+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5039 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM reconstruction of the giant Mimivirus using C5 symmetry | |||||||||
 Map data Map data | This is a volume map of Mimivirus reconstrument averaged by C5 symmetry | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Mimivirus / Acanthamoeba polyphaga / cryo-electron microscope / atomic force microscope / fiber / digestion / starfish-shaped feature / 5-fold symmetry / nucleocapsid / major capsid protein / capsomer arrangement | |||||||||
| Biological species |  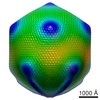 Acanthamoeba polyphaga mimivirus Acanthamoeba polyphaga mimivirus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 65.0 Å | |||||||||
 Authors Authors | Xiao C / Kuznetsov YG / Sun S / Hafenstein SL / Kostyuchenko VA / Chipman PR / Suzan-Monti M / Raoult D / McPherson A / Rossmann MG | |||||||||
 Citation Citation |  Journal: PLoS Biol / Year: 2009 Journal: PLoS Biol / Year: 2009Title: Structural studies of the giant mimivirus. Authors: Chuan Xiao / Yurii G Kuznetsov / Siyang Sun / Susan L Hafenstein / Victor A Kostyuchenko / Paul R Chipman / Marie Suzan-Monti / Didier Raoult / Alexander McPherson / Michael G Rossmann /  Abstract: Mimivirus is the largest known virus whose genome and physical size are comparable to some small bacteria, blurring the boundary between a virus and a cell. Structural studies of Mimivirus have been ...Mimivirus is the largest known virus whose genome and physical size are comparable to some small bacteria, blurring the boundary between a virus and a cell. Structural studies of Mimivirus have been difficult because of its size and long surface fibers. Here we report the use of enzymatic digestions to remove the surface fibers of Mimivirus in order to expose the surface of the viral capsid. Cryo-electron microscopy (cryoEM) and atomic force microscopy were able to show that the 20 icosahedral faces of Mimivirus capsids have hexagonal arrays of depressions. Each depression is surrounded by six trimeric capsomers that are similar in structure to those in many other large, icosahedral double-stranded DNA viruses. Whereas in most viruses these capsomers are hexagonally close-packed with the same orientation in each face, in Mimivirus there are vacancies at the systematic depressions with neighboring capsomers differing in orientation by 60 degrees . The previously observed starfish-shaped feature is well-resolved and found to be on each virus particle and is associated with a special pentameric vertex. The arms of the starfish fit into the gaps between the five faces surrounding the unique vertex, acting as a seal. Furthermore, the enveloped nucleocapsid is accurately positioned and oriented within the capsid with a concave surface facing the unique vertex. Thus, the starfish-shaped feature and the organization of the nucleocapsid might regulate the delivery of the genome to the host. The structure of Mimivirus, as well as the various fiber components observed in the virus, suggests that the Mimivirus genome includes genes derived from both eukaryotic and prokaryotic organisms. The three-dimensional cryoEM reconstruction reported here is of a virus with a volume that is one order of magnitude larger than any previously reported molecular assembly studied at a resolution of equal to or better than 65 Angstroms. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5039.map.gz emd_5039.map.gz | 101.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5039-v30.xml emd-5039-v30.xml emd-5039.xml emd-5039.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_5039_1.tif emd_5039_1.tif | 348.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5039 http://ftp.pdbj.org/pub/emdb/structures/EMD-5039 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5039 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5039 | HTTPS FTP |
-Validation report
| Summary document |  emd_5039_validation.pdf.gz emd_5039_validation.pdf.gz | 79.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_5039_full_validation.pdf.gz emd_5039_full_validation.pdf.gz | 78.2 KB | Display | |
| Data in XML |  emd_5039_validation.xml.gz emd_5039_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5039 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5039 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5039 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-5039 | HTTPS FTP |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_5039.map.gz / Format: CCP4 / Size: 300.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5039.map.gz / Format: CCP4 / Size: 300.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is a volume map of Mimivirus reconstrument averaged by C5 symmetry | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 15.875 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Mimivirus
| Entire | Name:   Mimivirus Mimivirus |
|---|---|
| Components |
|
-Supramolecule #1000: Mimivirus
| Supramolecule | Name: Mimivirus / type: sample / ID: 1000 / Number unique components: 1 |
|---|
-Supramolecule #1: Acanthamoeba polyphaga mimivirus
| Supramolecule | Name: Acanthamoeba polyphaga mimivirus / type: virus / ID: 1 / Name.synonym: Mimivirus / NCBI-ID: 212035 / Sci species name: Acanthamoeba polyphaga mimivirus / Database: NCBI / Virus type: VIRION / Virus isolate: SPECIES / Virus enveloped: No / Virus empty: No / Syn species name: Mimivirus |
|---|---|
| Host (natural) | Organism:  Acanthamoeba polyphaga (eukaryote) / synonym: PROTOZOA Acanthamoeba polyphaga (eukaryote) / synonym: PROTOZOA |
| Virus shell | Shell ID: 1 / Name: Mimivirus capsid / Diameter: 5000 Å |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 / Details: PBS (137 mM NaCl, 10 mM Phosphate, 2.7 mM KCl) |
|---|---|
| Grid | Details: 200 Copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 80 K / Instrument: REICHERT-JUNG PLUNGER Details: Vitrification instrument: Reichert plunger. Using Quantifoil S7 slash 2 grid Method: Blot for 1 seconds before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM300FEG/T |
|---|---|
| Temperature | Average: 80 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 98,000 times magnification |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: NIKON SUPER COOLSCAN 9000 / Digitization - Sampling interval: 38.1 µm / Average electron dose: 20 e/Å2 / Details: 6.35 um scanned then bined 6 times to 38.1 um / Bits/pixel: 16 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 24423 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus min: 4.505 µm / Nominal magnification: 24000 |
| Sample stage | Specimen holder: Side entry liquid nitrogen-cooled cryo specimen holder Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: Each particle |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 65.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: Frealign / Number images used: 30919 |
 Movie
Movie Controller
Controller




