide Sensitive Factor
| Macromolecule | Name: N-Ethylmaleimide Sensitive Factor / type: protein_or_peptide / ID: 1 / Name.synonym: NSF / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
 Processing Processing | single particle reconstruction |
|---|---|
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | May 20, 2011 |
| Image recording | Number real images: 858 / Average electron dose: 25 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: -2.0 µm / Nominal defocus min: -0.2 µm / Nominal magnification: 62000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: whole image |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C6 (6 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Xmipp, Appion, Imagic / Number images used: 12978 |
 Movie
Movie Controller
Controller